Abstract
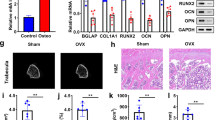
Alteration of N6-methyladenosine (m6A) is closely linked to spanning biological processes including osteoporosis (OP) development. This research focuses on the function of methyltransferase like 14 (METTL14) in bone turnover and its interaction with T cell factor 1 (TCF1). A mouse model of OP was established by ovariectomy (OVX). The bone mass parameters were evaluated by micro-CT analysis. Mouse MC3T3-E1 cells and mouse bone marrow macrophages (BMMs) were induced for osteogenic or osteoclastic differentiation, respectively, for in vitro experiments. The osteogenesis or osteoclasis activity was analyzed by measuring the biomarkers such as OPG, ALP, NFATC1, CTSK, RANKL, and TRAP. RT-qPCR and IHC assays identified reduced METTL14 expression in bone tissues of osteoporotic patients and ovariectomized mice. Artificial METTL14 overexpression increased bone mass of mice and promoted osteogenesis whereas suppressed osteoclasis both in vivo and in vitro. METTL14 promoted TCF1 expression through m6A mRNA methylation, and TCF1 increased the osteogenic activity by elevating the protein level of RUNX2, a key molecule linked to bone formation. In rescue experiments, TCF1 restored the RUNX2 level and osteogenic activity of cells suppressed by METTL14 silencing. In summary, this research demonstrates that METTL14 plays a protective role against OP by promoting the TCF1/RUNX2 axis.







Similar content being viewed by others
Availability of data and materials
The analyzed data sets generated during the study are available from the corresponding author on reasonable request.
Abbreviations
- ANOVA:
-
Analysis of variance
- BMMs:
-
Bone marrow macrophages
- CTSK:
-
Cathepsin K
- CTX-1:
-
Type I collagen cross-linked C-telopeptide
- FBS:
-
Fetal bovine serum
- GAPDH:
-
Glyceraldehyde-3-phosphate dehydrogenase
- HE:
-
Hematoxylin and eosin
- IgG:
-
Immunoglobulin G
- IHC:
-
Immunohistochemistry
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- METTL14:
-
Methyltransferase like 14
- mRNA:
-
Messenger RNA
- M-CSF:
-
Macrophage colony stimulating factor
- m6A:
-
N6-methyladenosine
- NC:
-
Negative control
- NFATC1:
-
Nuclear factor of activated T cells
- LEF:
-
Lymphoid enhancer factor
- Oe:
-
Overexpression
- OP:
-
Osteoporosis
- OPG:
-
Osteoprotegerin
- OVX:
-
Ovariectomy
- PFA:
-
Paraformaldehyde
- RIP:
-
RNA immunoprecipitation
- RT-qPCR:
-
Reverse transcription-quantitative polymerase chain reaction
- RUNX2:
-
Runt-related transcriptional factor 2
- shRNA:
-
Short hairpin RNA
- TCF1:
-
T cell factor 1
- TRAP:
-
Triiodothyronine receptor auxiliary protein
References
Briot K, Roux C, Thomas T, Blain H, Buchon D, Chapurlat R, Debiais F, Feron JM, Gauvain JB, Guggenbuhl P, Legrand E, Lehr-Drylewicz AM, Lespessailles E, Tremollieres F, Weryha G, Cortet B. 2018 update of French recommendations on the management of postmenopausal osteoporosis. Joint Bone Spine. 2018;85(5):519–30. https://doi.org/10.1016/j.jbspin.2018.02.009.
Anastasilakis AD, Polyzos SA, Makras P. therapy of endocrine disease: denosumab vs bisphosphonates for the treatment of postmenopausal osteoporosis. Eur J Endocrinol. 2018;179(1):R31–45. https://doi.org/10.1530/EJE-18-0056.
Jackson RD, Mysiw WJ. Insights into the epidemiology of postmenopausal osteoporosis: the women’s health initiative. Semin Reprod Med. 2014;32(6):454–62. https://doi.org/10.1055/s-0034-1384629.
Bijelic R, Milicevic S, Balaban J. Risk factors for osteoporosis in postmenopausal women. Med Arch. 2017;71(1):25–8. https://doi.org/10.5455/medarh.2017.71.25-28.
Lorentzon M, Cummings SR. Osteoporosis: the evolution of a diagnosis. J Intern Med. 2015;277(6):650–61. https://doi.org/10.1111/joim.12369.
Maeda SS, Lazaretti-Castro M. An overview on the treatment of postmenopausal osteoporosis. Arq Bras Endocrinol Metabol. 2014;58(2):162–71. https://doi.org/10.1590/0004-2730000003039.
Harvey N, Dennison E, Cooper C. Osteoporosis: impact on health and economics. Nat Rev Rheumatol. 2010;6(2):99–105. https://doi.org/10.1038/nrrheum.2009.260.
Si L, Winzenberg TM, Chen M, Jiang Q, Neil A, Palmer AJ. Screening for osteoporosis in Chinese post-menopausal women: a health economic modelling study. Osteoporos Int. 2016;27(7):2259–69. https://doi.org/10.1007/s00198-016-3502-1.
Parker MT, Knop K, Sherwood AV, Schurch NJ, Mackinnon K, Gould PD, Hall AJ, Barton GJ, Simpson GG. Nanopore direct RNA sequencing maps the complexity of arabidopsis mRNA processing and m(6)A modification. Elife. 2020. https://doi.org/10.7554/eLife.49658.
Wang X, Lu Z, Gomez A, Hon GC, Yue Y, Han D, Fu Y, Parisien M, Dai Q, Jia G, Ren B, Pan T, He C. N6-methyladenosine-dependent regulation of messenger RNA stability. Nature. 2014;505(7481):117–20. https://doi.org/10.1038/nature12730.
Ping XL, Sun BF, Wang L, Xiao W, Yang X, Wang WJ, Adhikari S, Shi Y, Lv Y, Chen YS, Zhao X, Li A, Yang Y, Dahal U, Lou XM, Liu X, Huang J, Yuan WP, Zhu XF, Cheng T, Zhao YL, Wang X, Rendtlew Danielsen JM, Liu F, Yang YG. Mammalian WTAP is a regulatory subunit of the RNA N6-methyladenosine methyltransferase. Cell Res. 2014;24(2):177–89. https://doi.org/10.1038/cr.2014.3.
Wang P, Doxtader KA, Nam Y. Structural basis for cooperative function of mettl3 and mettl14 methyltransferases. Mol Cell. 2016;63(2):306–17. https://doi.org/10.1016/j.molcel.2016.05.041.
Yang Y, Hsu PJ, Chen YS, Yang YG. Dynamic transcriptomic m(6)A decoration: writers, erasers, readers and functions in RNA metabolism. Cell Res. 2018;28(6):616–24. https://doi.org/10.1038/s41422-018-0040-8.
Jia G, Fu Y, Zhao X, Dai Q, Zheng G, Yang Y, Yi C, Lindahl T, Pan T, Yang YG, He C. N6-methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nat Chem Biol. 2011;7(12):885–7. https://doi.org/10.1038/nchembio.687.
Zheng G, Dahl JA, Niu Y, Fedorcsak P, Huang CM, Li CJ, Vagbo CB, Shi Y, Wang WL, Song SH, Lu Z, Bosmans RP, Dai Q, Hao YJ, Yang X, Zhao WM, Tong WM, Wang XJ, Bogdan F, Furu K, Fu Y, Jia G, Zhao X, Liu J, Krokan HE, Klungland A, Yang YG, He C. ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility. Mol Cell. 2013;49(1):18–29. https://doi.org/10.1016/j.molcel.2012.10.015.
Qin Y, Li L, Luo E, Hou J, Yan G, Wang D, Qiao Y, Tang C. Role of m6A RNA methylation in cardiovascular disease (Review). Int J Mol Med. 2020;46(6):1958–72. https://doi.org/10.3892/ijmm.2020.4746.
Zhou J, Wan J, Gao X, Zhang X, Jaffrey SR, Qian SB. Dynamic m(6)A mRNA methylation directs translational control of heat shock response. Nature. 2015;526(7574):591–4. https://doi.org/10.1038/nature15377.
Hu Y, Zhao X. Role of m6A in osteoporosis, arthritis and osteosarcoma (Review). Exp Ther Med. 2021;22(3):926. https://doi.org/10.3892/etm.2021.10358.
Peng J, Zhan Y, Zong Y. METTL3-mediated LINC00657 promotes osteogenic differentiation of mesenchymal stem cells via miR-144-3p/BMPR1B axis. Cell Tissue Res. 2022;388(2):301–12. https://doi.org/10.1007/s00441-022-03588-y.
Wu T, Tang H, Yang J, Yao Z, Bai L, Xie Y, Li Q, Xiao J. METTL3-m(6) A methylase regulates the osteogenic potential of bone marrow mesenchymal stem cells in osteoporotic rats via the Wnt signalling pathway. Cell Prolif. 2022;55(5): e13234. https://doi.org/10.1111/cpr.13234.
Wu Y, Xie L, Wang M, Xiong Q, Guo Y, Liang Y, Li J, Sheng R, Deng P, Wang Y, Zheng R, Jiang Y, Ye L, Chen Q, Zhou X, Lin S, Yuan Q. Mettl3-mediated m(6)A RNA methylation regulates the fate of bone marrow mesenchymal stem cells and osteoporosis. Nat Commun. 2018;9(1):4772. https://doi.org/10.1038/s41467-018-06898-4.
Lin Y, Shen X, Ke Y, Lan C, Chen X, Liang B, Zhang Y, Yan S. Activation of osteoblast ferroptosis via the METTL3/ASK1-p38 signaling pathway in high glucose and high fat (HGHF)-induced diabetic bone loss. FASEB J. 2022;36(3): e22147. https://doi.org/10.1096/fj.202101610R.
Kim C, Jin J, Weyand CM, Goronzy JJ. The transcription factor TCF1 in T cell differentiation and aging. Int J Mol Sci. 2020. https://doi.org/10.3390/ijms21186497.
Soderholm S, Cantu C. The WNT/beta-catenin dependent transcription: a tissue-specific business. WIREs Mech Dis. 2021;13(3): e1511. https://doi.org/10.1002/wsbm.1511.
Li Z, Xu Z, Duan C, Liu W, Sun J, Han B. Role of TCF/LEF transcription factors in bone development and osteogenesis. Int J Med Sci. 2018;15(12):1415–22. https://doi.org/10.7150/ijms.26741.
Krishnan V, Bryant HU, Macdougald OA. Regulation of bone mass by wnt signaling. J Clin Invest. 2006;116(5):1202–9. https://doi.org/10.1172/JCI28551.
Maeda K, Kobayashi Y, Koide M, Uehara S, Okamoto M, Ishihara A, Kayama T, Saito M, Marumo K. The regulation of bone metabolism and disorders by wnt signaling. Int J Mol Sci. 2019. https://doi.org/10.3390/ijms20225525.
Yao Y, Yang Y, Guo W, Xu L, You M, Zhang YC, Sun Z, Cui X, Yu G, Qi Z, Liu J, Wang F, Liu J, Zhao T, Ye L, Yang YG, Yu S. METTL3-dependent m(6)A modification programs T follicular helper cell differentiation. Nat Commun. 2021;12(1):1333. https://doi.org/10.1038/s41467-021-21594-6.
Tian Y, Ming J. Melatonin inhibits osteoclastogenesis via RANKL/OPG suppression mediated by rev-erbalpha in osteoblasts. J Cell Mol Med. 2022;26(14):4032–47. https://doi.org/10.1111/jcmm.17440.
Burja B, Kuret T, Janko T, Topalovic D, Zivkovic L, Mrak-Poljsak K, Spremo-Potparevic B, Zigon P, Distler O, Cucnik S, Sodin-Semrl S, Lakota K, Frank-Bertoncelj M. Olive leaf extract attenuates inflammatory activation and DNA damage in human arterial endothelial cells. Front Cardiovasc Med. 2019;6:56. https://doi.org/10.3389/fcvm.2019.00056.
Huang M, Xu S, Liu L, Zhang M, Guo J, Yuan Y, Xu J, Chen X, Zou J. m6A Methylation regulates osteoblastic differentiation and bone remodeling. Front Cell Dev Biol. 2021;9: 783322. https://doi.org/10.3389/fcell.2021.783322.
Sun Z, Wang H, Wang Y, Yuan G, Yu X, Jiang H, Wu Q, Yang B, Hu Z, Shi F, Cao X, Zhang S, Guo T, Zhao J. MiR-103-3p targets the m(6) A methyltransferase METTL14 to inhibit osteoblastic bone formation. Aging Cell. 2021;20(2): e13298. https://doi.org/10.1111/acel.13298.
Appelman-Dijkstra NM, Papapoulos SE. modulating bone resorption and bone formation in opposite directions in the treatment of postmenopausal osteoporosis. Drugs. 2015;75(10):1049–58. https://doi.org/10.1007/s40265-015-0417-7.
Zhao YL, Liu YH, Wu RF, Bi Z, Yao YX, Liu Q, Wang YZ, Wang XX. Understanding m(6)A function through uncovering the diversity roles of yth domain-containing proteins. Mol Biotechnol. 2019;61(5):355–64. https://doi.org/10.1007/s12033-018-00149-z.
Chen X, Xu M, Xu X, Zeng K, Liu X, Pan B, Li C, Sun L, Qin J, Xu T, He B, Pan Y, Sun H, Wang S. METTL14-mediated N6-methyladenosine modification of SOX4 mRNA inhibits tumor metastasis in colorectal cancer. Mol Cancer. 2020;19(1):106. https://doi.org/10.1186/s12943-020-01220-7.
Lin Z, Hsu PJ, Xing X, Fang J, Lu Z, Zou Q, Zhang KJ, Zhang X, Zhou Y, Zhang T, Zhang Y, Song W, Jia G, Yang X, He C, Tong MH. Mettl3-/Mettl14-mediated mRNA N(6)-methyladenosine modulates murine spermatogenesis. Cell Res. 2017;27(10):1216–30. https://doi.org/10.1038/cr.2017.117.
Yi D, Wang R, Shi X, Xu L, Yilihamu Y, Sang J. METTL14 promotes the migration and invasion of breast cancer cells by modulating N6methyladenosine and hsamiR146a5p expression. Oncol Rep. 2020;43(5):1375–86. https://doi.org/10.3892/or.2020.7515.
Yoon KJ, Ringeling FR, Vissers C, Jacob F, Pokrass M, Jimenez-Cyrus D, Su Y, Kim NS, Zhu Y, Zheng L, Kim S, Wang X, Dore LC, Jin P, Regot S, Zhuang X, Canzar S, He C, Ming GL, Song H. Temporal control of mammalian cortical neurogenesis by m(6) a methylation. Cell. 2017. https://doi.org/10.1016/j.cell.2017.09.003.
Zhang W, Ke T, Lin J, Liu P, Guan Z, Deng J, Wang D, Zeng H. The role of m6A in osteoporosis and in the differentiation of mesenchymal stem cells into osteoblasts and adipocytes. Curr Stem Cell Res Ther. 2022. https://doi.org/10.2174/1574888X17666220621155341.
Han J, Kong H, Wang X, Zhang XA. Novel insights into the interaction between N6-methyladenosine methylation and noncoding RNAs in musculoskeletal disorders. Cell Prolif. 2022;55(10): e13294. https://doi.org/10.1111/cpr.13294.
Yang C, Dong Z, Ling Z, Chen Y. The crucial mechanism and therapeutic implication of RNA methylation in bone pathophysiology. Ageing Res Rev. 2022;79: 101641. https://doi.org/10.1016/j.arr.2022.101641.
Li Y, Meng L, Zhao B. The roles of N6-methyladenosine methylation in the regulation of bone development, bone remodeling and osteoporosis. Pharmacol Ther. 2022;238: 108174. https://doi.org/10.1016/j.pharmthera.2022.108174.
Huang C, Wang Y. Downregulation of METTL14 improves postmenopausal osteoporosis via IGF2BP1 dependent posttranscriptional silencing of SMAD1. Cell Death Dis. 2022;13(11):919. https://doi.org/10.1038/s41419-022-05362-y.
Hong G, He X, Shen Y, Chen X, Yang F, Yang P, Pang F, Han X, He W, Wei Q. Chrysosplenetin promotes osteoblastogenesis of bone marrow stromal cells via Wnt/beta-catenin pathway and enhances osteogenesis in estrogen deficiency-induced bone loss. Stem Cell Res Ther. 2019;10(1):277. https://doi.org/10.1186/s13287-019-1375-x.
Baron R, Rawadi G. Targeting the Wnt/beta-catenin pathway to regulate bone formation in the adult skeleton. Endocrinology. 2007;148(6):2635–43. https://doi.org/10.1210/en.2007-0270.
Wallmen B, Schrempp M, Hecht A. Intrinsic properties of Tcf1 and Tcf4 splice variants determine cell-type-specific Wnt/beta-catenin target gene expression. Nucleic Acids Res. 2012;40(19):9455–69. https://doi.org/10.1093/nar/gks690.
Gaur T, Lengner CJ, Hovhannisyan H, Bhat RA, Bodine PV, Komm BS, Javed A, van Wijnen AJ, Stein JL, Stein GS, Lian JB. Canonical WNT signaling promotes osteogenesis by directly stimulating Runx2 gene expression. J Biol Chem. 2005;280(39):33132–40. https://doi.org/10.1074/jbc.M500608200.
Hu L, Su P, Yin C, Zhang Y, Li R, Yan K, Chen Z, Li D, Zhang G, Wang L, Miao Z, Qian A, Xian CJ. Microtubule actin crosslinking factor 1 promotes osteoblast differentiation by promoting beta-catenin/TCF1/Runx2 signaling axis. J Cell Physiol. 2018;233(2):1574–84. https://doi.org/10.1002/jcp.26059.
Li D, Tian Y, Yin C, Huai Y, Zhao Y, Su P, Wang X, Pei J, Zhang K, Yang C, Dang K, Jiang S, Miao Z, Li M, Hao Q, Zhang G, Qian A. Silencing of lncRNA AK045490 promotes osteoblast differentiation and bone formation via beta-catenin/TCF1/Runx2 signaling axis. Int J Mol Sci. 2019. https://doi.org/10.3390/ijms20246229.
Funding
This work was supported by Zhejiang Provincial Public Welfare Research Project (NO. LGF21H060004 and NO. LGF22H060025) and Zhejiang Provincial Medical and Health Technology Project (Grant. 2023KY175).
Author information
Authors and Affiliations
Contributions
XPW and CCZ: conception and design of the research, drafting the manuscript and obtaining funding; MQL: acquisition of data and drafting the manuscript; CJH: acquisition of data; WJ: analysis and interpretation of data; ZYB: statistical analysis; LLZ: obtaining funding, revision of manuscript for important intellectual content, and statistical analysis. All authors wrote and reviewed the manuscript.
Corresponding author
Ethics declarations
Conflict of interests
The authors declare no competing interests.
Ethical approval
The sample collection was approved by the Ethics Committee of Hangzhou First People's Hospital (Approval No: 2021170) and adhered to the Declaration of Helsinki. Signed informed consents were obtained from the patients. The usage of animals was ratified by the Animal Ethics Committee of Hangzhou First People’s Hospital (Approval No: 2021175) and the procedures were performed in strict accordance with the Guide for the Care and Use of Laboratory Animals.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, X., Zou, C., Li, M. et al. METTL14 upregulates TCF1 through m6A mRNA methylation to stimulate osteogenic activity in osteoporosis. Human Cell 36, 178–194 (2023). https://doi.org/10.1007/s13577-022-00825-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13577-022-00825-y