Abstract
Purpose
This study aims to analyze the effect of acrylamide (AC) on hepatic cells of fertilized eggs. The subclasses of glutathione S-transferases and histological variations in serum and liver samples were analyzed in AC-induced chick embryos.
Results
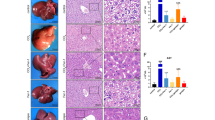
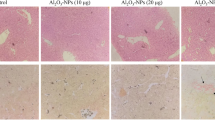
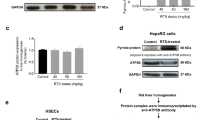
Our results showed that the levels of alanine aminotransferase, serum aspartate aminotransferase, and alkaline phosphatase were significantly (p < 0.05) elevated in the liver of AC-treated chick embryo groups compared to control. In addition, AC-treated groups exhibited a significant (p < 0.05) decreased levels of glutathione content (GSH), glutathione reductase, GSTs, and increased malondialdehyde (MDA levels) in hepatic tissue of chick embryos. Furthermore, liver GSTs binding to various substrates such as CDNB, EPNP, pNPA, BSP, pNBC, ∆5-androstene-3, 17-dione, and EA was significantly (p < 0.05) decreased in AC-treated chick embryo groups compared to control. Histopathology examination (H&E staining) of the AC-treated chick embryo liver showed liver damage to a larger extent, specifically cytoplasmic vacuolation, pyknotic nuclei and blood vein congestion with patchy degeneration were observed. Further, the obtained in vitro results of α-GST, μ-GST, and π-GST were subjected to rigid receptor docking and quantum-polarized-ligand docking, which predicted that the in silico binding affinities of these classes of GSTs are in good correlation with strong hydrogen and van der Waal interactions to AC with −32.760, −26.254, −33.594 and −36.287, −33.980 and −36.363 kcal/mol MMGBSA (ΔGbind) scores respectively. The long-range molecular dynamics (50 ns) simulations have shown more consistency by AC.
Conclusion
The in vivo and in silico findings conclude that AC resulted in the dysfunction of the liver by altering its biochemical markers, α-GST, μ-GST, and π-GST, as well as by histological changes, and thus, AC is suggested as a more harmful agent in developing chick embryo.
Graphical abstract









Similar content being viewed by others
Availability of data and material
All data and materials are available upon request.
Abbreviations
- AC:
-
Acrylamide
- CYP2E1:
-
Cellular enzyme cytochrome P450 2E1
- GST:
-
Glutathione-S-transferase
- GSH:
-
Glutathione
- AST:
-
Aspartate aminotransferase
- ALT:
-
Alanine aminotransferase
- ALP:
-
Alkaline phosphatase
- RRD:
-
Rigid receptor docking
- QPLD:
-
Quantum-polarized-ligand docking
- GR:
-
Glutathione reductase
- EPNP:
-
1,2-Epoxy-3-(p-nitro phenoxy)-propane
- pNPA:
-
P-nitrophenyl acetate
- BSP:
-
Bromosulfophthalein
- pNBC:
-
P-nitrobenzol chloride
- EA:
-
Ethacrynic acid
- CHP:
-
Cumene hydroperoxide
- MDS:
-
Molecular dynamic simulations
- RMSD:
-
Root-mean-square deviation
- PE:
-
Potential of the system
- RMSF:
-
Root-mean-square fluctuations
- rGyr:
-
Radius of gyration
References
Shipp A, Lawrence G, Gentry R, McDonald T, Bartow H, Bounds J, Macdonald N, Clewell H, Allen B, VanLandingham C (2006) Acrylamide: review of toxicity data and dose-response analyses for cancer and noncancer effects. Crit Rev Toxicol 36(6–7):481–608. https://doi.org/10.1080/10408440600851377
El-Mehi AE, El-Sherif NM (2015) Influence of acrylamide on the gastric mucosa of adult albino rats and the possible protective role of rosemary. Tissue Cell 47(3):273–83.https://doi.org/10.1016/j.tice.2015.03.005.
Elblehi SS, El Euony OI, El-Sayed YS (2020) Apoptosis and astrogliosis perturbations and expression of regulatory inflammatory factors and neurotransmitters in acrylamide-induced neurotoxicity under ω3 fatty acids protection in rats. Neurotoxicology 76:44–57. https://doi.org/10.1016/j.neuro.2019.10.004
Hamdy SM, El-Khayat Z, Farrag AR, Sayed ON, El-Sayed MM, Massoud D (2022) Hepatoprotective effect of Raspberry ketone and white tea against acrylamide-induced toxicity in rats. Drug ChemToxicol 45(2):722–730.https://doi.org/10.1080/01480545.2020.1772279.
Zhang L, Zhang H, Miao Y, Wu S, Ye H, Yuan Y (2012) Protective effect of allicin against acrylamide-induced hepatocyte damage in vitro and in vivo. Food ChemToxicol 50(9):3306–12.https://doi.org/10.1016/j.fct.2012.05.060.
Uthra C, Shrivastava S, Jaswal A, Sinha N, Reshi MS, Shukla S (2017) Therapeutic potential of quercetin against acrylamide induced toxicity in rats. Biomed Pharmacother 86:705–714.https://doi.org/10.1016/j.biopha.2016.12.065.
Zamani E, Shokrzadeh M, Ziar A, Abedian-Kenari S, Shaki F (2018) Acrylamide attenuated immune tissues’ function via induction of apoptosis and oxidative stress: Protection by l-carnitine. Hum ExpToxicol 37(8):859–869. https://doi.org/10.1177/0960327117741753
Acaroz U, Ince S, Arslan-Acaroz D, Gurler Z, Kucukkurt I, Demirel HH, Arslan HO, Varol N, Zhu K (2018) The ameliorative effects of boron against acrylamide-induced oxidative stress, inflammatory response, and metabolic changes in rats. Food ChemToxicol 118:745–752. https://doi.org/10.1016/j.fct.2018.06.029
Sun J, Li M, Zou F, Bai S, Jiang X, Tian L, Ou S, Jiao R, Bai W (2018) Protection of cyanidin-3-O-glucoside against acrylamide- and glycidamide-induced reproductive toxicity in leydig cells. Food ChemToxicol 119:268–274. https://doi.org/10.1016/j.fct.2018.03.027
Matoso V, Bargi-Souza P, Ivanski F, Romano MA, Romano RM (2019) Acrylamide: A review about its toxic effects in the light of developmental origin of health and disease (DOHaD) concept. Food Chem 283:422–430. https://doi.org/10.1016/j.foodchem.2019.01.054.
Lee KS, Buck M, Houglum K, Chojkier M (1995) Activation of hepatic stellate cells by TGF alpha and collagen type I is mediated by oxidative stress through c-myb expression. J Clin Invest 96(5):2461–2468. https://doi.org/10.1172/JCI118304
Fennell TR, Sumner SC, Snyder RW, Burgess J, Friedman MA (2006) Kinetics of elimination of urinary metabolites of acrylamide in humans. ToxicolSci 93(2):256–267. https://doi.org/10.1093/toxsci/kfl069
Jiang L, Cao J, An Y, Geng C, Qu S, Jiang L, Zhong L (2007) Genotoxicity of acrylamide in human hepatoma G2 (HepG2) cells. ToxicolIn Vitro 21(8):1486–92. https://doi.org/10.1016/j.tiv.2007.06.011.
Dai Y, Rashba-Step J, Cederbaum AI (1993) Stable expression of human cytochrome P4502E1 in HepG2 cells: characterization of catalytic activities and production of reactive oxygen intermediates. Biochemistry 32(27):6928–6937. https://doi.org/10.1021/bi00078a017
Doroshyenko O, Fuhr U, Kunz D, Frank D, Kinzig M, Jetter A, Reith Y, Lazar A, Taubert D, Kirchheiner J, Baum M, Eisenbrand G, Berger FI, Bertow D, Berkessel A, SörgelF SE, Tomalik-Scharte D (2009) In vivo role of cytochrome P450 2E1 and glutathione-S-transferase activity for acrylamide toxicokinetics in humans. Cancer Epidemiol Biomarkers Prev 18(2):433–443. https://doi.org/10.1158/1055-9965.EPI-08-0832
Hayes JD, Pulford DJ (1995) The glutathione S-transferase supergene family: regulation of GST and the contribution of the isoenzymes to cancer chemoprotection and drug resistance. Crit Rev BiochemMolBiol 30(6):445–600. https://doi.org/10.3109/10409239509083491
Shi C, Wang X, Xiao Z, Wang R, Qiao Y, Kan G (2020) Cloning, characterization and expression analysis of glutathione S-transferase from the Antarctic yeast Rhodotorulamucilaginosa AN5. Protein ExprPurif 167:105518. https://doi.org/10.1016/j.pep.2019.105518.
Lee BY, Choi BS, Kim MS, Park JC, Jeong CB, Han J, Lee JS (2019) The genome of the freshwater water flea Daphnia magna: a potential use for freshwater molecular ecotoxicology. AquatToxicol 210:69–84. https://doi.org/10.1016/j.aquatox.2019.02.009
Flanagan JU, Smythe ML (2011) Sigma-class glutathione transferases. Drug Metab Rev 43(2):194–214. https://doi.org/10.3109/03602532.2011.560157
Howie AF, Forrester LM, Glancey MJ, Schlager JJ, Powis G, Beckett GJ, Hayes JD, Wolf CR (1990) Glutathione S-transferase and glutathione peroxidase expression in normal and tumour human tissues. Carcinogenesis 11(3):451–458. https://doi.org/10.1093/carcin/11.3.451
Thyagaraju K, Hemavathi B, Vasundhara K, Rao AD, Devi KN (2005) Comparative study on glutathione transferases of rat brain and testis under the stress of phenobarbitol and beta-methylcholanthrene. J Zhejiang UnivSci B 6(8):759–769. https://doi.org/10.1631/jzus.2005.B0759
Listowsky I, Rowe JD, Patskovsky YV, Tchaikovskaya T, Shintani N, Novikova E, Nieves E (1998) Human testicular glutathione S-transferases: insights into tissue-specific expression of the diverse subunit classes. ChemBiol Interact 24(111–112):103–112. https://doi.org/10.1016/s0009-2797(97)00154-3
Shang W, Liu WH, Zhao XH, Sun QJ, Bi JZ, Chi ZF (2008) Expressions of glutathione S-transferase alpha, mu, and pi in brains of medically intractable epileptic patients. BMC Neurosci 18(9):67. https://doi.org/10.1186/1471-2202-9-67
Hayes JD, Mantle TJ (1986) Inhibition of hepatic and extrahepatic glutathione S-transferases by primary and secondary bile acids. Biochem J 15;233(2):407–15. https://doi.org/10.1042/bj2330407
Dasari S, Ganjayi MS, Yellanurkonda P, Basha S, Meriga B (2018) Role of glutathione S-transferases in detoxification of a polycyclic aromatic hydrocarbon, methylcholanthrene. ChemBiol Interact 294:81–90. https://doi.org/10.1016/j.cbi.2018.08.023
Barbosa VB, Costa-Silva PF, Araújo-Silva H, Miranda G, Correia-Pinto J, Moura RS (2022) Molecular insights of Hippo signaling in the chick developing lung. Biochim Biophys Acta Gene Regul Mech 1866(1):194904. https://doi.org/10.1016/j.bbagrm.2022.194904
Song R, Shurson GC (2013) Evaluation of lipid peroxidation level in corn dried distillers grains with solubles. J Anim Sci 91(9):4383–4388. https://doi.org/10.2527/jas.2013-6319
Tekin S, Seven E (2022) Assessment of serum catalase, reduced glutathione, and superoxide dismutase activities and malondialdehyde levels in keratoconus patients. Eye (Lond) 36(10):2062–2066. https://doi.org/10.1038/s41433-021-01753-1
Mannervik B, Danielson UH (1988) Glutathione transferases–structure and catalytic activity. CRC Crit Rev Biochem 23(3):283–337. https://doi.org/10.3109/10409238809088226
Pasala C, Katari SK, Nalamolu RM, Bitla AR, Amineni U (2019) In silico probing exercises, bioactive-conformational and dynamic simulations strategies for designing and promoting selective therapeutics against Helicobacter pylori strains. J Mol Graph Model 92:167–179. https://doi.org/10.1016/j.jmgm.2019.07.015
Pasala C, Chilamakuri CSR, Katari SK, Nalamolu RM, Bitla AR, Amineni U (2019b) Epitope-driven common subunit vaccine design against H. pylori strains. J Biomol Struct Dyn 37(14):3740–3750. https://doi.org/10.1080/07391102.2018.1526714.
Pasala C, Katari SK, Nalamolu RM, Bitla AR, Amineni U (2019) Hierarchical-clustering, scaffold-mining exercises and dynamics simulations for effectual inhibitors against lipid-a biosynthesis of Helicobacter pylori. Cell MolBioeng 12(3):255–274. https://doi.org/10.1007/s12195-019-00572-5
Friesner RA, Murphy RB, Repasky MP, Frye LL, Greenwood JR, Halgren TA, Sanschagrin PC, Mainz DT (2006) Extra precision glide: docking and scoring incorporating a model of hydrophobic enclosure for protein-ligand complexes. J Med Chem 49(21):6177–6196. https://doi.org/10.1021/jm051256o
Nagalakshmamma V, Venkataswamy M, Pasala C, Umamaheswari A, Thyagaraju K, Nagaraju C, Chalapathi PV (2020) Design, synthesis, anti-tobacco mosaic viral and molecule docking simulations of urea/thiourea derivatives of 2-(piperazine-1-yl)-pyrimidine and 1-(4-Fluoro/4-Chloro phenyl)-piperazine and 1-(4-Chloro phenyl)-piperazine—a study. Bioorg Chem 102:104084. https://doi.org/10.1016/j.bioorg.2020.104084.
Elfimova N, Schlattjan M, Sowa JP, Dienes HP, Canbay A, Odenthal M (2012) Circulating microRNAs: promising candidates serving as novel biomarkers of acute hepatitis. Front Physiol 3:476. https://doi.org/10.3389/fphys.2012.00476
Honig LS, Rosenberg RN (2000) Apoptosis and neurologic disease. Am J Med 108(4):317–330. https://doi.org/10.1016/s0002-9343(00)00291-6
Lopachin RM, Decaprio AP (2005) Protein adduct formation as a molecular mechanism in neurotoxicity. ToxicolSci 86(2):214–225. https://doi.org/10.1093/toxsci/kfi197
Ahmadabad MA, Naeimi A, Keymoradzadeh A, Faghani S, Ahmadabad MA, Boroujeni NA, Mohammadpour H, Saberi A. Evaluation of De Ritis (AST/ALT), ALP/ALT, and AST/ALP ratios as prognostic factors in patients with acute ischemic stroke. BMC Neurol 22(1):450. https://doi.org/10.1186/s12883-022-02989-4
Dasari S, Gonuguntla S, Yellanurkonda P, Nagarajan P, Meriga B (2019) Sensitivity of glutathione S-transferases to high doses of acrylamide in albino wistar rats: affinity purification, biochemical characterization and expression analysis. Ecotoxicol Environ Saf 182:109416. https://doi.org/10.1016/j.ecoenv.2019.109416.
Acknowledgements
The authors are thankful to DST-FIST and DST-PURSE, S. V. University, Tirupati for providing facilities.The authors are grateful to the University Grants Commission (UGC), Government of India, New Delhi, for providing a fellowship to carry out the research (File No. F/PDFSS-2015-17-AND-10953)
Funding
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors. Also, no sponsorships in any way have been existed.
Author information
Authors and Affiliations
Contributions
VM contributed to methodology, investigation, data curation, writing-original draft, visualization. JBP and BM contributed to investigation, methodology, data curation. NV and REM contributed to investigation, formal analysis. CP and UMA contributed to formal analysis, investigation, methodology. TK contributed to conceptualization; methodology; supervision; review and editing.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Mallepogu, V., Mallapu, R.E., Vadabingi, N. et al. Acrylamide toxicity inhibits chick embryo hepatic alpha, mu, and pi-glutathione S-transferases: molecular dynamic simulations, substrate specificity, and docking. Toxicol. Environ. Health Sci. 15, 289–302 (2023). https://doi.org/10.1007/s13530-023-00183-8
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13530-023-00183-8