Abstract
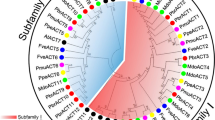
Although plant actins have highly conserved sequences, they are classified as vegetative or reproductive actin based on genetic relationship and the role of the tissue. To characterize the 1,590 bp actin (ACT) gene cloned from the petals of the Ascocenda Princess Mikasa ‘Blue’ orchid, the encoded protein sequence of Ascocenda ACT was aligned to 31 related proteins from orchids and other plant species. The comparison of the amino acids of the actin proteins showed that Ascocenda ACT1, Cymbidium ACT, Phalaenopsis ACT1, Phalaenopsis ACT2, and Phalaenopsis ACT4 contained 13 positions different in amino acid residues in actin polypeptides. Consequently, Ascocenda ACT, Cymbidium ACT, and Phalaenopsis ACT2 were classified as vegetative actins. The classification of Ascocenda ACT was consistent with the results of qualtitative real-time polymerase chain reaction (qPCR) analysis, which demonstrated that actin is highly expressed in all vegetative tissues of the Ascocenda orchid but not in pollen. The predicted three-dimensional structure of the orchid actins revealed that most of the 13 variable residues were located outside the dynamic functional loops, with the exception of Thr16 and Pro353, which were located in the S-loop and target binding site of Ascocenda ACT, respectively. We suggest that the DNA sequence in actin may play a role in the functions of each actin isoform in a tissue-specific manner.





Similar content being viewed by others
Abbreviations
- ACT:
-
Actin
- D-loop:
-
DNase-I binding loop
- G-actin:
-
Globular actin
- H-loop:
-
Histidine-loop
- S-loop:
-
Serine-loop
- W-loop:
-
Tryptophan-loop
References
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucl Acids Res 25:3389–3402
Altschul SF, Wootton JC, Gertz EM, Agarwala R, Morgulis A, Schaffer AA, Yu YK (2005) Protein database searches using compositionally adjusted substitution matrices. FEBS J 272:5101–5109
An YQ, Huang S, McDowell JM, McKinney EC, Meagher RB (1996) Conserved expression of the Arabidopsis ACT1 and ACT3 actin subclass in organ primordia and mature pollen. Plant Cell 8:15–30
An SS, Mopps B, Weber K, Bhattacharya D (1999) The origin and evolution of green algal and plant actins. Mol Biol Evol 16:275–285
Artimo P, Jonnalagedda M, Arnold K, Baratin D, Csardi G, de Castro E, Duvaud S, Flegel V, Fortier A, Gasteiger E, Grosdidier A, Hernandez C, Ioannidis V, Kuznetsov D, Liechti R, Moretti S, Mostaguir K, Redaschi N, Rossier G, Xenarios I, Stockinger H (2012) ExPASy: SIB bioinformatics resource portal. Nucl Acids Res 40:W597–W603
Aspenstrom P, Lindberg U, Karlsson R (1992) Site-specific amino-terminal mutants of yeast-expressed beta-actin. Characterization of the interaction with myosin and tropomyosin. FEBS Lett 303:59–63
Benkert P, Biasini M, Schwede T (2011) Toward the estimation of the absolute quality of individual protein structure models. Bioinformatics 27:343–350
Betts MJ, Russell RB (2003) Amino acid properties and consequences of subsitutions. In: Barnes MR, Gray IC (eds) Bioinformatics for Geneticists. Wiley, United Kingdom
Bhattacharya D, Aubry J, Twait EC, Jurk S (2000) Actin gene duplication and the evolution of morphorlogical complexity in land plants. J Phycol 36:813–820
Bordoli L, Schwede T (2012) Automated protein structure modeling with SWISS-MODEL workspace and the protein model portal. Methods Mol Biol 857:107–136
Chik JK, Lindberg U, Schutt CE (1996) The structure of an open state of β-actin at 2.65 Å resolution. J Mol Biol 263:607–623
de Almeida Engler J, Rodiuc N, Smertenko A, Abad P (2010) Plant actin cytoskeleton re-modeling by plant parasitic nematodes. Plant Signaling Behav 5:213–217
Di Giammartino DC, Nishida K, Manley JL (2011) Mechanisms and consequences of alternative polyadenylation. Mol Cell 43:853–866
Dominguez R, Holmes KC (2011) Actin structure and function. Annu Rev Biophys 40:169–186
Dos Remedios CG, Chhabra D, Kekic M, Dedova IV, Tsubakihara M, Berry DA, Nosworthy NJ (2003) Actin binding proteins: regulation of cytoskeletal microfilaments. Physiol Rev 83:433–473
Edgar RC (2004) MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucl Acids Res 32:1792–1797
Fedorov AA, Lappalainen P, Fedorov EV, Drubin DG, Almo SC (1997) Structure determination of yeast cofilin. Nat Struct Biol 4:366–369
Hennessey ES, Harrison A, Drummond DR, Sparrow JC (1992) Mutant actin: a dead end? J Muscle Res Cell Motil 13:127–131
Hennessey ES, Drummond DR, Sparrow JC (1993) Molecular genetics of actin function. Biochem J 282:657–671
Hu J, Lutz CS, Wilusz J, Tian B (2005) Bioinformatic identification of candidate cis-regulatory elements involved in human mRNA polyadenylation. RNA 11:1485–1493
Huang S, An YQ, McDowell JM, McKinney EC, Meagher RB (1996) The Arabidopsis thaliana ACT4/ACT12 actin gene subclass is strongly expressed throughout pollen development. Plant J 10:189–202
Huang S, An YQ, McDowell JM, McKinney EC, Meagher RB (1997) The Arabidopsis ACT11 actin gene is strongly expressed in tissues of the emerging inflorescence, pollen, and developing ovules. Plant Mol Biol 33:125–139
Hussey PJ, Ketelaar T, Deeks MJ (2006) Control of the actin cytoskeleton in plant cell growth. Annu Rev Plant Biol 57:109–125
Ji G, Zheng J, Shen Y, Wu X, Jiang R, Lin Y, Loke JC, Davis KM, Reese GJ, Li QQ (2007) Predictive modeling of plant messenger RNA polyadenylation sites. BMC Bioinformatics 8:43
Johannes FJ, Gallwitz D (1991) Site-directed mutagenesis of the yeast actin gene: a test for actin function in vivo. EMBO J 10:3951–3958
Jones DT, Taylor WR, Thornton JM (1992) The rapid generation of mutation data matrices from protein sequences. CABIOS 8:275–282
Kabsch W, Mannherz HG, Suck D, Pai EF, Holmes KC (1990) Atomic structure of the actin:DNase I complex. Nature 347:37–44
Kandasamy MK, McKinney EC, Meagher RB (1999) The late pollen-specific actins in angiosperms. Plant J 18:681–691
Kandasamy MK, McKinney EC, Meagher RB (2009) A single vegetative actin isovariant overexpressed under the control of multiple regulatory sequences is sufficient for normal Arabidopsis development. Plant Cell 21:701–718
Kandasamy MK, McKinney EC, Roy E, Meagher RB (2012) Plant vegetative and animal cytoplasmic actins share functional competence for spatial development with protists. Plant Cell 24:2041–2057
Kisselev AF, Songyang Z, Goldberg AL (2000) Why does threonine, and not serine, function as the active site nucleophile in proteasomes? J Biol Chem 275:14831–14837
Kudryashov DS, Grintsevich EE, Rubenstein PA, Reisler E (2010) A nucleotide state-sensing region on actin. J Biol Chem 285:25591–25601
Li XB, Fan XP, Wang XL, Cai L, Yang WC (2005) The cotton ACTIN1 gene is functionally expressed in fibers and participates in fiber elongation. Plant Cell 17:859–875
Lievens S, Goormachtig S, Holsters M (1997) Identification of differentially expressed mRNAs using the differential display technique, workshop on genome diversity and genome expression in plants EMBO course. Ghent, Belgium, pp 1–17
Liu F, Marquardt S, Lister C, Swiezewski S, Dean C (2010) Targeted 3’ processing of antisense transcripts triggers Arabidopsis FLC chromatin silencing. Science 327:94–97
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔC T Method. Methods 25:402–408
Marchler-Bauer A, Lu S, Anderson JB, Chitsaz F, Derbyshire MK, DeWeese-Scott C, Fong JH, Geer LY, Geer RC, Gonzales NR, Gwadz M, Hurwitz DI, Jackson JD, Ke Z, Lanczycki CJ, Lu F, Marchler GH, Mullokandov M, Omelchenko MV, Robertson CL, Song JS, Thanki N, Yamashita RA, Zhang D, Zhang N, Zheng C, Bryant SH (2011) CDD: a Conserved Domain Database for the functional annotation of proteins. Nucl Acids Res 39:D225–D229
McDowell JM, An YQ, Huang S, McKinney EC, Meagher RB (1996a) The Arabidopsis ACT7 actin gene is expressed in rapidly developing tissues and responds to several external stimuli. Plant Physiol 111:699–711
McDowell JM, Huang S, McKinney EC, An YQ, Meagher RB (1996b) Structure and evolution of the actin gene family in Arabidopsis thaliana. Genetics 142:587–602
McKinney EC, Meagher RB (1998) Members of the Arabidopsis actin gene family are widely dispersed in the genome. Genetics 149:663–675
Meagher RB, McKinney EC, Vitale AV (1999) The evolution of new structures: clues from plant cytoskeletal genes. Trends Genet 15:278–284
Miralles F, Visa N (2006) Actin in transcription and transcription regulation. Curr Opin Cell Biol 18:261–266
Mounier N, Arrigo AP (2002) Actin cytoskeleton and small heat shock proteins: How do they interact. Cell Stress Chaperones 7:167–176
Nair UB, Joel PB, Wan Q, Lowey S, Rould MA, Trybus KM (2008) Crystal structures of monomeric actin bound to cytochalasin D. J Mol Biol 384:848–864
Novick P, Botstein D (1985) Phenotypic analysis of temperature-sensitive yeast actin mutants. Cell 40:405–416
Otterbein LR, Graceffa P, Dominguez R (2001) The crystal structure of uncomplexed actin in the ADP state. Science 293:708–711
Oztug Durer ZA, Diraviyam K, Sept D, Kudryashov DS, Reisler E (2010) F-actin structure destabilization and DNase I binding loop: Fluctuations mutational cross-linking and electron microscopy analysis of loop states and effects on F-actin. J Mol Biol 395:544–557
Pearlman SM, Serber Z, Ferrell JE Jr (2011) A mechanism for the evolution of phosphorylation sites. Cell 147:934–946
Reece KS, McElroy D, Wu R (1990) Genomic nucleotide sequence of four rice (Oryza sativa) actin genes. Plant Mol Biol 14:621–624
Reedy M, Beall C, Fyrberg E (1991) Do variant residues among the six actin isoforms of Drosophila reflect functional specialization? Biophys J. 59:187(abstract).
Reisler E (1993) Actin molecular structure and function. Curr Opin Cell Biol 5:41–47
Rothnie HM, Reid J, Hohn T (1994) The contribution of AAUAAA and the upstream element UUUGUA to the efficiency of mRNA 3’-end formation in plants. EMBO J 13:2200–2210
Shah DM, Hightower RC, Meagher RB (1982) Complete nucleotide sequence of a soybean actin gene. Proc Natl Acad Sci U S A 79:1022–1026
Shen Y, Ji G, Haas BJ, Wu X, Zheng J, Reese GJ, Li QQ (2008) Genome level analysis of rice mRNA 3’-end processing signals and alternative polyadenylation. Nucl Acids Res 36:3150–3161
Sheterline P, Clayton J, Sparrow J (1995) Actin. Protein Profile 2:1–103
Slajcherova K, Fiserova J, Fischer L, Schwarzerova K (2012) Multiple actin isotypes in plants: diverse genes for diverse roles. Front Plant Sci. doi:10.3389/fpls.2012.00226
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Vorobiev S, Strokopytov B, Drubin DG, Frieden C, Ono S, Condeelis J, Rubenstein PA, Almo SC (2003) The structure of nonvertebrate actin: Implications for the ATP hydrolytic mechanism. Proc Natl Acad Sci U S A 100:5760–5765
Wertman KF, Drubin DG, Botstein D (1992) Systematic mutational analysis of the yeast ACT1 gene. Genetics 132:337–350
Zheng X, Diraviyam K, Sept D (2007) Nucleotide effects on the structure and dynamics of actin. Biophys J 93:1277–1283
Acknowledgments
This project was supported by the Science Achievement Scholarship of Thailand (SAST) contract no.02/2551, Chulabhorn Research Institute (CRI) and Kasetsart University Research and Development Institute (KURDI), Thailand.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Fig. 1
(PPTX 286 kb)
Supplementary Table 1
(DOCX 37 kb)
Supplementary Table 2
(DOCX 546 kb)
Supplementary Table 3
(DOCX 12 kb)
Rights and permissions
About this article
Cite this article
Pangjai, W., Huehne, P.S. Variable amino acid sequences in the S-loop and target binding site of vegetative actin in flowers of the Ascocenda orchid. J. Plant Biochem. Biotechnol. 24, 408–416 (2015). https://doi.org/10.1007/s13562-014-0291-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13562-014-0291-5