Abstract
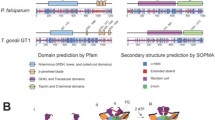
Gyrases are a class of Topoisomerase II that facilitate ATP dependent negative supercoiling of DNA during transcription and replication. They aid in strand breakage during chromosome condensation and take part in separation of intertwined DNA during mitosis. The present study reports gyrase-A (PsGryA) CDS clone from Pisum sativum (L.) comprising of 2817 bp that encodes for a peptide of 939 amino acids. The estimated molecular weight of PsGryA is approximately 104 kDa with pI as 6.99. The protein sequence showed maximum similarity with GyrA of Medicago truncatula. The PsGyrA polypeptide has approximately 42% overall sequence identity with Escherichia coli gyrase-A. The three-dimensional structure of PsGyrA generated based on the X-ray structure of E. coli gyrase-A revealed a putative active site region with amino acid (AA) sequence AAMRYTE located between 208th and 214th AA with a tyrosine residue at position 212. In silico and In vivo immunolocalization studies establishes the dual targeting of PsGyrA to both chloroplast and mitochondria.




Similar content being viewed by others
Abbreviations
- Gyr:
-
Gyrase
- DAPI:
-
4′,6-diamidino-2-phenylindole
- PAGE:
-
Polyacrylamide gel electrophoresis
- CAP:
-
Cysteine-rich secretory proteins, antigen 5, and pathogenesis-related 1 protein
- UTR:
-
Untranslated region
References
Blanchard JL, Lynch M (2000) Organellar genes: Why do they end up in the nucleus? Trends Genet 16:315–320. https://doi.org/10.1016/S0168-9525(00)02053-9
Champoux JJ (2001) DNA topoisomerases: structure, function, and mechanism. Ann Rev Biochem 70:369–413. https://doi.org/10.1146/annurev.biochem.70.1.369
Chen SH, Chan NL, Hsieh TS (2013) New mechanistic and functional insights into DNA topoisomerases. Ann Rev Biochem 82:139–170. https://doi.org/10.1146/annurev-biochem-061809-100002
Cho HS, Lee SS, Kim KD, Hwang I, Lim JS, Park YI, Pai HS (2004) DNA gyrase is involved in chloroplast nucleoid partitioning. Plant Cell 16:2665–2682. https://doi.org/10.1105/tpc.104.024281
Collin F, Karkare S, Maxwell A (2011) Exploiting bacterial DNA gyrase as a drug target: current state and perspectives. Appl Microbiol Biotechnol 92:479–497. https://doi.org/10.1007/s00253-011-3557-z
Emanuelsson O, Nielsen H, von Heijne G (1999) ChloroP, a neural network-based method for predicting chloroplast transit peptides and their cleavage sites. Protein Sci 8:978–984. https://doi.org/10.1110/ps.8.5.978
Evans-Roberts KM, Breuer C, Wall MK, Sugimoto-Shirasu K, Maxwell A (2010) Arabidopsis thaliana GYRB3 does not encode a DNA gyrase subunit. PLoS One 5:e9899. https://doi.org/10.1371/journal.pone.0009899
Evans-Roberts KM, Mitchenall LA, Wall MK, Leroux J, Mylne JS, Maxwell A (2016) DNA gyrase is the target for the quinolone drug ciprofloxacin in Arabidopsis thaliana. J Biol Chem 291:3136–3144. https://doi.org/10.1074/jbc.M115.689554
Gadelle D, Filée J, Buhler C, Forterre P (2003) Phylogenomics of type II DNA topoisomerases. BioEssays 25:232–242. https://doi.org/10.1002/bies.10245
Gould SB, Waller RF, McFadden GI (2008) Plastid evolution. Ann Rev Plant Biol 59:491–517. https://doi.org/10.1146/annurev.arplant.59.032607.092915
Heinhorst S, Cannon G, Weissbach A (1985) Chloroplast DNA synthesis during the cell cycle in cultured cells of Nicotianatabucum:inhibition by nalidixic acid and hydroxyurea. Arch Biochem Biophys 239:475–479. https://doi.org/10.1016/0003-9861(85)90714-3
Hiratsuka J, Shimada H, Whittier R, Ishibashi T, Sakamoto M, Mori M, Kondo C, Honji Y, Sun CR, Meng BY, Li YQ, Kanno A, Nishizawa Y, Hirai A, Shinozaki K, Sugiura M (1989) The complete sequence of the rice (Oryza sativa) chloroplast genome: intermolecular recombination between distinct tRNA genes accounts for a major plastid DNA inversion during the evolution of the cereals. Mol Gen Genet 217:185–194. https://doi.org/10.1007/BF02464880
Itoh R, Takahashi H, Toda K, Kuroiwa H, Kuroiwa T (1997) DNA gyrase involvementin chloroplast-nucleoid division in Cyanidioschyzon merolae. Eur J Cell Biol 73:252–258
Keeling PJ, Archibald JM (2008) Organelle evolution: What’s in a name? Curr Biol 18:345–347. https://doi.org/10.1016/j.cub.2008.02.065
Lam E, Chua NH (1987) Chloroplast DNA gyrase and in vitro regulation of transcription by template topology and novobiocin. Plant Mol Biol 8:415–424. https://doi.org/10.1007/BF00015819
MiIls WR, Reeves M, Fowler DL, Capo SF (1989) DNA synthesis in chloroplasts III: The DNA gyrase inhibitors nalidixic acid andnovobiocin inhibit both thymidine incorporation into DNA and photosynthetic oxygen evolution by isolated chloroplasts. J Exp Bot 40:425–429
Moriyama T, Tajima N, Sekine K, Sato N (2014) Localization and phylogenetic analysis of enzymes related to organellar genome replication in the unicellular rhodophyte Cyanidio schyzon merolae. Genome Biol Evol 6:228–237. https://doi.org/10.1093/gbe/evu009
Pommier Y, Leo E, Zhang H, Marchand C (2010) DNA topoisomerases and their poisoning by anticancer and antibacterial drugs. Chem Biol 17:421–433. https://doi.org/10.1016/j.chembiol.2010.04.012
Pyke KA, Marrlson JL, Leech RM (1989) Evidence for a type II topoisomerase in wheat chloroplasts. FEBS Lett 242:305–308. https://doi.org/10.1016/0014-5793(89)80490-9
Reece RJ, Maxwell A (1991) DNA gyrase: structure and function. Crit Rev Biochemn Mol Biol 26:335–375. https://doi.org/10.3109/10409239109114072
Sato S, Nakamura Y, Kaneko T, Asamizu E, Tabata S (1999) Complete structure of the chloroplast genome of Arabidopsis thaliana. DNA Res 6:283–290. https://doi.org/10.1093/dnares/6.5.283
Shen HB, Chou KC (2007) Signal-3L: a 3-layer approach for predicting signal peptides. Biochem Biophys Res Commun 363:297–303. https://doi.org/10.1016/j.bbrc.2007.08.140
Shen HB, Chou KC (2010) Plant-mPLoc: a top-down strategy to augment the power for predicting plant protein subcellular localization. PLoS One 5:e11335. https://doi.org/10.1371/journal.pone.0011335
Shinozaki K, Ohme M, Tanaka M, Wakasugi T, Hayashida N, Matsubayashi T, Zaita N, Chunwongse J, Obokata J, Yamaguchi-Shinozaki K, Ohto C, Torazawa K, Meng BY, Sugita M, Deno H, Kamogashira T, Yamada K, Kusuda J, Takaiwa F, Kato A, Tohdoh N, Shimada H, Sugiura M (1986) The complete nucleotide sequence of the tobacco chloroplast genome: its gene organization and expression. EMBO J 5:2043–2049
Singh BN, Sopory SK, Reddy MK (2004) Plant DNA topoisomerases: structure, function and cellular roles in plant development. Crit Rev Plant Sci 23:251–269. https://doi.org/10.1080/07352680490452816
Thompson RJ, Mosig G (1985) An ATP-dependent supercoiling topoisomerase of Chlamydomonas reinhardtii affects accumulation of specific chloroplast transcripts. Nucleic Acids Res 13:873–891
Unseld M, Marienfeld JR, Brandt P, Brennicke A (1997) The mitochondrial genome of Arabidopsis thaliana contains 57 genes in 366,924 nucleotides. Nat Genet 15:57–61. https://doi.org/10.1038/ng0197-57
Wall M, Mitchenall L, Maxwell A (2004) Arabidopsis thaliana DNA gyrase is targeted to chloroplasts and mitochondria. Proc Natl Acad Sci USA 101:7821–7826. https://doi.org/10.1073/pnas.0400836101
Wang JC (1996) DNA topoisomerases. Ann Rev Biochem 65:635–692. https://doi.org/10.1146/annurev.bi.65.070196.003223
Woelfle MA, Thompson RJ, Mosig G (1993) Roles of novobiocin-sensitive topoisomerases in chloroplast DNA replication in Chlamydomonas reinhardtii. Nucleic Acids Res 18:4231–4238
Acknowledgements
We acknowledge the research facilities provided by ICGEB, New Delhi for carrying out this piece of work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Reddy, M.K., Achary, V.M.M., Singh, B.N. et al. Molecular characterization of pea DNA gyrase-A reveals dual localization of protein in plastid and mitochondria. J. Plant Biochem. Biotechnol. 28, 291–300 (2019). https://doi.org/10.1007/s13562-018-0478-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13562-018-0478-2