Abstract
Objective and design
Pancreatic cancer is a highly malignant tumor that is well known for its poor prognosis. Based on glycosylation, we performed integrated quantitative N-glycoproteomics to investigate the synergistic anti-tumor effects of aspirin and gemcitabine on pancreatic cancer cells and explore the potential molecular mechanisms of chemotherapy in pancreatic cancer.
Methods and results
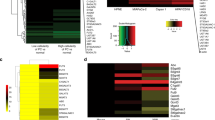
Two pancreatic cancer cell lines (PANC-1 and BxPC-3) were treated with gemcitabine, aspirin, and a combination (gemcitabine + aspirin). We found that the addition of aspirin enhanced the inhibitory effect of gemcitabine on the activity of PANC-1 and BxPC-3 cells. Quantitative N-glycoproteome, proteome, phosphorylation, and transcriptome data were obtained from integrated multi-omics analysis to evaluate the anti-tumor effects of aspirin and gemcitabine on pancreatic cancer cells. Mfuzz analysis of intact N-glycopeptide profiles revealed two consistent trends associated with the addition of aspirin, which showed a strong relationship between N-glycosylation and the synergistic effect of aspirin. Further analysis demonstrated that the dynamic regulation of sialylation and high-mannose glycoforms on ECM-related proteins (LAMP1, LAMP2, ITGA3, etc.) was a significant factor for the ability of aspirin to promote the anti-tumor activity of gemcitabine and the drug resistance of pancreatic cancer cells.
Conclusions
In-depth analysis of N-glycosylation-related processes and pathways in pancreatic cancer cells can provide new insight for future studies regarding pancreatic cancer therapeutic targets and drug resistance mechanisms.





Similar content being viewed by others
Data Availability
The data used and/or analyzed during the present study are available.
Abbreviations
- PDAC:
-
Pancreatic ductal adenocarcinoma
- NSAIDs:
-
Non-steroidal anti-inflammatory drugs
- COX-2:
-
Cyclooxygenase-2
- NF-κB:
-
Nuclear factor kappa-B
- TNF:
-
Tumor necrosis factor
- ECM:
-
Extremely dense extracellular matrix
- EMT:
-
Mesenchymal transition
- TME:
-
Tumor microenvironment
- TMT:
-
Tandem mass tag
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- GO:
-
Gene ontology
- INGPs:
-
Intact N-glycopeptides
- IC50:
-
Half maximal inhibitory concentration
- GTs:
-
Glycosyltransferases
- PTM:
-
Post-translational modification
- PPI:
-
Protein-protein interaction
References
R.L. Siegel, K.D. Miller, Cancer statistics, 2020. CA Cancer J Clin. 70, 7–30 (2020)
T. Kamisawa, L.D. Wood, T. Itoi, K. Takaori, Pancreatic cancer. Lancet. 388, 73–85 (2016)
D.D. Von Hoff, T. Ervin, F.P. Arena, E.G. Chiorean, J. Infante, M. Moore, T. Seay, S.A. Tjulandin, W.W. Ma, M.N. Saleh, M. Harris, M. Reni, S. Dowden, D. Laheru, N. Bahary, R.K. Ramanathan, J. Tabernero, M. Hidalgo, D. Goldstein, E. Van Cutsem, X. Wei, J. Iglesias, M.F. Renschler, Increased survival in pancreatic cancer with nab-paclitaxel plus gemcitabine. N Engl. J. Med. 369, 1691–1703 (2013)
A. Stathis, M.J. Moore, Advanced pancreatic carcinoma: current treatment and future challenges. Nat. Rev. Clin. Oncol. 7, 163–172 (2010)
K.K.F. Tsoi, J.M.W. Ho, F.C.H. Chan, J.J.Y. Sung, Long-term use of low-dose aspirin for cancer prevention: a 10-year population cohort study in Hong Kong. Int. J. Cancer. 145, 267–273 (2019)
Y. Zhang, L. Liu, P. Fan, N. Bauer, J. Gladkich, E. Ryschich, A.V. Bazhin, N.A. Giese, O. Strobel, T. Hackert, U. Hinz, W. Gross, F. Fortunato, I. Herr, Aspirin counteracts cancer stem cell features, desmoplasia and gemcitabine resistance in pancreatic cancer. Oncotarget. 6, 9999–10015 (2015)
K.P. Olive, M.A. Jacobetz, C.J. Davidson, A. Gopinathan, D. McIntyre, D. Honess, B. Madhu, M.A. Goldgraben, M.E. Caldwell, D. Allard, K.K. Frese, G. Denicola, C. Feig, C. Combs, S.P. Winter, H. Ireland-Zecchini, S. Reichelt, W.J. Howat, A. Chang, M. Dhara, L. Wang, F. Rückert, R. Grützmann, C. Pilarsky, K. Izeradjene, S.R. Hingorani, P. Huang, S.E. Davies, W. Plunkett, M. Egorin, R.H. Hruban, N. Whitebread, K. McGovern, J. Adams, C. Iacobuzio-Donahue, J. Griffiths, D.A. Tuveson, Inhibition of hedgehog signaling enhances delivery of chemotherapy in a mouse model of pancreatic cancer. Science 324, 1457–1461 (2009)
M.V. Apte, J.S. Wilson, A. Lugea, S.J. Pandol, A starring role for stellate cells in the pancreatic cancer microenvironment. Gastroenterology 144, 1210–1219 (2013)
C.J. Tape, S. Ling, M. Dimitriadi, K.M. McMahon, J.D. Worboys, H.S. Leong, I.C. Norrie, C.J. Miller, G. Poulogiannis, D.A. Lauffenburger, C. Jørgensen, Oncogenic KRAS regulates Tumor Cell Signaling via Stromal Reciprocation. Cell 165, 910–920 (2016)
C. Feig, A. Gopinathan, A. Neesse, D.S. Chan, N. Cook, D.A. Tuveson, The pancreas cancer microenvironment. Clin. Cancer Res. 18, 4266–4276 (2012)
S. Pan, Y. Tamura, R. Chen, D. May, M.W. McIntosh, T.A. Brentnall, Large-scale quantitative glycoproteomics analysis of site-specific glycosylation occupancy. Mol. Biosyst. 8, 2850–2856 (2012)
H.M. Park, M.P. Hwang, Y.W. Kim, K.J. Kim, J.M. Jin, Y.H. Kim, Y.H. Yang, K.H. Lee, Y.G. Kim, Mass spectrometry-based N-linked glycomic profiling as a means for tracking pancreatic cancer metastasis. Carbohydr. Res. 413, 5–11 (2015)
P. Sharma, S. Hu-Lieskovan, J.A. Wargo, A. Ribas, Primary, adaptive, and Acquired Resistance to Cancer Immunotherapy. Cell 168, 707–723 (2017)
Y. Binenbaum, S. Na’ara, Z. Gil, Gemcitabine resistance in pancreatic ductal adenocarcinoma. Drug Resist. Updat. 23, 55–68 (2015)
P. Saftig, R. Puertollano, How lysosomes sense, integrate, and cope with stress. Trends Biochem. Sci. 46, 97–112 (2021)
S. Sahni, J. Gillson, K.C. Park, S. Chiang, L.Y.W. Leck, P.J. Jansson, D.R. Richardson, NDRG1 suppresses basal and hypoxia-induced autophagy at both the initiation and degradation stages and sensitizes pancreatic cancer cells to lysosomal membrane permeabilization. Biochim. Biophys. Acta Gen. Subj. 1864, 129625 (2020)
R. Raman, M. Venkataraman, S. Ramakrishnan, W. Lang, S. Raguram, R. Sasisekharan, Advancing glycomics: implementation strategies at the consortium for functional glycomics. Glycobiology 16, 82r–90r (2006)
D.F. Quail, J.A. Joyce, Microenvironmental regulation of tumor progression and metastasis. Nat. Med. 19, 1423–1437 (2013)
M.H. Tan, N.J. Nowak, R. Loor, H. Ochi, A.A. Sandberg, C. Lopez, J.W. Pickren, R. Berjian, H.O. Douglass, T.M. Chu, Characterization of a new primary human pancreatic tumor line. Cancer Invest. 4, 15–23 (1986)
E.L. Deer, J. González-Hernández, J.D. Coursen, J.E. Shea, J. Ngatia, C.L. Scaife, M.A. Firpo, S.J. Mulvihill, Phenotype and genotype of pancreatic cancer cell lines. Pancreas 39, 425–435 (2010)
D.A. Drew, Y. Cao, A.T. Chan, Aspirin and colorectal cancer: the promise of precision chemoprevention. Nat. Rev. Cancer. 16, 173–186 (2016)
D.L. Tao, Tassi Yunga, aspirin and antiplatelet treatments in cancer. Blood 137, 3201–3211 (2021)
Y. Zhang, L. Liu, P. Fan, N. Bauer, J. Gladkich, E. Ryschich, A.V. Bazhin, N.A. Giese, O. Strobel, T. Hackert, Aspirin counteracts cancer stem cell features, desmoplasia and gemcitabine resistance in pancreatic cancer. Oncotarget 6, 9999 (2015)
X. Zhu, X. Shen, J. Qu, R.M. Straubinger, W.J. Jusko, Proteomic analysis of combined Gemcitabine and Birinapant in Pancreatic Cancer cells. Front. Pharmacol. 9, 84 (2018)
T. Arumugam, V. Ramachandran, K.F. Fournier, H. Wang, L. Marquis, J.L. Abbruzzese, G.E. Gallick, C.D. Logsdon, D.J. McConkey, W. Choi, Epithelial to mesenchymal transition contributes to drug resistance in pancreatic cancer. Cancer Res. 69, 5820–5828 (2009)
J. Xu, S. Liu, X. Yang, S. Cao, Y. Zhou, Paracrine HGF promotes EMT and mediates the effects of PSC on chemoresistance by activating c-Met/PI3K/Akt signaling in pancreatic cancer in vitro. Life Sci. 263, 118523 (2020)
H. Sumiyoshi, A. Matsushita, Y. Nakamura, Y. Matsuda, T. Ishiwata, Z. Naito, E. Uchida, Suppression of STAT5b in pancreatic cancer cells leads to attenuated gemcitabine chemoresistance, adhesion and invasion. Oncol. Rep. 35, 3216–3226 (2016)
S. Zhao, C. Chen, K. Chang, A. Karnad, J. Jagirdar, A.P. Kumar, J.W. Freeman, CD44 expression level and isoform contributes to pancreatic Cancer cell plasticity, invasiveness, and response to TherapyCD44 and Gemcitabine Resistance. Clin. Cancer Res. 22, 5592–5604 (2016)
C. Yang, M. Cao, Y. Liu, Y. He, Y. Du, G. Zhang, F. Gao, Inducible formation of leader cells driven by CD44 switching gives rise to collective invasion and metastases in luminal breast carcinomas. Oncogene 38, 7113–7132 (2019)
C. Chen, S. Zhao, X. Zhao, L. Cao, A. Karnad, A.P. Kumar, J.W. Freeman, Gemcitabine resistance of pancreatic cancer cells is mediated by IGF1R dependent upregulation of CD44 expression and isoform switching. Cell. Death Dis. 13, 682 (2022)
M. Liu, Y. Zhang, J. Yang, X. Cui, Z. Zhou, H. Zhan, K. Ding, X. Tian, Z. Yang, K.A. Fung, B.H. Edil, R.G. Postier, M.S. Bronze, M.E. Fernandez-Zapico, M.P. Stemmler, T. Brabletz, Y.P. Li, C.W. Houchen, M. Li, ZIP4 increases expression of transcription factor ZEB1 to promote integrin α3β1 signaling and inhibit expression of the Gemcitabine Transporter ENT1 in Pancreatic Cancer cells. Gastroenterology 158, 679–692e671 (2020)
F.M. Platt, A. d’Azzo, B.L. Davidson, E.F. Neufeld, C.J. Tifft, Lysosomal storage diseases. Nat. Rev. Dis. Primers. 4, 27 (2018)
X. Zhai, Y. El, Hiani, Getting lost in the cell–lysosomal entrapment of chemotherapeutics. Cancers 12, 3669 (2020)
A.M. Chamoun-Emanuelli, L.K. Bryan, N.D. Cohen, T.L. Tetrault, J.A. Szule, R. Barhoumi, Whitfield-Cargile, NSAIDs disrupt intestinal homeostasis by suppressing macroautophagy in intestinal epithelial cells. Sci. Rep. 9, 14534 (2019)
F.Y. Ke, W.Y. Chen, M.C. Lin, Y.C. Hwang, K.T. Kuo, H.C. Wu, Novel monoclonal antibody against integrin α3 shows therapeutic potential for ovarian cancer. Cancer Sci. 111, 3478–3492 (2020)
L. Shi, B. Liu, D.-. Shen, P. Yan, Y. Zhang, Y. Tian, L. Hou, G. Jiang, Y. Zhu, Y. Liang, A tumor-suppressive circular RNA mediates uncanonical integrin degradation by the proteasome in liver cancer. Sci. Adv. 7, eabe5043 (2021)
A. Okato, Y. Goto, A. Kurozumi, M. Kato, S. Kojima, R. Matsushita, M. Yonemori, K. Miyamoto, T. Ichikawa, N. Seki, Direct regulation of LAMP1 by tumor-suppressive microRNA-320a in prostate cancer. Int. J. Oncol. 49, 111–122 (2016)
Y. Hu, J. Pan, P. Shah, M. Ao, S.N. Thomas, Y. Liu, L. Chen, M. Schnaubelt, D.J. Clark, H. Rodriguez, Integrated proteomic and glycoproteomic characterization of human high-grade serous ovarian carcinoma. Cell. Rep. 33, 108276 (2020)
M.K. McKenna, A. Ozcan, D. Brenner, N. Watanabe, M. Legendre, D.G. Thomas, C. Ashwood, R.D. Cummings, C. Bonifant, D.M. Markovitz, Novel banana lectin CAR-T cells to target pancreatic tumors and tumor-associated stroma. J. Immunother Cancer 11, e005891 (2023)
N. Taniguchi, Y. Kizuka, Glycans and cancer: role of N-glycans in cancer biomarker, progression and metastasis, and therapeutics. Adv. Cancer Res. 126, 11–51 (2015)
E. RodrÍguez, S.T.T. Schetters, Y. van Kooyk, The tumour glyco-code as a novel immune checkpoint for immunotherapy. Nat. Rev. Immunol. 18, 204–211 (2018)
R. Gupta, F. Leon, C.M. Thompson, R. Nimmakayala, S. Karmakar, P. Nallasamy, S. Chugh, D.R. Prajapati, S. Rachagani, S. Kumar, Global analysis of human glycosyltransferases reveals novel targets for pancreatic cancer pathogenesis. Br. J. Cancer. 122, 1661–1672 (2020)
Y. Li, Y. Lin, L. Aye, L. Dong, C. Zhang, F. Chen, Y. Liu, J. Fan, Q. Gao, H. Lu, An integrative pan-cancer analysis of the molecular and biological features of glycosyltransferases. Clin. Transl Med. 12, (2022)
M.U. Rajagopal, S. Bansal, P. Kaur, S.K. Jain, T. Altadil, C.P. Hinzman, Y. Li, J. Moulton, B. Singh, S. Bansal, TGFβ drives metabolic perturbations during epithelial mesenchymal transition in pancreatic Cancer: TGFβ Induced EMT in PDAC. Cancers. 13, 6204 (2021)
D.A. Polasky, F. Yu, G.C. Teo, A.I. Nesvizhskii, Fast and comprehensive N-and O-glycoproteomics analysis with MSFragger-Glyco. Nat. methods. 17, 1125–1132 (2020)
Y. Liao, J. Wang, E.J. Jaehnig, Z. Shi, B. Zhang, WebGestalt 2019: gene set analysis toolkit with revamped UIs and APIs. Nucleic Acids Res. 47, W199–w205 (2019)
Funding
This work was supported by the National Key Program for Basic Research of China (grant numbers 2020YFC2002700, 2020YFE0202200); and Research Program of the State Key Laboratory of Proteomics [grant number SKLP-K201901].
Author information
Authors and Affiliations
Contributions
X.L. and R.K. contributed to the work equally and should be regarded as co-first authors.X.L : Conceptualization; Experimenter; Data curation; Formal analysis; Visualization; Roles/Writing original draft;R.K : Investigation; Roles/Writing original draft; Experimenter;W.H : Writing editing;L.Z : Formal analysis; J.C : Experimenter;X.Q : Project administration; Resources; Funding acquisition;L.Z : Project administrationW.Y : Project administration; Resources; Funding acquisition; Writing review & editing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
Not applicable.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, X., Kong, R., Hou, W. et al. Integrative proteomics and n-glycoproteomics reveal the synergistic anti-tumor effects of aspirin- and gemcitabine-based chemotherapy on pancreatic cancer cells. Cell Oncol. 47, 141–156 (2024). https://doi.org/10.1007/s13402-023-00856-z
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13402-023-00856-z