Abstract
Purpose
Transcriptional addiction plays a pivotal role in maintaining the hallmarks of cancer cells. Thus, targeting super-enhancers (SEs), which modulate the transcriptional activity of oncogenes, has become an attractive strategy for cancer therapy. As yet, however, the molecular mechanisms of this process in bladder cancer (BC) remain to be elucidated. Here, we aimed to provide detailed information regarding the SE landscape in BC and to investigate new potential pharmaceutical targets for BC therapy.
Methods
We employed THZ1 as a potent and specific CDK7 inhibitor. In vitro and in vivo studies were carried out to investigate the anticancer and apoptosis-inducing effects of THZ1 on BC cells. Whole-transcriptome sequencing (RNA-seq) and chromatin immunoprecipitation sequencing (ChIP-seq) were performed to investigate the mechanism and function of SE-linked oncogenic transcription in BC cells.
Results
We found that THZ1 serves as an effective and potent inhibitor with suppressive activity against BC cells. An integrative analysis of THZ1-sensitive and SE-associated oncogenes yielded potential new pharmaceutical targets, including DDIT4, B4GALT5, PSRC1 and MED22. Combination treatment with THZ1 and the DDIT4 inhibitor rapamycin effectively suppressed BC cell growth. In addition, we found that THZ1 and rapamycin sensitized BC cells to conventional chemotherapy.
Conclusions
Our data indicate that exploring BC gene regulatory mechanisms associated with SEs through integrating RNA-seq and ChIP-seq data improves our understanding of BC biology and provides a basis for innovative therapies.







Similar content being viewed by others
References
R.L. Siegel, K.D. MillerA, Jemal, Cancer statistics, 2019. CA Cancer J. Clin. 69, 7–34 (2019)
S. Antoni, J. Ferlay, I. Soerjomataram, A. Znaor, A. Jemal, F. Bray, Bladder cancer incidence and mortality: a global overview and recent trends. Eur. Urol. 71, 96–108 (2017)
N. Howlader, K.M. Na, D. Miller, A. Brest, M. Yu, J. Ruhl, Z. Tatalovich, A. Mariotto, D. Lewis, H. Chen, SEER cancer statistics review, 1975–2016. Natl. Cancer Inst. 26,1423-37 (2019)
J.I. Warrick, G. Sjödahl, M. Kaag, J.D. Raman, S. Merrill, L. Shuman, G. Chen, V. Walter, D.J. DeGraff, Intratumoral heterogeneity of bladder cancer by molecular subtypes and histologic variants. Eur. Urol. 75, 18–22 (2019)
D. Hanahan, R.A. Weinberg, Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011)
J.E. Bradner, D. Young, R.A. Hnisz, Transcriptional addiction in cancer. Cell 168, 629–643 (2017)
T.I. Lee, R.A. Young, Transcriptional regulation and its misregulation in disease. Cell 152, 1237–1251 (2013)
B. Donati, E. Lorenzini, A. Ciarrocchi, BRD4 and cancer: going beyond transcriptional regulation. Mol. Cancer 17, 164 (2018)
S. Sengupta, R.E. George, Super-enhancer-driven transcriptional dependencies in cancer. Trends Cancer 3, 269–281 (2017)
J. Yuan, Y.-Y. Jiang, A. Mayakonda, M. Huang, L.-W. Ding, H. Lin, F. Yu, Y. Lu, T.K. Loh, M. Chow, Super-enhancers promote transcriptional dysregulation in nasopharyngeal carcinoma. Cancer Res. 77, 6614–6626 (2017)
L. Lin, M. Huang, X. Shi, A. Mayakonda, K. Hu, Y.-Y. Jiang, X. Guo, L. Chen, B. Pang, N. Doan, Super-enhancer-associated MEIS1 promotes transcriptional dysregulation in Ewing sarcoma in co-operation with EWS-FLI1. Nucleic Acids Res. 47, 1255–1267 (2019)
Y. Wang, T. Zhang, N. Kwiatkowski, B.J. Abraham, T.I. Lee, S. Xie, H. Yuzugullu, T. Von, H. Li, Z. Lin, CDK7-dependent transcriptional addiction in triple-negative breast cancer. Cell 163, 174–186 (2015)
J. Chou, D.A. Quigley, T.M. Robinson, F.Y. FengA, A. Ashworth, Transcription-associated cyclin-dependent kinases as targets and biomarkers for cancer therapy. Cancer Discov. 10, 351–370 (2020)
B.-B. Li, B. Wang, C.-M. Zhu, D. Tang, J. Pang, J. Zhao, C.-H. Sun, M.-J. Qiu, Z.-R. Qian, Cyclin-dependent kinase 7 inhibitor THZ1 in cancer therapy. Chronic Dis. Transl. Med. 5, 155–169 (2019)
K.A. Nilson, J. Guo, M.E. Turek, J.E. Brogie, E. Delaney, D.S. Luse, D.H. Price, THZ1 reveals roles for Cdk7 in co-transcriptional capping and pausing. Mol. Cell 59, 576–587 (2015)
N. Kwiatkowski, T. Zhang, P.B. Rahl, B.J. Abraham, J. Reddy, S.B. Ficarro, A. Dastur, A. Amzallag, S. Ramaswamy, B. Tesar, Targeting transcription regulation in cancer with a covalent CDK7 inhibitor. Nature 511, 616–620 (2014)
J. Wang, R. Zhang, Z. Lin, S. Zhang, Y. Chen, J. Tang, J. Hong, X. Zhou, Y. Zong, Y. Xu, CDK7 inhibitor THZ1 enhances antiPD-1 therapy efficacy via the p38α/MYC/PD-L1 signaling in non-small cell lung cancer. J. Hematol. Oncol. 13, 1–16 (2020)
D. Hnisz, B.J. Abraham, T.I. Lee, A. Lau, V. Saint-André, A.A. Sigova, H.A. Hoke, R.A. Young, Super-enhancers in the control of cell identity and disease. Cell 155, 934–947 (2013)
P. Thandapani, Super-enhancers in cancer. Pharmacol. Ther. 199, 129–138 (2019)
B.J. Greber, J.M. Perez-Bertoldi, K. Lim, A.T. Iavarone, D.B. TosoE, E. Nogales, The cryo-electron microscopy structure of the human CDK-activating kinase. Proc. Natl. Acad. Sci. 37, 22849–22857 (2020)
M.S. Akhtar, M. Heidemann, J.R. Tietjen, D.W. Zhang, R.D. Chapman, D. Eick, A.Z. Ansari, TFIIH kinase places bivalent marks on the carboxy-terminal domain of RNA polymerase II. Mol. Cell 34, 387–393 (2009)
S. Larochelle, R. Amat, K. Glover-Cutter, M. Sansó, C. Zhang, J.J. Allen, K.M. Shokat, D.L. Bentley, R.P. Fisher, Cyclin-dependent kinase control of the initiation-to-elongation switch of RNA polymerase II. Nat. Struct. Mol. Biol. 19, 1108 (2012)
K. Glover-Cutter, S. Larochelle, B. Erickson, C. Zhang, K. Shokat, R.P. Fisher, D.L. Bentley, TFIIH-associated Cdk7 kinase functions in phosphorylation of C-terminal domain Ser7 residues, promoter-proximal pausing, and termination by RNA polymerase II. Mol. Cell. Biol. 29, 5455–5464 (2009)
X. Cao, L. Dang, X. Zheng, Y. Lu, Y. Lu, R. Ji, T. Zhang, X. Ruan, J. Zhi, X. Hou, Targeting super-enhancer-driven oncogenic transcription by CDK7 inhibition in anaplastic thyroid carcinoma. Thyroid 29, 809–823 (2019)
Y.-Y. Jiang, D.-C. Lin, A. Mayakonda, M. Hazawa, L.-W. Ding, W.-W. Chien, L. Xu, Y. Chen, W. Senapedis, Targeting super-enhancer-associated oncogenes in oesophageal squamous cell carcinoma. Gut 66, 1358–1368 (2017)
E. Chipumuro, E. Marco, C.L. Christensen, N. Kwiatkowski, T. Zhang, C.M. Hatheway, B.J. Abraham, B. Sharma, C. Yeung, A. Altabef, CDK7 inhibition suppresses super-enhancer-linked oncogenic transcription in MYCN-driven cancer. Cell 159, 1126–1139 (2014)
H.E. Pelish, B.B. Liau, I.I. Nitulescu, A. Tangpeerachaikul, Z.C. Poss, D.H. Da Silva, B.T. Caruso, A. Arefolov, O. Fadeyi, A.L. Christie, Mediator kinase inhibition further activates super-enhancer-associated genes in AML. Nature 526, 273–276 (2015)
M. Noda, M. Vallon, C.J. Kuo, The Wnt7’s Tale: a story of an orphan who finds her tie to a famous family. Cancer Sci. 107, 576–582 (2016)
J. Bellmunt, Stem-like signature predicting disease progression in early stage bladder cancer. The role of E2F3 and SOX4. Biomedicines 6, 85 (2018)
B.A. Mooso, R.L. Vinall, M. Mudryj, S.A. Yap, RWd. White, P.M. Ghosh, The role of EGFR family inhibitors in muscle invasive bladder cancer: a review of clinical data and molecular evidence. J. Urol. 193, 19–29 (2015)
N. Zhang, X. Zeng, C. Sun, H. Guo, T. Wang, L. Wei, Y. Zhang, J.Z.X. Ma, LncRNA LINC00963 promotes tumorigenesis and radioresistance in breast cancer by sponging miR-324-3p and inducing ACK1 expression. Mol. Ther.–Nucleic Acids 18, 871–881 (2019)
Y.S.L. Ma, New insights into long non-coding RNA MALAT1 in cancer and metastasis. Cancers 11, 216 (2019)
S. Shen, J. Wang, B. Zheng, Y. Tao, M. Li, Y. Wang, X. Ni, T. Suo, H. Liu, H. Liu, LINC01714 enhances gemcitabine sensitivity by modulating FOXO3 phosphorylation in cholangiocarcinoma. Mol. Ther.–Nucleic Acids 19, 446–457 (2020)
Q. He, L. Huang, D. Yan, J. Bi, M. Yang, J.H.T. Lin, CircPTPRA acts as a tumor suppressor in bladder cancer by sponging miR-636 and upregulating KLF9. Aging 11, 11314 (2019)
E. Lesovaya, S. Agarwal, B. Readhead, E. Vinokour, G. Baida, P. Bhalla, K. Kirsanov, M. Yakubovskaya, L.C. Platanias, J.T. Dudley, Rapamycin modulates glucocorticoid receptor function, blocks atrophogene REDD1, and protects skin from steroid atrophy. J. Invest. Derm. 138, 1935–1944 (2018)
T.-C. Chou, Drug combination studies and their synergy quantification using the Chou-Talalay method. Cancer Res. 70, 440–446 (2010)
M. Zarei, H. Du, A.H. Nassar, R.E. Yan, K. Giannikou, S.H. Johnson, H.C. Lam, E.P. Henske, Y. Wang, T. Zhang, Tumors with TSC mutations are sensitive to CDK7 inhibition through NRF2 and glutathione depletion. J. Exp. Med. 216, 2635–2652 (2019)
J. Earl, D. Rico, E. Carrillo-de-Santa-Pau, B. Rodríguez-Santiago, M. Méndez-Pertuz, H. Auer, G. Gómez, H.B. Grossman, D.G. Pisano, W.A. Schulz, The UBC-40 Urothelial Bladder Cancer cell line index: a genomic resource for functional studies. BMC Genom. 16, 403 (2015)
F. Du, L. Sun, Y. Chu, T. Li, C. Lei, X. Wang, M. Jiang, Y. Min, Y. Lu, X. Zhao, DDIT4 promotes gastric cancer proliferation and tumorigenesis through the p53 and MAPK pathways. Cancer Commun. 38, 45 (2018)
R.N.J. Bellmunt, Management of metastatic bladder cancer. Cancer Treat. Rev. 76, 10–21 (2019)
P.J. Loehrer Sr, L.H. Einhorn, P.J. Elson, E.D. Crawford, P. Kuebler, I. Tannock, D. Raghavan, R. Stuart-Harris, M.F. Sarosdy, B.A. Lowe, A randomized comparison of cisplatin alone or in combination with methotrexate, vinblastine, and doxorubicin in patients with metastatic urothelial carcinoma: a cooperative group study. J. Clin. Oncol. 10, 1066–1073 (1992)
M. Yu, T. Ozaki, D. Sun, H. Xing, B. Wei, J. An, J. Yang, Y. Gao, S.L.C. Kong, HIF-1α-dependent miR-424 induction confers cisplatin resistance on bladder cancer cells through down-regulation of pro-apoptotic UNC5B and SIRT4. J. Exp. Clin. Cancer Res. 39, 1–13 (2020)
M.A. Knowles, C.D. Hurst, Molecular biology of bladder cancer: new insights into pathogenesis and clinical diversity. Nat. Rev. Cancer 15, 25–41 (2015)
J. Earl, D. Rico, E. Carrillo-de-Santa-Pau, B. Rodríguez-Santiago, M. Méndez-Pertuz, H. Auer, G. Gómez, H.B. Grossman, D.G. Pisano, W.A. Schulz, The UBC-40 Urothelial Bladder Cancer cell line index: a genomic resource for functional studies. BMC Genom. 16, 1–16 (2015)
M. Razmara, A.M.B. Skogseid, Reduced menin expression impairs rapamycin effects as evidenced by an increase in mTORC2 signaling and cell migration. Cell. Commun. Signal 16, 64 (2018)
J.J. Oh, S.H. Ji, D.K. Choi, I.H. Gong, T.H. Kim, D.S. Park, A six-week course of bacillus Calmette-Guerin prophylaxis is insufficient to prevent tumor recurrence in nonmuscle invasive bladder cancer with strong-positive expression of p53. Oncology 79, 440–446 (2010)
X. Zhou, G. Zhang, Y. Tian, p53 status correlates with the risk of recurrence in non-muscle invasive bladder cancers treated with bacillus Calmette–Guérin: a meta-analysis. PLoS One 10, e0119476 (2015)
E.M. Alexandrova, A.R. Yallowitz, D. Li, S. Xu, R. Schulz, D.A. Proia, G. Lozano, M. Dobbelstein, U.M. Moll, Improving survival by exploiting tumour dependence on stabilized mutant p53 for treatment. Nature 523, 352–356 (2015)
Acknowledgements
The present study was funded by the National Key R&D Program of China (2018YFA0902800), the National Natural Science Foundation of China (81772754), the Major Basic Research and Cultivation Program of Natural Science Foundation of Guangdong Province (2017A03038009), the Shenzhen Basic Science Research program (JCYJ20190809164617205), the Sanming Project of Medicine in Shenzhen (SZSM202011011), Shenzhen Key Laboratory Program (ZDSYS20190902092857146), the Hospital Research Fund of SAHSYSU (ZSQYLCKYJJ202019) and a Research Start-up Fund of part-time PI, SAHSYSU (ZSQYJZPI202003).
Author information
Authors and Affiliations
Contributions
Song Wu and Jun Pang designed the experiments. Yafei Yang, Donggen Jiang and Ziyu Zhou collected the data and analyzed the results. Haiyun Xiong, Xiangwei Yang, Guoyu Peng, Wuchao Xia, Shang Wang, Hanqi Lei, Jing Zhao and Zhirong Qian provided conceptual advice. Yafei Yang, Donggen Jiang and Ziyu Zhou wrote the paper. Song Wu and Jun Pang reviewed and edited the manuscript. Song Wu and Jun Pang obtained funding. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Research involving human participants and/or animals
This study was approved by the ethics committee of the Seventh Affiliated Hospital of Sun Yat-sen University. All animal experiments were performed in accordance with the established guidelines of the Animal Research Ethics Committee of the Seventh Affiliated Hospital of Sun Yat-sen University.
Informed consent
Not applicable.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Supplementary Fig. 1
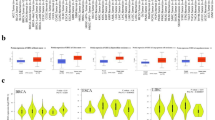
(a) Docking model of THZ1 in the ATP-binding pocket of CDK7. CDK7 marked with green and THZ1 depicted with iridescence, the binding site Cys312 is in red. (b) MGH-U3, 5637 and human urothelial SV-HUC-1 cells were treated with 100 nM THZ1, cell morphology and (c) colony forming ability were recorded. (d) Apoptosis analysis in MGH-U3 and 5637 cell lines treated with escalating doses for 48 h. (e) mRNA expression of representative transcripts (DDIT4, B4GALT5, EGFR and MED22) across various types of human cancer cells. Data were retrieved from the TCGA and GTEX projects (PNG 3783 kb)
Supplementary Fig. 2
The correlation between selected candidate oncogene transcript levels and the OS of BC patients from public data sets plotted by the Kaplan-Meier method (PNG 514 kb)
Supplementary Fig. 3
Sequence verification of CRISPR targeting regions in the indicated genes (PNG 1457 kb)
Table S1
(XLSX 13 kb)
Table S2
(XLSX 8216 kb)
Table S3
(XLSX 285 kb)
Table S4
(XLSX 9 kb)
ESM 1
(DOCX 15 kb)
Rights and permissions
About this article
Cite this article
Yang, Y., Jiang, D., Zhou, Z. et al. CDK7 blockade suppresses super‐enhancer‐associated oncogenes in bladder cancer. Cell Oncol. 44, 871–887 (2021). https://doi.org/10.1007/s13402-021-00608-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13402-021-00608-x