Abstract
Purpose
Pancreatic ductal adenocarcinoma (PDAC) is an aggressive disease with a high mortality rate, in which about 90% of patients harbor somatic oncogenic point mutations in KRAS. SPRY family genes have been recognized as crucial negative regulators of Ras/Raf/ERK signaling. Here, we investigate the expression and role of SPRY proteins in PDAC.
Methods
Expression of SPRY genes in human and mice PDAC was analyzed using The Cancer Genome Atlas and Gene Expression Omnibus datasets, and by immunohistochemistry analysis. Gain-of-function, loss-of-function of Spry1 and orthotopic xenograft model were adopted to investigate the function of Spry1 in mice PDAC. Bioinformatics analysis, transwell and flowcytometry analysis were used to identify the effects of SPRY1 on immune cells. Co-immunoprecipitation and K-ras4B G12V overexpression were used to identify molecular mechanism.
Results
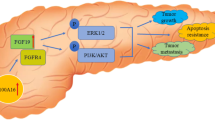
SPRY1 expression was remarkably increased in PDAC tissues and positively associated with poor prognosis of PDAC patients. SPRY1 knockdown suppressed tumor growth in mice. SPRY1 was found to promote CXCL12 expression and facilitate neutrophil and macrophage infiltration via CXCL12-CXCR4 axis. Pharmacological inhibition of CXCL12-CXCR4 largely abrogated the oncogenic functions of SPRY1 by suppressing neutrophil and macrophage infiltration. Mechanistically, SPRY1 interacted with ubiquitin carboxy-terminal hydrolase L1 to induce activation of nuclear factor κB signaling and ultimately increase CXCL12 expression. Moreover, SPRY1 transcription was dependent on KRAS mutation and was mediated by MAPK-ERK signaling.
Conclusion
High expression of SPRY1 can function as an oncogene in PDAC by promoting cancer-associated inflammation. Targeting SPRY1 might be an important approach for designing new strategy of tumor therapy.
Highlights
The expression of SPRY1 is increased during malignant transformation and predicts a poor prognosis in PDAC.
SPRY1 promotes neutrophil and macrophage recruitment via CXCL12-CXCR4 axis.
SPRY1 modulates CXCL12 expression through promoting nuclear factor κB signaling activation via interaction with ubiquitin carboxy-terminal hydrolase L1.
SPRY1 transcription is dependent on KRAS mutation and mediated by MAPK-ERK signaling in PDAC.







Similar content being viewed by others
Data availability
The data are available within the Article, Supplementary Information, or available from the authors upon request. Source data are provided with this paper.
References
J.M. Mason, D.J. Morrison, M.A. Basson, J.D. Licht, Trends Cell Biol. 16, 45–54 (2006). https://doi.org/10.1016/j.tcb.2005.11.004
S. Masoumi-Moghaddam, A. Amini, D.L. Morris, Cancer Metastasis Rev. 33, 695–720 (2014). https://doi.org/10.1007/s10555-014-9497-1
K. Bundschu, U. Walter, K. Schuh, BioEssays. 29, 897–907 (2007). https://doi.org/10.1002/bies.20632
K. Ozaki, R. Kadomoto, K. Asato, S. Tanimura, N. Itoh, M. Kohno, Biochem. Biophys. Res. Commun. 285, 1084–1088 (2001). https://doi.org/10.1006/bbrc.2001.5295
Z. Koledova, X. Zhang, C. Streuli, R.B. Clarke, O.D. Klein, Z. Werb, P. Lu, Proc. Natl. Acad. Sci. U. S. A. 113, E5731–E5740 (2016). https://doi.org/10.1073/pnas.1611532113
A.K. Perl, I. Hokuto, M.A. Impagnatiello, G. Christofori, J.A. Whitsett, Dev. Biol. 258, 154–168 (2003). https://doi.org/10.1016/s0012-1606(03)00106-4
V. Sigurdsson, S. Ingthorsson, B. Hilmarsdottir, S.M. Gustafsdottir, S.R. Franzdottir, A.J. Arason, E. Steingrimsson, M.K. Magnusson, T. Gudjonsson, PLoS One 8, e60798 (2013). https://doi.org/10.1371/journal.pone.0060798
K.B. Sylvestersen, P.L. Herrera, P. Serup, C. Rescan, Gene Expr. Patterns 11, 105–111 (2011). https://doi.org/10.1016/j.gep.2010.10.001
T. Kawazoe, K. Taniguchi, Cancer Sci. 110, 1525–1535 (2019). https://doi.org/10.1111/cas.13999
T.L. Lo, P. Yusoff, C.W. Fong, K. Guo, B.J. McCaw, W.A. Phillips, H. Yang, E.S. Wong, H.F. Leong, Q. Zeng, T.C. Putti, G.R. Guy, Cancer Res. 64, 6127–6136 (2004). https://doi.org/10.1158/0008-5472.CAN-04-1207
M.A. Tennis, M.M. Van Scoyk, S.V. Freeman, K.M. Vandervest, R.A. Nemenoff, R.A. Winn, Mol. Cancer Res.: MCR 8, 833–843 (2010). https://doi.org/10.1158/1541-7786.MCR-09-0400
S.A. Lee, C. Ho, R. Roy, C. Kosinski, M.A. Patil, A.D. Tward, J. Fridlyand, X. Chen, Hepatology 47, 1200–1210 (2008). https://doi.org/10.1002/hep.22169
P. Lito, B.D. Mets, S. Kleff, S. O'Reilly, V.M. Maher, J.J. McCormick, J. Biol. Chem. 283, 2002–2009 (2008). https://doi.org/10.1074/jbc.M709046200
C. Holgren, U. Dougherty, F. Edwin, D. Cerasi, I. Taylor, A. Fichera, L. Joseph, M. Bissonnette, S. Khare, Oncogene 29, 5241–5253 (2010). https://doi.org/10.1038/onc.2010.264
S. Bloethner, B. Chen, K. Hemminki, J. Muller-Berghaus, S. Ugurel, D. Schadendorf, R. Kumar, Carcinogenesis 26, 1224–1232 (2005). https://doi.org/10.1093/carcin/bgi066
K.M. Mann, H. Ying, J. Juan, N.A. Jenkins, N.G. Copeland, Pharmacol. Ther. 168, 29–42 (2016). https://doi.org/10.1016/j.pharmthera.2016.09.003
T.J. Grant, K. Hua, A. Singh, Prog. Mol. Biol. Transl. Sci. 144, 241–275 (2016). https://doi.org/10.1016/bs.pmbts.2016.09.008
J. Vivian, A.A. Rao, F.A. Nothaft, C. Ketchum, J. Armstrong, A. Novak, J. Pfeil, J. Narkizian, A.D. Deran, A. Musselman-Brown, H. Schmidt, P. Amstutz, B. Craft, M. Goldman, K. Rosenbloom, M. Cline, B. O'Connor, M. Hanna, C. Birger, et al., Nat. Biotechnol. 35, 314–316 (2017). https://doi.org/10.1038/nbt.3772
J. Yang, A. Kumar, A.E. Vilgelm, S.C. Chen, G.D. Ayers, S.V. Novitskiy, S. Joyce, A. Richmond, Cancer Immunol. Res. 6, 1186–1198 (2018). https://doi.org/10.1158/2326-6066.CIR-18-0045
A. Schernberg, P. Blanchard, C. Chargari, E. Deutsch, Acta Oncol. 56, 1522–1530 (2017). https://doi.org/10.1080/0284186X.2017.1348623
Z. Zhang, N. Liu, X. Chen, F. Zhang, T. Kong, X. Tang, Q. Yang, W. Chen, X. Xiong, X. Chen, Cell Biol. Int. 45, 2107–2117 (2021). https://doi.org/10.1002/cbin.11662
B. Montico, F. Colizzi, G. Giurato, A. Rizzo, A. Salvati, L. Baboci, D. Benedetti, E. Pivetta, A. Covre, M.D. Bo, A. Weisz, A. Steffan, M. Maio, L. Sigalotti, E. Fratta, Cell Death Dis. 11, 392 (2020). https://doi.org/10.1038/s41419-020-2585-y
Q. He, H. Jing, L. Liaw, L. Gower, C. Vary, S. Hua, X. Yang, Sci. Rep. 6, 23216 (2016). https://doi.org/10.1038/srep23216
Q. Zhang, T. Wei, K. Shim, K. Wright, K. Xu, H.L. Palka-Hamblin, A. Jurkevich, S. Khare, Oncogene 35, 3151–3162 (2016). https://doi.org/10.1038/onc.2015.365
G. Schaaf, M. Hamdi, D. Zwijnenburg, A. Lakeman, D. Geerts, R. Versteeg, M. Kool, Cancer Res. 70, 762–771 (2010). https://doi.org/10.1158/0008-5472.CAN-09-2532
Y. Xiao, M. Cong, J. Li, D. He, Q. Wu, P. Tian, Y. Wang, S. Yang, C. Liang, Y. Liang, J. Wen, Y. Liu, W. Luo, X. Lv, Y. He, D.D. Cheng, T. Zhou, W. Zhao, P. Zhang, et al., Cancer Cell 39, 423–437 e427 (2021). https://doi.org/10.1016/j.ccell.2020.12.012
F. Mollinedo, Trends Immunol. 40, 228–242 (2019). https://doi.org/10.1016/j.it.2019.01.006
M. Huber, C.U. Brehm, T.M. Gress, M. Buchholz, B. Alashkar Alhamwe, E.P. von Strandmann, E.P. Slater, J.W. Bartsch, C. Bauer, M. Lauth, Int. J. Mol. Sci. 21 (2020). https://doi.org/10.3390/ijms21197307
B.A. Teicher, S.P. Fricker, Clin. Cancer Res. 16, 2927–2931 (2010). https://doi.org/10.1158/1078-0432.CCR-09-2329
Z. Peng, C. Liu, A.R. Victor, D.Y. Cao, L.C. Veiras, E.A. Bernstein, Z. Khan, J.F. Giani, X. Cui, K.E. Bernstein, D. Okwan-Duodu, Oncoimmunology 10, 1870811 (2021). https://doi.org/10.1080/2162402X.2020.1870811
R.V. Sionov, Z.G. Fridlender, Z. Granot, Cancer Microenviron. 8, 125–158 (2015). https://doi.org/10.1007/s12307-014-0147-5
A. Macia, M. Vaquero, M. Gou-Fabregas, E. Castelblanco, J.M. Valdivielso, C. Anerillas, D. Mauricio, X. Matias-Guiu, J. Ribera, M. Encinas, Cell Death Differ. 21, 333–343 (2014). https://doi.org/10.1038/cdd.2013.161
S.H. Chiou, P. Shahi, R.T. Wagner, H. Hu, N. Lapteva, M. Seethammagari, S.C. Sun, J.M. Levitt, D.M. Spencer, EMBO Rep. 12, 971–979 (2011). https://doi.org/10.1038/embor.2011.143
Acknowledgments
We thank Prof. Xiongjun Wang for technical support.
Funding
This study was supported by the National Natural Science Foundation of China (No. 31801212 to LY). Beijing Natural Science Foundation (No. 5232003 to LW).
Author information
Authors and Affiliations
Contributions
LY, JY and LW conceived the project; LY and JY designed experiments, interpreted data in the manuscript and wrote the manuscript; LW performed bioinformatics analyses, data analysis and wrote the manuscript; XL, JZ, LY and TS performed cell and animal experiments; JY provided clinical specimens and made clinical pathology evaluations; XL, JZ and ZD performed western blotting analysis and IHC analysis; OYY, YG and QW reviewed the manuscript and discussed the data. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethical approval
All procedures related to patients were carried out in accordance with International Ethical Guidelines for Biomedical Research Involving Human Subjects (CIOMS). The study was approved by the Research Ethics Committee of Ren Ji Hospital, School of Medicine, Shanghai Jiao Tong University. All the mouse experiments were approved by the Research Ethics Committee of Shanghai Jiao Tong University.
Consent for publication
All authors had agreed to publish this manuscript.
Competing interests
The authors have declared that no conflict of interest exists.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Shi, T., Li, X., Zheng, J. et al. Increased SPRY1 expression activates NF-κB signaling and promotes pancreatic cancer progression by recruiting neutrophils and macrophages through CXCL12-CXCR4 axis. Cell Oncol. 46, 969–985 (2023). https://doi.org/10.1007/s13402-023-00791-z
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13402-023-00791-z