Abstract
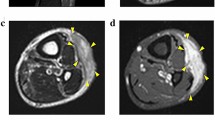
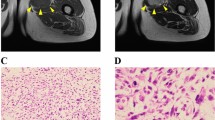
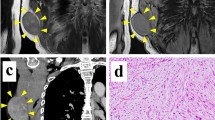
Myxofibrosarcoma (MFS) is an aggressive sarcoma that requires novel therapeutic approaches to improve its clinical outcome. Cell lines are a valuable tool for pre-clinical research; however, there is a lack of patient-derived cell lines of MFS available from public cell banks. This study aimed to develop a patient-derived cell line of MFS. A cell line designated NCC-MFS1-C1 was established from the primary tumor tissue of an 82-year-old male patient with MFS. The short tandem repeat pattern of NCC-MFS1-C1 cells was identical to that of the original tumor, but distinct from that of any other cell lines in public cell banks. NCC-MFS1-C1 cells were maintained as a monolayer culture for over 20 passages in 19 months; the cells exhibited spindle-like morphology, continuous growth, and ability for spheroid formation and invasion. Genomic assay showed that NCC-MFS1-C1 cells had gain and loss of genetic loci. Proteomic profiling revealed that the original tumor and the derived NCC-MFS1-C1 cells had similar, but distinct protein expression patterns. Screening of anti-cancer drugs in NCC-MFS1-C1 cells identified five candidate drugs for MFS. In conclusion, we established a novel MFS cell line, NCC-MFS1-C1, which could be used to study tumor development and effects of anti-cancer drugs.





Similar content being viewed by others
References
Fletcher CDM, Bridge JA, Hogendoorn P, Mertens F. WHO classification of tumours of soft tissue and bone. Fourth edn. Geneva: WHO Press; 2013.
Haglund KE, Raut CP, Nascimento AF, Wang Q, George S, Baldini EH. Recurrence patterns and survival for patients with intermediate- and high-grade myxofibrosarcoma. Int J Radiat Oncol Biol Phys. 2012;82:361–7.
Sanfilippo R, Miceli R, Grosso F, et al. Myxofibrosarcoma: prognostic factors and survival in a series of patients treated at a single institution. Ann Surg Oncol. 2011;18:720–5.
Mentzel T, Calonje E, Wadden C, et al. Myxofibrosarcoma. Clinicopathologic analysis of 75 cases with emphasis on the low-grade variant. Am J Surg Pathol. 1996;20:391–405.
Merck C, Angervall L, Kindblom LG, Oden A. Myxofibrosarcoma. A malignant soft tissue tumor of fibroblastic-histiocytic origin. A clinicopathologic and prognostic study of 110 cases using multivariate analysis. Acta pathologica, microbiologica, et immunologica Scandinavica Supplement. 1983; 282:1–40.
Weiss SW, Enzinger FM. Myxoid variant of malignant fibrous histiocytoma. Cancer. 1977;39:1672–85.
Willems SM, Debiec-Rychter M, Szuhai K, Hogendoorn PC, Sciot R. Local recurrence of myxofibrosarcoma is associated with increase in tumour grade and cytogenetic aberrations, suggesting a multistep tumour progression model. Mod Pathol. 2006;19:407–16.
Lin CN, Chou SC, Li CF, et al. Prognostic factors of myxofibrosarcomas: implications of margin status, tumor necrosis, and mitotic rate on survival. J Surg Oncol. 2006;93:294–303.
Barretina J, Caponigro G, Stransky N, et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature. 2012;483:603–7.
Iorio F, Knijnenburg TA, Vis DJ, et al. A landscape of pharmacogenomic interactions in cancer. Cell. 2016;166:740–54.
Kawashima H, Ogose A, Gu W, et al. Establishment and characterization of a novel myxofibrosarcoma cell line. Cancer Genet Cytogenet. 2005;161:28–35.
Huang HY, Wu WR, Wang YH, et al. ASS1 as a novel tumor suppressor gene in myxofibrosarcomas: aberrant loss via epigenetic DNA methylation confers aggressive phenotypes, negative prognostic impact, and therapeutic relevance. Clin Cancer Res. 2013;19:2861–72.
Lohberger B, Stuendl N, Wolf E, Liegl-Atzwanger B, Leithner A, Rinner B. The novel myxofibrosarcoma cell line MUG-Myx1 expresses a tumourigenic stem-like cell population with high aldehyde dehydrogenase 1 activity. BMC Cancer. 2013;13:563.
Salawu A, Fernando M, Hughes D, et al. Establishment and molecular characterisation of seven novel soft-tissue sarcoma cell lines. Br J Cancer. 2016;115:1058–68.
Lohberger B, Stuendl N, Leithner A, et al. Establishment of a novel cellular model for myxofibrosarcoma heterogeneity. Scientific reports. 2017;7:44700.
Ariizumi T, Ogose A, Kawashima H, Hotta T, Umezu H, Endo N. Multinucleation followed by an acytokinetic cell division in myxofibrosarcoma with giant cell proliferation. J Exp Clin Cancer Res. 2009;28:44.
Willems SM, Mohseny AB, Balog C, et al. Cellular/intramuscular myxoma and grade I myxofibrosarcoma are characterized by distinct genetic alterations and specific composition of their extracellular matrix. J Cell Mol Med. 2009;13:1291–301.
Willems SM, van Remoortere A, van Zeijl R, Deelder AM, McDonnell LA, Hogendoorn PC. Imaging mass spectrometry of myxoid sarcomas identifies proteins and lipids specific to tumour type and grade, and reveals biochemical intratumour heterogeneity. J Pathol. 2010;222:400–9.
Jones EA, van Remoortere A, van Zeijl RJ, et al. Multiple statistical analysis techniques corroborate intratumor heterogeneity in imaging mass spectrometry datasets of myxofibrosarcoma. PLoS One. 2011;6:e24913.
Workgroup ATCCSDO. Cell line misidentification: the beginning of the end. Nat Rev Cancer. 2010;10:441–8.
Capes-Davis A, Reid YA, Kline MC, et al. Match criteria for human cell line authentication: where do we draw the line? Int J Cancer. 2013;132:2510–9.
Capes-Davis A, Dirks W, MacLeod R, Uphoff C. Quality matters: cell lines and their use in research. GIT Lab J Eur. 2014;17:12–3.
Kondo T, Hirohashi S. Application of highly sensitive fluorescent dyes (CyDye DIGE Fluor saturation dyes) to laser microdissection and two-dimensional difference gel electrophoresis (2D-DIGE) for cancer proteomics. Nat Protoc. 2007;1:2940–56.
Wisniewski JR, Zougman A, Nagaraj N, Mann M. Universal sample preparation method for proteome analysis. Nat Methods. 2009;6:359–62.
Baehrecke EH, Dang N, Babaria K, Shneiderman B. Visualization and analysis of microarray and gene ontology data with treemaps. BMC Bioinformatics. 2004;5:84.
Domcke S, Sinha R, Levine DA, Sander C, Schultz N. Evaluating cell lines as tumour models by comparison of genomic profiles. Nat Commun. 2013;4:2126.
Arai K, Sakamoto R, Kubota D, Kondo T. Proteomic approach toward molecular backgrounds of drug resistance of osteosarcoma cells in spheroid culture system. Proteomics. 2013;13:2351–60.
Ben-David U, Ha G, Tseng YY, et al. Patient-derived xenografts undergo mouse-specific tumor evolution. Nat Genet. 2017.
Acknowledgements
We thank Drs. M. Endo, Y. Minami, K. Shimizu, T. Mori, T. Uehara, M. Sugawara, Y. Araki, and Ms. R. Nakano of the Division of Musculoskeletal Oncology, National Cancer Center Hospital for sampling tumor tissue specimens from surgically resected materials. Fundamental Innovative Oncology Core at the National Cancer Center provided support for the SNP array experiment. We would like to thank Editage (http://www.editage.jp) for English language editing and constructive comments on the manuscript.
Funding
This study was funded by the National Cancer Center Research and Development Fund (26-A-9 and 29-A-2).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kito, F., Oyama, R., Sakumoto, M. et al. Establishment and characterization of a novel cell line, NCC-MFS1-C1, derived from a patient with myxofibrosarcoma. Human Cell 32, 214–222 (2019). https://doi.org/10.1007/s13577-018-00233-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13577-018-00233-1