Abstract
Diacylglycerol kinase (DGK) is a kind of phosphokinase that catalyzes the formation of signaling molecule phosphatidic acid. In this study, seven maize (Zea mays) DGK gene family members were identified by an exploration of maize genome via multiple online databases, and designated as ZmDGK1-7, respectively. The proteins encoded by ZmDGKs ranged from 487 to 716 amino acids, and had a molecular weight (MWs) between 54.6 and 80.2 kDa. Phylogenetic analysis revealed that ZmDGKs grouped into three clusters as described for known plant DGK families: Cluster I was composed of three maize DGKs, ZmDGK1, ZmDGK4 and ZmDGK5, cluster II contained ZmDGK6, and the isoforms ZmDGK2, ZmDGK3 and ZmDGK7 fell into cluster III. ZmDGK proteins featured the typical functional domains, while all seven ZmDGKs have a conserved catalytic domain DGKc, only the cluster I ZmDGKs have the DAG/PE binding domain. Most ZmDGK genes showed ubiquitous expression profiles at various developmental stages, while a high relative expression was observed at the tasseling stage. ZmDGK genes exhibited differential expression patterns in response to abiotic stresses including cold, salinity and drought, and all ZmDGK genes were found obviously up-regulated by cold. The distinct roles of ZmDGKs in cold response was also supported by the finding that an accumulation of DGK products–PA under low temperature. This study will help to better understand the roles of DGKs in the development and abiotic stress responses in major crops.




Similar content being viewed by others
Abbreviations
- DAG:
-
Diacylglycerol
- DGK:
-
Diacylglycerol kinase
- PA:
-
Phosphatidic acid
- PC:
-
Phosphatidylcholine
- PE:
-
Phosphatidylethanolamine
- PLD:
-
Phospholipase D
- PLC:
-
Phospholipase C
References
Arisz SA, Testerink C, Munnik T (2009) Plant PA signaling via diacylglycerol kinase. Biochim et Biophys Acta (BBA) Mol Cell Biol Lipids 1791:869–875. doi:10.1016/j.bbalip.2009.04.006
Arisz SA, van Wijk R, Roels W, Zhu JK, Haring MA, Munnik T (2013) Rapid phosphatidic acid accumulation in response to low temperature stress in Arabidopsis is generated through diacylglycerol kinase. Front Plant Sci 4:1. doi:10.3389/fpls.2013.00001
Cacas JL et al (2017) Diacylglycerol kinases activate tobacco NADPH oxidase-dependent oxidative burst in response to cryptogein. Plant Cell Environ 40:585–598. doi:10.1111/pce.12771
Chen J, Zhang W, Song F, Zheng Z (2007) Phospholipase C/diacylglycerol kinase-mediated signalling is required for benzothiadiazole-induced oxidative burst and hypersensitive cell death in rice suspension-cultured cells. Protoplasma 230:13–21. doi:10.1007/s00709-006-0195-x
Colon-Gonzalez F, Kazanietz MG (2006) C1 domains exposed: from diacylglycerol binding to protein-protein interactions. Biochim Biophys Acta 1761:827–837. doi:10.1016/j.bbalip.2006.05.001
Donaldson JG (2009) Phospholipase D in endocytosis and endosomal recycling pathways. Biochim Biophys Acta 1791:845–849. doi:10.1016/j.bbalip.2009.05.011
Escobar-Sepulveda HF, Trejo-Tellez LI, Perez-Rodriguez P, Hidalgo-Contreras JV, Gomez-Merino FC (2017) Diacylglycerol kinases are widespread in higher plants and display inducible gene expression in response to beneficial elements, metal, and metalloid ions. Front Plant Sci 8:129. doi:10.3389/fpls.2017.00129
Ge H, Chen C, Jing W, Zhang Q, Wang H, Wang R, Zhang W (2012) The rice diacylglycerol kinase family: functional analysis using transient RNA interference. Front Plant Sci 3:60. doi:10.3389/fpls.2012.00060
Gomez-Merino FC, Brearley CA, Ornatowska M, Abdel-Haliem ME, Zanor MI, Mueller-Roeber B (2004) AtDGK2, a novel diacylglycerol kinase from Arabidopsis thaliana, phosphorylates 1-stearoyl-2-arachidonoyl-sn-glycerol and 1,2-dioleoyl-sn-glycerol and exhibits cold-inducible gene expression. J Biol Chem 279:8230–8241. doi:10.1074/jbc.M312187200
Hall BG (2013) Building phylogenetic trees from molecular data with MEGA. Mol Biol Evol 30:1229–1235. doi:10.1093/molbev/mst012
Higo K, Ugawa Y, Iwamoto M, Korenaga T (1999) Plant cis-acting regulatory DNA elements (PLACE) database: 1999. Nucleic Acids Res 27:297–300. doi:10.1093/nar/27.1.297
Hirayama T, Ohto C, Mizoguchi T, Shinozaki K (1995) A gene encoding a phosphatidylinositol-specific phospholipase C is induced by dehydration and salt stress in Arabidopsis thaliana. Proc Natl Acad Sci USA 92:3903–3907. doi:10.1073/pnas.92.9.3903
Hong Y et al (2016) Plant phospholipases D and C and their diverse functions in stress responses. Prog Lipid Res 62:55–74. doi:10.1016/j.plipres.2016.01.002
Hou Q, Ufer G, Bartels D (2016) Lipid signalling in plant responses to abiotic stress. Plant, Cell Environ 39:1029–1048. doi:10.1111/pce.12666
Laxalt AM, Munnik T (2002) Phospholipid signalling in plant defence. Curr Opin Plant Biol 5:332–338. doi:10.1016/s1369-5266(02)00268-6
Lee BH, Henderson DA, Zhu JK (2005) The Arabidopsis cold-responsive transcriptome and its regulation by ICE1. Plant Cell 17:3155–3175. doi:10.1105/tpc.105.035568
Lescot M et al (2002) PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res 30:325–327. doi:10.1093/nar/30.1.325
Li W, Li M, Zhang W, Welti R, Wang X (2004) The plasma membrane-bound phospholipase Ddelta enhances freezing tolerance in Arabidopsis thaliana. Nat Biotechnol 22:427–433. doi:10.1038/nbt949
Li M, Hong Y, Wang X (2009) Phospholipase D- and phosphatidic acid-mediated signaling in plants. Biochim Biophys Acta 1791:927–935. doi:10.1016/j.bbalip.2009.02.017
Li Y, Tan Y, Shao Y, Li M, Ma F (2015) Comprehensive genomic analysis and expression profiling of diacylglycerol kinase gene family in Malus prunifolia (Willd.) Borkh. Gene 561:225–234. doi:10.1016/j.gene.2015.02.029
Liu W et al (2015) IBS: an illustrator for the presentation and visualization of biological sequences. Bioinformatics 31:3359–3361. doi:10.1093/bioinformatics/btv362
Meijer HJ, Munnik T (2003) Phospholipid-based signaling in plants. Annu Rev Plant Biol 54:265–306. doi:10.1146/annurev.arplant.54.031902.134748
Mishkind M, Vermeer JE, Darwish E, Munnik T (2009) Heat stress activates phospholipase D and triggers PIP accumulation at the plasma membrane and nucleus. Plant J Cell Mol Biol 60:10–21. doi:10.1111/j.1365-313X.2009.03933.x
Munnik T, Testerink C (2009) Plant phospholipid signaling: “in a nutshell”. J Lipid Res 50(Suppl):S260–265. doi:10.1194/jlr.R800098-JLR200
Munnik T, Vermeer JEM (2010) Osmotic stress-induced phosphoinositide and inositol phosphate signalling in plants. Plant, Cell Environ 33:655–669. doi:10.1111/j.1365-3040.2009.02097.x
Narayanan S, Prasad PV, Welti R (2016) Wheat leaf lipids during heat stress: II. Lipids experiencing coordinated metabolism are detected by analysis of lipid co-occurrence. Plant, Cell Environ 39:608–617. doi:10.1111/pce.12648
Peters C, Kim SC, Devaiah S, Li M, Wang X (2014) Non-specific phospholipase C5 and diacylglycerol promote lateral root development under mild salt stress in Arabidopsis. Plant, Cell Environ 37:2002–2013. doi:10.1111/pce.12334
Raghu P, Manifava M, Coadwell J, Ktistakis NT (2009) Emerging findings from studies of phospholipase D in model organisms (and a short update on phosphatidic acid effectors). Biochim Biophys Acta 1791:889–897. doi:10.1016/j.bbalip.2009.03.013
Ruelland E, Cantrel C, Gawer M, Kader JC, Zachowski A (2002) Activation of phospholipases C and D is an early response to a cold exposure in Arabidopsis suspension cells. Plant Physiol 130:999–1007. doi:10.1104/pp.006080
Saucedo-Garcia M, Gavilanes-Ruiz M, Arce-Cervantes O (2015) Long-chain bases, phosphatidic acid, MAPKs, and reactive oxygen species as nodal signal transducers in stress responses in Arabidopsis. Frontiers in plant science 6:55. doi:10.3389/fpls.2015.00055
Sui Z, Niu L, Yue G, Yang A, Zhang J (2008) Cloning and expression analysis of some genes involved in the phosphoinositide and phospholipid signaling pathways from maize (Zea mays L.). Gene 426:47–56. doi:10.1016/j.gene.2008.09.004
Testerink C, Munnik T (2005) Phosphatidic acid: a multifunctional stress signaling lipid in plants. Trends Plant Sci 10:368–375. doi:10.1016/j.tplants.2005.06.002
Testerink C, Munnik T (2011) Molecular, cellular, and physiological responses to phosphatidic acid formation in plants. J Exp Bot 62:2349–2361. doi:10.1093/jxb/err079
Vaultier MN et al (2008) The hydrophobic segment of Arabidopsis thaliana cluster I diacylglycerol kinases is sufficient to target the proteins to cell membranes. FEBS Lett 582:1743–1748. doi:10.1016/j.febslet.2008.04.042
Wang X (2004) Lipid signaling. Curr Opin Plant Biol 7:329–336. doi:10.1016/j.pbi.2004.03.012
Wang Y et al (2013) PIECE: a database for plant gene structure comparison and evolution. Nucleic Acids Res 41:D1159–1166. doi:10.1093/nar/gks1109
Welti R et al (2002) Profiling membrane lipids in plant stress responses. Role of phospholipase D alpha in freezing-induced lipid changes in Arabidopsis. J Biol Chem 277:31994–32002. doi:10.1074/jbc.M205375200
Xie S, Naslavsky N, Caplan S (2015) Diacylglycerol kinases in membrane trafficking. Cell Logist 5:e1078431. doi:10.1080/21592799.2015.1078431
Xue HW, Chen X, Mei Y (2009) Function and regulation of phospholipid signalling in plants. Biochem J 421:145–156. doi:10.1042/BJ20090300
Yu L et al (2010) Phosphatidic acid mediates salt stress response by regulation of MPK6 in Arabidopsis thaliana. New Phytol 188:762–773. doi:10.1111/j.1469-8137.2010.03422.x
Yue R et al (2015) Genome-wide identification and expression profiling analysis of ZmPIN, ZmPILS, ZmLAX and ZmABCB auxin transporter gene families in maize (Zea mays L.) under various abiotic stresses. PLoS ONE 10:e0118751. doi:10.1371/journal.pone.0118751
Zhang W, Qin C, Zhao J, Wang X (2004) Phospholipase D alpha 1-derived phosphatidic acid interacts with ABI1 phosphatase 2C and regulates abscisic acid signaling. Proc Natl Acad Sci USA 101:9508–9513. doi:10.1073/pnas.0402112101
Acknowledgement
This work was supported by National science and technology support plan (2015BAD23B05-04); Natural Science Foundation of Heilongjiang Province(C201446); National science and technology support plan of china (2013BAD07B01); Heilongjiang Bayi Agricultural University graduate student innovation fund projects (YJSCX2015-Z01).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing financial interests.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Fig. S1
Gene structural analysis of maize DGK genes. (a) Phylogenetic clusters. (b) Exon-intron architecture of ZmDGKs. Genome sequences of ZmDGK genes were analyzed, and a diagram of the exon-intron distribution was generated by the online PIECE program. The blue boxes represent exons, and the gray lines represent the introns. The number 0, 1, 2 indicate the phase of the introns. (TIFF 327 kb)
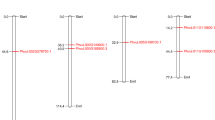
Fig. S2
Chromosomal localization of maize DGK genes. All non-redundant maize DGK genes were mapped on the 10 maize chromosomes and a sketch map was created with MapDraw. (TIFF 916 kb)
Fig. S3
Multiple sequence alignment and conserved domain analysis of maize and Arabidopsis DGKs. (a) DGKc: Diacylglycerol kinase catalytic domain in maize and Arabidopsis. (b) DAG/PE-binding domain in maize, rice, apple and Arabidopsis. The consensus GXGXXG sequence in ATP-binding site in DGKc domain was boxed. (The accession numbers of all the plant DGKs are listed in Table S1). (TIFF 2569 kb)
Fig. S4
Multiple sequence alignment and conserved domain analysis of maize and Arabidopsis DGKs. (a) DGKa: Diacylglycerol kinase accessory domain in maize and Arabidopsis. (b) DAG/PE-binding domain c in maize, rice, apple and Arabidopsis. Multiple sequences alignments were conducted using DNAMAN software (The accession numbers of all the DGKs are listed in Table S1). (TIFF 2948 kb)
Fig. S5
Analysis of Cis- elements in the promoter regions of ZmDGKs. The Scale bar -1500 to -1 represents the upstream region of the promoter of ZmDGKs. The regulatory elements analyzed are LTR (low temperature responsive), C-repeat/DRE (cold/dehydration responsive), ABRE (abscisic acid responsive), MBS (MYB binding site), and TC-rich repeat (defense/ stress responsive). (TIFF 288 kb)
Fig. S6
Expression analysis of ZmDGKs genes in different tissues/developmental stages. Quantitive real-time PCR was conducted with the specific primers designed according to individual ZmDGK gene sequence, and the expression profile of maize DGKs was detected in various maize tissues/developmental stages, including L1: Seedling Stage, L2: Elongation Stage, L3: Huge Bellbottom Period, L4: Tasseling Stage, ST: Stem, ES: Endosperm, SE: Seed. The maize 18s-rRNA gene was used as an endogenous control, and values from roots were normalized to 0 as experimental control. Mean values were obtained from 3 replicates. Vertical bars indicate standard deviation. Lowercase letter on top of the error bars indicate the significant difference (α=0.05) compared with DGK genes. (TIFF 270 kb)
Table S1
Accession numbers of all the DGK genes in this study. (PDF 191 kb)
Table S2
Primers used for quantitive real-time PCR. (PDF 145 kb)
Table S3
Accession numbers of genes in PLD and PLC/DGK pathway. (PDF 219 kb)
Table S4
Sequences producing significant alignments (AtDGK1). (PDF 222 kb)
Table S5
Sequences producing significant alignments (AtDGK2). (PDF 229 kb)
Table S6
Sequences producing significant alignments (AtDGK3). (PDF 228 kb)
Table S7
Sequences producing significant alignments (AtDGK4). (PDF 225 kb)
Table S8
Sequences producing significant alignments (AtDGK5). (PDF 209 kb)
Table S9
Sequences producing significant alignments (AtDGK6). (PDF 202 kb)
Table S10
Sequences producing significant alignments (AtDGK7). (PDF 246 kb)
Table S11
List of PF00130 found in phytozome and NCBI databases. (PDF 142 kb)
Table S12
List of PF00781 found in phytozome database. (PDF 153 kb)
Table S13
List of PF00609 found in phytozome database. (PDF 144 kb)
Rights and permissions
About this article
Cite this article
Gu, Y., Zhao, C., He, L. et al. Genome-wide identification and abiotic stress responses of DGK gene family in maize. J. Plant Biochem. Biotechnol. 27, 156–166 (2018). https://doi.org/10.1007/s13562-017-0424-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13562-017-0424-8