Abstract
Purpose
Glucose not only provides energy for tumor cells, but also provides various biomolecules that are essential for their survival, proliferation and invasion. Therefore, it is of great clinical significance to understand the mechanism of how tumor cells adapt to metabolic stress and maintain their survival. The aim of this research was to study the critical role of OGT and TRIM29 O-GlcNAc modification driven adaptability of PDAC cells to low glucose stress, which might have important medical implications for PDAC therapy.
Methods
Western blotting, mass spectrometry and WGA-immunoprecipitation were used to examined the levels of OGT and O-GlcNAc glycosylated proteins in BxPC3 and SW1990 cells in normal culture and under glucose deprivation conditions. Crystal violet assay, flow cytometry, RIP, RT-qPCR, protein stability assay, biotin pull down were used to investigate the mechanism of OGT and TRIM29-mediated adaptive response to glucose deficiency in PDAC cells.
Results
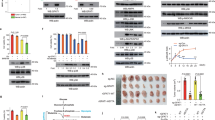
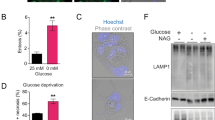
The current study found that under the condition of low glucose culture, the levels of OGT and O-GlcNAc glycosylation in PDAC cells were significantly higher than those in normal culture. Moreover, the high expression of OGT has a protective effect on PDAC cells under low glucose stress. This study confirmed that there was no significant change in mRNA level and protein degradation of OGT under low glucose stress, which was mainly reflected in the increase of protein synthesis. In addition, O-GlcNAc modification at T120 site plays a critical role in the metabolic adaptive responses mediated by TRIM29.
Conclusions
Taken together, our study indicated that O-GlcNAcylation of TRIM29 at T120 site and OGT translation forms a loop feedback to facilitate survival of PDAC under glucose deficiency.








Similar content being viewed by others
Data availability
All data in this study will be available from the corresponding author on reasonable request.
References
E. Hessmann, S.M. Buchholz, I.E. Demir, S.K. Singh, T.M. Gress, V. Ellenrieder, A. Neesse, Physiol. Rev. 100, 1707–1751 (2020). https://doi.org/10.1152/physrev.00042.2019
Y. Qian, Y. Gong, Z. Fan, G. Luo, Q. Huang, S. Deng, H. Cheng, K. Jin, Q. Ni, X. Yu, C. Liu, J. Hematol. Oncol. 13, 130 (2020). https://doi.org/10.1186/s13045-020-00958-3
T. Seufferlein, T.J. Ettrich, Transl. Gastroenterol. Hepatol. 4, 21 (2019). https://doi.org/10.21037/tgh.2019.03.05
Z. Nsingwane, G. Candy, J. Devar, J. Omoshoro-Jones, M. Smith, E. Nweke, Mol. Biol. Rep. 47, 6269–6280 (2020). https://doi.org/10.1007/s11033-020-05648-4
V. Purohit, L. Wang, H. Yang, J. Li, G.M. Ney, E.R. Gumkowski, A.J. Vaidya, A. Wang, A. Bhardwaj, E. Zhao, I. Dolgalev, A. Zamperone, E.V. Abel, M.P.D. Magliano, H.C. Crawford, D. Diolaiti, T.Y. Papagiannakopoulos, C.A. Lyssiotis, D.M. Simeone, Genes Dev. 35, 218–233 (2021). https://doi.org/10.1101/gad.344184.120
M. Ghashghaeinia, M. Koberle, U. Mrowietz, I. Bernhardt, Cell Cycle 18, 1316–1334 (2019). https://doi.org/10.1080/15384101.2019.1618125
Y. Zhao, X. Hu, Y. Liu, S. Dong, Z. Wen, W. He, S. Zhang, Q. Huang, M. Shi, Mol. Cancer 16, 79 (2017). https://doi.org/10.1186/s12943-017-0648-1
N. Hay, Nat. Rev. Cancer 16, 635–649 (2016). https://doi.org/10.1038/nrc.2016.77
M.G. Vander Heiden, L.C. Cantley, C.B. Thompson, Science 324, 1029–1033 (2009). https://doi.org/10.1126/science.1160809
M.G. Vander Heiden, R.J. DeBerardinis, Cell 168, 657–669 (2017). https://doi.org/10.1016/j.cell.2016.12.039
K. Ohshima, E. Morii, Metabolites 11 (2021). https://doi.org/10.3390/metabo11010028
L. Yan, P. Raj, W. Yao, H. Ying, Cancers (Basel) 11 (2019). https://doi.org/10.3390/cancers11101460
N.M. Akella, L. Ciraku, M.J. Reginato, BMC Biol. 17, 52 (2019). https://doi.org/10.1186/s12915-019-0671-3
J.B. Lee, K.H. Pyo, H.R. Kim, Cancers (Basel) 13 (2021). https://doi.org/10.3390/cancers13215365
F. Chiaradonna, F. Ricciardiello, R. Palorini, Cells 7 (2018). https://doi.org/10.3390/cells7060053
E. Ju Kim, Chembiochem 21, 3026–3035 (2020). https://doi.org/10.1002/cbic.202000194
D.T. King, A. Males, G.J. Davies, D.J. Vocadlo, Curr. Opin. Chem. Biol. 53, 131–144 (2019). https://doi.org/10.1016/j.cbpa.2019.09.001
H. Nie, W. Yi, J. Zhejiang Univ. Sci. B 20, 437–448 (2019). https://doi.org/10.1631/jzus.B1900150
C. Slawson, G.W. Hart, Nat. Rev. Cancer 11, 678–684 (2011). https://doi.org/10.1038/nrc3114
Q. Ong, W. Han, X. Yang, Front. Endocrinol. (Lausanne) 9, 599 (2018). https://doi.org/10.3389/fendo.2018.00599
J.C. Chatham, J. Zhang, A.R. Wende, Physiol. Rev. 101, 427–493 (2021). https://doi.org/10.1152/physrev.00043.2019
G.W. Hart, J. Biol. Chem. 294, 2211–2231 (2019). https://doi.org/10.1074/jbc.AW119.003226
X. Li, Z. Wu, J. He, Y. Jin, C. Chu, Y. Cao, F. Gu, H. Wang, C. Hou, X. Liu, Q. Zou, Oncogene 40, 4859–4871 (2021). https://doi.org/10.1038/s41388-021-01901-7
H.G. Seo, H.B. Kim, J.Y. Yoon, T.H. Kweon, Y.S. Park, J. Kang, J. Jung, S. Son, E.C. Yi, T.H. Lee, W.H. Yang, J.W. Cho, Cell Death Dis. 11, 815 (2020). https://doi.org/10.1038/s41419-020-02999-5
J. Xing, L. Weng, B. Yuan, Z. Wang, L. Jia, R. Jin, H. Lu, X.C. Li, Y.J. Liu, Z. Zhang, Nat. Immunol. 17, 1479 (2016). https://doi.org/10.1038/ni1216-1479a
X. Deng, X. Fu, H. Teng, L. Fang, B. Liang, R. Zeng, L. Chen, Y. Zou, J. Transl. Med. 19, 332 (2021). https://doi.org/10.1186/s12967-021-03007-w
R. Huang, Y. Fu, Y. Deng, Biochem. Biophys. Reports 28, 101117 (2021). https://doi.org/10.1016/j.bbrep.2021.101117
Q. Li, L. Lin, Y. Tong, Y. Liu, J. Mou, X. Wang, X. Wang, Y. Gong, Y. Zhao, Y. Liu, B. Zhong, L. Dai, Y.Q. Wei, H. Zhang, H. Hu, Cell Discov. 4, 13 (2018). https://doi.org/10.1038/s41421-018-0010-9
J. Xing, A. Zhang, H. Zhang, J. Wang, X.C. Li, M.S. Zeng, Z. Zhang, Nat. Commun. 8, 945 (2017). https://doi.org/10.1038/s41467-017-00101-w
L. Ai, W.J. Kim, M. Alpay, M. Tang, C.E. Pardo, S. Hatakeyama, W.S. May, M.P. Kladde, C.D. Heldermon, E.M. Siegel, K.D. Brown, Cancer Res. 74, 4875–4887 (2014). https://doi.org/10.1158/0008-5472.CAN-13-3579
Y. Kanno, M. Watanabe, T. Kimura, K. Nonomura, S. Tanaka, S. Hatakeyama, Acta Histochem. 116, 708–712 (2014). https://doi.org/10.1016/j.acthis.2013.12.009
N. Jiang, W.J. Chen, J.W. Zhang, C. Xu, X.C. Zeng, T. Zhang, Y. Li, G.Y. Wang, Oncotarget 6, 7866–7879 (2015). https://doi.org/10.18632/oncotarget.3492
H. Sun, X. Dai, B. Han, Dis. Markers 2014, 317817 (2014). https://doi.org/10.1155/2014/317817
A.D. Santin, F. Zhan, S. Bellone, M. Palmieri, S. Cane, E. Bignotti, S. Anfossi, M. Gokden, D. Dunn, J.J. Roman, T.J. O’Brien, E. Tian, M.J. Cannon, J. Shaughnessy Jr., S. Pecorelli, Int. J. Cancer 112, 14–25 (2004). https://doi.org/10.1002/ijc.20408
Z. Yuan, A. Villagra, L. Peng, D. Coppola, M. Glozak, E.M. Sotomayor, J. Chen, W.S. Lane, E. Seto, Mol. Cell. Biol. 30, 3004–3015 (2010). https://doi.org/10.1128/MCB.01023-09
S. Chandana, H.M. Babiker, D. Mahadevan, Expert Opin. Investig. Drugs 28, 161–177 (2019). https://doi.org/10.1080/13543784.2019.1557145
R. Buono, V.D. Longo, Trends Endocrinol. Metab. 29, 271–280 (2018). https://doi.org/10.1016/j.tem.2018.01.008
J.M. Wang, B.Q. Liu, Z.X. Du, C. Li, J. Sun, J. Yan, J.Y. Jiang, H.Q. Wang, J. Cell Mol. Med. 24, 562–572 (2020). https://doi.org/10.1111/jcmm.14764
C. Lam, J.Y. Low, P.T. Tran, H. Wang, Cancer Lett. 503, 11–18 (2021). https://doi.org/10.1016/j.canlet.2021.01.010
X. Yang, K. Qian, Nat. Rev. Mol. Cell. Biol. 18, 452–465 (2017). https://doi.org/10.1038/nrm.2017.22
K. Qian, S. Wang, M. Fu, J. Zhou, J.P. Singh, M.D. Li, Y. Yang, K. Zhang, J. Wu, Y. Nie, H.B. Ruan, X. Yang, J. Biol. Chem. 293, 13989–14000 (2018). https://doi.org/10.1074/jbc.RA118.004709
A.K. Olson, B. Bouchard, W.Z. Zhu, J.C. Chatham, C. Des Rosiers, J. Biol. Chem. 295, 2018–2033 (2020). https://doi.org/10.1074/jbc.RA119.010565
C. Jia, H. Li, D. Fu, Y. Lan, BioMed. Res. Int. 2020, 1921609 (2020). https://doi.org/10.1155/2020/1921609
M. Aquino-Gil, A. Pierce, Y. Perez-Cervera, E. Zenteno, T. Lefebvre, Biochem. Soc. Trans. 45, 365–370 (2017). https://doi.org/10.1042/BST20160404
N.S. Sharma, V.K. Gupta, P. Dauer, K. Kesh, R. Hadad, B. Giri, A. Chandra, V. Dudeja, C. Slawson, S. Banerjee, S.M. Vickers, A. Saluja, S. Banerjee, Theranostics 9, 3410–3424 (2019). https://doi.org/10.7150/thno.32615
N. Sonenberg, A.G. Hinnebusch, Mol. Cell 28, 721–729 (2007). https://doi.org/10.1016/j.molcel.2007.11.018
W.C. Merrick, G.D. Pavitt, Cold Spring Harb. Perspect. Biol. 10 (2018). https://doi.org/10.1101/cshperspect.a033092
A. Fukao, T. Tomohiro, T. Fujiwara, Cells 10 (2021). https://doi.org/10.3390/cells10071711
H. Ramos, A. Monette, M. Niu, A. Barrera, B. Lopez-Ulloa, Y. Fuentes, P. Guizar, K. Pino, L. DesGroseillers, A.J. Mouland, M. Lopez-Lastra, Nucleic Acids Res. 50, 411–429 (2022). https://doi.org/10.1093/nar/gkab1188
S. Li, J. Shao, G. Lou, C. Wu, Y. Liu, M. Zheng, J. Exp. Clin. Cancer Res. 40, 53 (2021). https://doi.org/10.1186/s13046-021-01853-6
S.A. Haizel, U. Bhardwaj, G. R.l. Jr., S. Mitra, D.J. Goss, J. Biol. Chem. 295, 11693–11706 (2020). https://doi.org/10.1074/jbc.RA120.013678
V.V. Smirnova, E.D. Shestakova, D.V. Bikmetov, A.A. Chugunova, I.A. Osterman, M.V. Serebryakova, O.V. Sergeeva, T.S. Zatsepin, I.N. Shatsky, I.M. Terenin, RNA 25, 757–767 (2019). https://doi.org/10.1261/rna.065623.118
G. Hernandez, J.L. Ramirez, A. Pedroza-Torres, L.A. Herrera, M.A. Jimenez-Rios, Front. Genet. 10, 14 (2019). https://doi.org/10.3389/fgene.2019.00014
V.V. Smirnova, E.D. Shestakova, D.S. Nogina, P.A. Mishchenko, T.A. Prikazchikova, T.S. Zatsepin, I.V. Kulakovskiy, I.N. Shatsky, I.M. Terenin, Nucleic Acids Res. 50, 1111–1127 (2022). https://doi.org/10.1093/nar/gkab1286
L. Marash, N. Liberman, S. Henis-Korenblit, G. Sivan, E. Reem, O. Elroy-Stein, A. Kimchi, Mol. Cell 30, 447–459 (2008). https://doi.org/10.1016/j.molcel.2008.03.018
C. Sanchez-Jimenez, J.M. Izquierdo, Cell Cycle 14, 2033–2043 (2015). https://doi.org/10.1080/15384101.2015.1053668
I. Carrascoso, B.R. Velasco, J.M. Izquierdo, Int. J. Mol. Sci. 22 (2021). https://doi.org/10.3390/ijms222312775
T. Kawai, A. Lal, X. Yang, S. Galban, K. Mazan-Mamczarz, M. Gorospe, Mol. Cell. Biol. 26, 3295–3307 (2006). https://doi.org/10.1128/MCB.26.8.3295-3307.2006
S. Yamasaki, G. Stoecklin, N. Kedersha, M. Simarro, P. Anderson, J. Biol. Chem. 282, 30070–30077 (2007). https://doi.org/10.1074/jbc.M706273200
R. Reyes, J. Alcalde, J.M. Izquierdo, Genome Biol. 10, R87 (2009). https://doi.org/10.1186/gb-2009-10-8-r87
J.M. Izquierdo, N. Majos, S. Bonnal, C. Martinez, R. Castelo, R. Guigo, D. Bilbao, J. Valcarcel, Mol. Cell 19, 475–484 (2005). https://doi.org/10.1016/j.molcel.2005.06.015
C. Sanchez-Jimenez, M.D. Ludena, J.M. Izquierdo, Cell Death Dis. 6, e1669 (2015). https://doi.org/10.1038/cddis.2015.43
S. Hofmann, N. Kedersha, P. Anderson, P. Ivanov, Biochim. Biophys. Acta Mol. Cell. Res. 1868, 118876 (2021). https://doi.org/10.1016/j.bbamcr.2020.118876
A. Khong, T. Matheny, S. Jain, S.F. Mitchell, J.R. Wheeler, R. Parker, Mol. Cell 68, 808–820 e805 (2017). https://doi.org/10.1016/j.molcel.2017.10.015
S. Hatakeyama, Expert Opin. Ther. Targets 20, 767–770 (2016). https://doi.org/10.1517/14728222.2016.1148687
L. Hao, J.M. Wang, B.Q. Liu, J. Yan, C. Li, J.Y. Jiang, F.Y. Zhao, H.Y. Qiao, H.Q. Wang, Biochim. Biophys. Acta Mol. Cell. Res. 1868, 118878 (2021). https://doi.org/10.1016/j.bbamcr.2020.118878
T.M. Harris, P. Du, N. Kawachi, T.J. Belbin, Y. Wang, N.F. Schlecht, T.J. Ow, C.E. Keller, G.J. Childs, R.V. Smith, R.H. Angeletti, M.B. Prystowsky, J. Lim, Arch. Pathol. Lab. Med. 139, 494–507 (2015). https://doi.org/10.5858/arpa.2014-0131-OA
W. Lai, J. Zhao, C. Zhang, D. Cui, J. Lin, Y. He, H. Zheng, X. Wu, M. Yang, Dis. Esophagus 26, 817–822 (2013). https://doi.org/10.1111/j.1442-2050.2012.01400.x
L. Hao, Q. Zhang, H.Y. Qiao, F.Y. Zhao, J.Y. Jiang, L.Y. Huyan, B.Q. Liu, J. Yan, C. Li, H.Q. Wang, Mol. Ther. Nucleic Acids 24, 579–590 (2021). https://doi.org/10.1016/j.omtn.2021.01.027
J. Sun, J. Yan, H.Y. Qiao, F.Y. Zhao, C. Li, J.Y. Jiang, B.Q. Liu, X.N. Meng, H.Q. Wang, Oncogene 39, 546–559 (2020). https://doi.org/10.1038/s41388-019-0992-2
Acknowledgements
We thank Key Laboratory of Cell Biology in Ministry of Public Health, and Key Laboratory of Medical Cell Biology in Ministry of Education of China Medical University for providing experimental platform.
Funding
This work was partly supported by National Natural Science Foundation of China (32071258, 82203337); Natural Science Foundation of the Higher Education Institutions of Jiangsu Province, China (22KJB320017); Supporting project of Nanjing University of Chinese Medicine to NSFC (XPT82203337); Doctoral Start-up Foundation of Liaoning Province (2021-BS-093); Doctoral Start-up Foundation of Liaoning Province (2020-BS-101).
Author information
Authors and Affiliations
Contributions
JS and HW designed this study; FZ, XC, JW and YY performed the experiments; XC, JS, JW, CL analyzed the data; HW and FZ wrote the manuscript. All authors have contributed to the final version of the manuscript.
Corresponding authors
Ethics declarations
Consent for publication
Written informed consent for publication was obtained from all participants.
Conflict of interest
There is no conflict of interests in this work.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhao, FY., Chen, X., Wang, JM. et al. O-GlcNAcylation of TRIM29 and OGT translation forms a feedback loop to promote adaptive response of PDAC cells to glucose deficiency. Cell Oncol. (2024). https://doi.org/10.1007/s13402-023-00915-5
Accepted:
Published:
DOI: https://doi.org/10.1007/s13402-023-00915-5