Abstract
Purpose
The recent focus on the roles of N-linked glycoproteins in carcinogenesis across various malignancies has prompted our exploration of aberrantly expressed glycoproteins responsible for HCC progression and potential therapeutic strategy.
Methods
Mass spectrometry was applied to initially identify abnormally expressed glycoproteins in HCC, which was further assessed by immunohistochemistry (IHC) staining. The role of selected glycoprotein on HCC development and underlying mechanism was systematically investigated by colony formation, mouse xenograft, RNA-sequencing and western blot assays, etc. Chromatin immunoprecipitation (ChIP) and luciferase assays were performed to explore potential transcription factors (TFs) of selected glycoprotein. The regulation of repaglinide (RPG) on expression of lumican and downstream effectors was assessed by western blot and IHC, while its impact on malignant phenotypes of HCC was explored through in vitro and in vivo analyses, including a murine NASH-HCC model established using western diet and carbon tetrachloride (CCl4).
Results
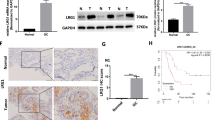
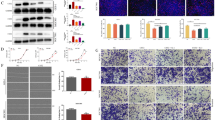
Lumican exhibited upregulation in both serum and tumor tissue, with elevated expression associated with an inferior prognosis in HCC patients. Knockdown of lumican resulted in significantly reduced growth of HCC in vitro and in vivo. Mechanically, lumican promoted HCC malignant phenotypes by inhibiting the p53/p21 signaling pathway. Forkhead Box O3 (FOXO3) was identified as the TF of lumican that transcriptionally enhanced its expression. Without silencing FOXO3, RPG blocked the binding of FOXO3 to the promoter region of lumican, thereby inhibiting the activation of lumican/p53/p21 axis. Mice treated with RPG developed fewer and smaller HCCs than those in the control group at 24 weeks after establishment.
Conclusion
Our results indicate that RPG prevented the development and progression of HCC via alteration of FOXO3/lumican/p53 axis.







Similar content being viewed by others
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- AFP:
-
Alpha-fetoprotein
- ALT:
-
Alanine aminotransferase
- ALB:
-
Albumin
- APS:
-
Ammonium persulfate
- AST:
-
Aspartate aminotransferase
- AUC:
-
Area under curve
- BCLC:
-
Barcelona Clinic Liver Cancer
- BS:
-
Binding site
- CF:
-
Core-fucosylation
- ChIP:
-
Chromatin immunoprecipitation
- CCl4 :
-
Carbon tetrachloride
- DDA:
-
Data dependent acquisition
- DMSO:
-
Dimethylsulfoxide
- EMT:
-
Epithelial-mesenchymal transition
- FOXO3:
-
Forkhead Box O3
- HCC:
-
Hepatocellular carcinoma
- IC50:
-
Half maximal inhibitory concentration
- IHC:
-
Immunohistochemistry
- MVI:
-
Microvascular invasion
- PBS:
-
Phosphate buffer saline
- PCR:
-
Polymerase chain reaction
- PRM:
-
Parallel Reaction Monitoring
- PVTT:
-
Portal vein tumor thrombosis
- ROC:
-
Receiver operating characteristic curve
- shRNA:
-
Short hairpin RNA
- siRNA:
-
Small interfering RNA
- SLRP:
-
Small leucine-rich proteoglycan
- STAT3:
-
Signal transducer and activator of transcription 3
- TBL:
-
Total bilirubin
- TCGA:
-
The Cancer Genome Atlas
- TF:
-
Transcription Factor
- WD:
-
Western diet
References
H. Sung, J. Ferlay, R.L. Siegel, M. Laversanne, I. Soerjomataram, A. Jemal, F. Bray, CA Cancer J. Clin. 71, 209–249 (2021). https://doi.org/10.3322/caac.21660
Q. Zhang, Y. Lou, J. Yang, J. Wang, J. Feng, Y. Zhao, L. Wang, X. Huang, Q. Fu, M. Ye, X. Zhang, Y. Chen, C. Ma, H. Ge, J. Wang, J. Wu, T. Wei, Q. Chen, J. Wu, C. Yu, Y. Xiao, X. Feng, G. Guo, T. Liang, X. Bai, Gut 68, 2019–2031 (2019). https://doi.org/10.1136/gutjnl-2019-318912
S.S. Pinho, C.A. Reis, Nat. Rev. Cancer 15, 540–555 (2015). https://doi.org/10.1038/nrc3982
C. Liao, J. An, S. Yi, Z. Tan, H. Wang, H. Li, X. Guan, J. Liu, Q. Wang, J. Cancer 12, 4109–4120 (2021). https://doi.org/10.7150/jca.58268
G. Antonarelli, V. Pieri, F.M. Porta, N. Fusco, G. Finocchiaro, G. Curigliano, C. Criscitiello, Cells 12 (2023). https://doi.org/10.3390/cells12060840
Y. Zhou, T. Fukuda, Q. Hang, S. Hou, T. Isaji, A. Kameyama, J. Gu, Sci. Rep. 7, 11563 (2017). https://doi.org/10.1038/s41598-017-11911-9
F. Li, S. Zhao, Y. Cui, T. Guo, J. Qiang, Q. Xie, W. Yu, W. Guo, W. Deng, C. Gu, T. Wu, Am. J. Cancer Res. 10, 816–837 (2020)
P. Agrawal, B. Fontanals-Cirera, E. Sokolova, S. Jacob, C.A. Vaiana, D. Argibay, V. Davalos, M. McDermott, S. Nayak, F. Darvishian, M. Castillo, B. Ueberheide, I. Osman, D. Fenyö, L.K. Mahal, E. Hernando, Cancer Cell 31, 804–819 e807 (2017)
G.S.M. Kammeijer, J. Nouta, J. de la Rosette, T.M. de Reijke, M. Wuhrer, Anal. Chem. 90, 4414–4421 (2018). https://doi.org/10.1021/acs.analchem.7b04281
H. Yin, Z. Lin, S. Nie, J. Wu, Z. Tan, J. Zhu, J. Dai, Z. Feng, J. Marrero, D.M. Lubman, J. Proteome Res. 13, 2887–2896 (2014)
H. Yin, Z. Tan, J. Wu, J. Zhu, K.A. Shedden, J. Marrero, D.M. Lubman, J. Proteome Res. 14, 4876–4884 (2015). https://doi.org/10.1021/acs.jproteome.5b00718
S. Shang, A. Plymoth, S. Ge, Z. Feng, H.R. Rosen, S. Sangrajrang, P. Hainaut, J.A. Marrero, L. Beretta, Hepatology 55, 483–490 (2012). https://doi.org/10.1002/hep.24703
A.D. Nardo, N.G. Grun, M. Zeyda, M. Dumanic, G. Oberhuber, E. Rivelles, T.H. Helbich, D.F. Markgraf, M. Roden, T. Claudel, M. Trauner, T.M. Stulnig, Liver Int. 40, 1620–1633 (2020). https://doi.org/10.1111/liv.14464
X. Gao, Y. Sheng, J. Yang, C. Wang, R. Zhang, Y. Zhu, Z. Zhang, K. Zhang, S. Yan, H. Sun, J. Wei, X. Wang, X. Yu, Y. Zhang, Q. Luo, Y. Zheng, P. Qiao, Y. Zhao, Q. Dong, L. Qin, J. Exp. Clin. Cancer Res. 37, 179 (2018)
K. Stavenhagen, H. Hinneburg, M. Thaysen-Andersen, L. Hartmann, D. Varón Silva, J. Fuchser, S. Kaspar, E. Rapp, P.H. Seeberger, D. Kolarich, J. Mass Spectrom. 48, 627–639 (2013). https://doi.org/10.1002/jms.3210
L. Cao, N. Tolić, Y. Qu, D. Meng, R. Zhao, Q. Zhang, R.J. Moore, E.M. Zink, M.S. Lipton, L. Paša-Tolić, S. Wu, Anal. Biochem. 452, 96–102 (2014)
Y. Tan, J. Zhu, C.D. Gutierrez Reyes, Y. Lin, Z. Tan, Z. Wu, J. Zhang, A. Cano, S. Verschleisser, Y. Mechref, A.G. Singal, N.D. Parikh, D.M. Lubman, ACS Omega 8, 12467–12480 (2023). https://doi.org/10.1021/acsomega.3c00519
L. Yang, W.L. Deng, B.G. Zhao, Y. Xu, X.W. Wang, Y. Fang, H.J. Xiao, Cancer Gene Ther. 29, 326–340 (2022). https://doi.org/10.1038/s41417-021-00312-w
W. Chen, J. Jiang, L. Gong, Z. Shu, D. Xiang, X. Zhang, K. Bi, H. Diao, J. Exp. Clin. Cancer Res. 40(1) (2021). https://doi.org/10.1186/s13046-020-01803-8
Y. Cheng, P. Zhan, J. Lu, Y. Lu, C. Luo, X. Cen, F. Wang, C. Xie, Z. Yin, Liver Int. 43, 1577–1592 (2023). https://doi.org/10.1111/liv.15611
M. Ballmann, D. Hubert, B.M. Assael, D. Staab, A. Hebestreit, L. Naehrlich, T. Nickolay, N. Prinz, R.W. Holl, Lancet Diabetes Endocrinol. 6, 114–121 (2018). https://doi.org/10.1016/s2213-8587(17)30400-x
S. Salcher, G. Spoden, J.M. Huber, G. Golderer, H. Lindner, M.J. Ausserlechner, U. Kiechl-Kohlendorfer, K. Geiger, P. Obexer, Cells 9 (2019). https://doi.org/10.3390/cells9010001
A. Jin, Y. Hong, Y. Yang, H. Xu, X. Huang, X. Gao, X. Gong, Q. Dai, L. Jiang, J. Dent. Res. 101, 196–205 (2022). https://doi.org/10.1177/00220345211021534
Y. Zhou, Y. Chen, X. Zhang, Q. Xu, Z. Wu, X. Cao, M. Shao, Y. Shu, T. Lv, C. Lu, M. Xie, T. Wen, J. Yang, Y. Shi, H. Bu, Hepatology 74, 797–815 (2021). https://doi.org/10.1002/hep.31780
T. Tsuchida, Y.A. Lee, N. Fujiwara, M. Ybanez, B. Allen, S. Martins, M.I. Fiel, N. Goossens, H.I. Chou, Y. Hoshida, S.L. Friedman, J. Hepatol. 69, 385–395 (2018)
S. Sribenja, K. Sawanyawisuth, R. Kraiklang, C. Wongkham, K. Vaeteewoottacharn, S. Obchoei, Q. Yao, S. Wongkham, C. Chen, BMC Cancer 13, 430 (2013). https://doi.org/10.1186/1471-2407-13-430
K. Engeland, Cell Death Differ. 29, 946–960 (2022). https://doi.org/10.1038/s41418-022-00988-z
F. Fontana, M. Marzagalli, M. Raimondi, V. Zuco, N. Zaffaroni, P. Limonta, Cell Prolif. 54, e13111 (2021). https://doi.org/10.1111/cpr.13111
E.M. Kim, C.H. Jung, J. Kim, S.G. Hwang, J.K. Park, H.D. Um, Cancer Res. 77, 3092–3100 (2017). https://doi.org/10.1158/0008-5472.Can-16-2098
Z. Ma, Z. Li, S. Wang, Z. Zhou, C. Liu, H. Zhuang, Q. Zhou, S. Huang, C. Zhang, B. Hou, J. Exp. Clin. Cancer Res. 41, 130 (2022). https://doi.org/10.1186/s13046-022-02310-8
E.S. Wong, D. Thybert, B.M. Schmitt, K. Stefflova, D.T. Odom, P. Flicek, Genome Res. 25, 167–178 (2015). https://doi.org/10.1101/gr.177840.114
O. Fornes, J.A. Castro-Mondragon, A. Khan, R. van der Lee, X. Zhang, P.A. Richmond, B.P. Modi, S. Correard, M. Gheorghe, D. Baranasic, W. Santana-Garcia, G. Tan, J. Cheneby, B. Ballester, F. Parcy, A. Sandelin, B. Lenhard, W.W. Wasserman, A. Mathelier, Nucleic Acids Res. 48, D87–D92 (2020). https://doi.org/10.1093/nar/gkz1001
S. Li, X. Wei, J. He, Q. Cao, D. Du, X. Zhan, Y. Zeng, S. Yuan, L. Sun, Cancer Metastasis Rev. 40, 925–948 (2021). https://doi.org/10.1007/s10555-021-09973-3
H. Hu, Y.R. Miao, L.H. Jia, Q.Y. Yu, Q. Zhang, A.Y. Guo, Nucleic Acids Res. 47, D33–D38 (2019)
Y. Wu, H. Du, M. Zhan, H. Wang, P. Chen, D. Du, X. Liu, X. Huang, P. Ma, D. Peng, L. Sun, S. Yuan, J. Ding, L. Lu, J. Jiang, Cell Death Discov. 11, 248 (2020). https://doi.org/10.1038/s41419-020-2471-7
X. Wang, Q. Zhou, Z. Yu, X. Wu, X. Chen, J. Li, C. Li, M. Yan, Z. Zhu, B. Liu, L. Su, Int. J. Cancer 141, 998–1010 (2017). https://doi.org/10.1002/ijc.30801
W. Mao, M. Luo, X. Huang, Q. Wang, J. Fan, L. Gao, Y. Zhang, J. Geng, Transl. Oncol. 12, 1072–1078 (2019). https://doi.org/10.1016/j.tranon.2019.05.014
K. Karamanou, M. Franchi, D. Vynios, S. Brezillon, Semin. Cancer Biol. 62, 125–133 (2020). https://doi.org/10.1016/j.semcancer.2019.08.003
C.T. Yang, J.M. Li, W.K. Chu, S.E. Chow, Cell Death Discov. 9, 414 (2018). https://doi.org/10.1038/s41419-017-0212-3
X. Chen, X. Li, X. Hu, F. Jiang, Y. Shen, R. Xu, L. Wu, P. Wei, X. Shen, Front. Oncol. 10, 605 (2020). https://doi.org/10.3389/fonc.2020.00605
L.R. Pack, L.H. Daigh, M. Chung, T. Meyer, Nat. Commun. 12, 3356 (2021). https://doi.org/10.1038/s41467-021-23612-z
A. Papoutsidakis, E.M. Giatagana, A. Berdiaki, I. Spyridaki, D.A. Spandidos, A. Tsatsakis, G.N. Tzanakakis, D. Nikitovic, Int. J. Oncol. 57, 791–803 (2020). https://doi.org/10.3892/ijo.2020.5094
D. Dhar, L. Antonucci, H. Nakagawa, J.Y. Kim, E. Glitzner, S. Caruso, S. Shalapour, L. Yang, M.A. Valasek, S. Lee, K. Minnich, E. Seki, J. Tuckermann, M. Sibilia, J. Zucman-Rossi, M. Karin, Cancer Cell 33, 1061–1077 e1066 (2018). https://doi.org/10.1016/j.ccell.2018.05.003
M. Lu, D. Hartmann, R. Braren, A. Gupta, B. Wang, Y. Wang, C. Mogler, Z. Cheng, T. Wirth, H. Friess, J. Kleeff, N. Huser, Y. Sunami, BMC Cancer 19, 887 (2019). https://doi.org/10.1186/s12885-019-6110-6
F. Fondevila, P. Fernandez-Palanca, C. Mendez-Blanco, T. Payo-Serafin, E. Lozano, J.J.G. Marin, J. Gonzalez-Gallego, J.L. Mauriz, Cancers (Basel) 13 (2021). https://doi.org/10.3390/cancers13215349
S. de Oliveira, R.A. Houseright, A.L. Graves, N. Golenberg, B.G. Korte, V. Miskolci, A. Huttenlocher, J. Hepatol. 70, 710–721 (2019). https://doi.org/10.1016/j.jhep.2018.11.034
Acknowledgements
We acknowledge Guang Yang and Guangneng Liao for their help with the mice management. We also acknowledge the support provided by LC-Bio company (Hangzhou, China) with the RNA sequencing and related analysis experiments. We also acknowledge the support provided by Fangfang Wang (Institute of hematology, West China Hospital) with flow cytometry assay and related analysis.
Funding
This work was supported by grants from the Natural Science Foundation of China (NO. 82173255, 82270691 and 82273330), the Key R&D projects of Sichuan Provincial Department of Science and Technology (NO. 2022YFS0253, 2023YFS0026).
Author information
Authors and Affiliations
Contributions
Conception and design: Y. T., Y. Z., and Y. S.; Development of methodology: Y. T., Y. Z., W. Z., Z. W., Q. X.; Acquisition of data (provided animals, acquired and managed patients, provided facilities, etc.): Y. T., Y. Z., W. Z., Q. W., Jian Y. and T. L.; Analysis and interpretation of data: Y. T., Y. Z., L. Y. and Y. S.; Writing, review, and/or revision of the manuscript: Y. T., Y. Z., Jia. Y and H. L.; Administrative, technical, or material support: Y. Z., Q. W. and Y. S.; Study supervision: Jia. Y., Y. S. and H. L.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
This study was approved by the ethics committee of the West China Hospital, Sichuan University. The protocols applied in the current study conformed to the ethical guidelines of 1975 Declaration of Helsinki, and written informed consent were provided by all patients.
Consent for publication
All authors have reviewed the final version of the manuscript and approved it for publication.
Competing interests
None of the authors have any competing interests to declare.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tan, Y., Zhou, Y., Zhang, W. et al. Repaglinide restrains HCC development and progression by targeting FOXO3/lumican/p53 axis. Cell Oncol. (2024). https://doi.org/10.1007/s13402-024-00919-9
Accepted:
Published:
DOI: https://doi.org/10.1007/s13402-024-00919-9