Abstract
Background
MicroRNAs (miRNAs) are known to play an important role in cancer development by post-transcriptionally affecting the expression of critical genes. The aims of this study were two-fold: (i) to develop a robust method to isolate miRNAs from small volumes of saliva and (ii) to develop a panel of saliva-based diagnostic biomarkers for the detection of head and neck squamous cell carcinoma (HNSCC).
Methods
Five differentially expressed miRNAs were selected from miScript™ miRNA microarray data generated using saliva from five HNSCC patients and five healthy controls. Their differential expression was subsequently confirmed by RT-qPCR using saliva samples from healthy controls (n = 56) and HNSCC patients (n = 56). These samples were divided into two different cohorts, i.e., a first confirmatory cohort (n = 21) and a second independent validation cohort (n = 35), to narrow down the miRNA diagnostic panel to three miRNAs: miR-9, miR-134 and miR-191. This diagnostic panel was independently validated using HNSCC miRNA expression data from The Cancer Genome Atlas (TCGA), encompassing 334 tumours and 39 adjacent normal tissues. Receiver operating characteristic (ROC) curve analysis was performed to assess the diagnostic capacity of the panel.
Results
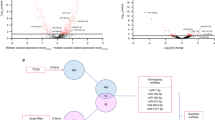
On average 60 ng/μL miRNA was isolated from 200 μL of saliva. Overall a good correlation was observed between the microarray data and the RT-qPCR data. We found that miR-9 (P <0.0001), miR-134 (P <0.0001) and miR-191 (P <0.001) were differentially expressed between saliva from HNSCC patients and healthy controls, and that these miRNAs provided a good discriminative capacity with area under the curve (AUC) values of 0.85 (P <0.0001), 0.74 (P < 0.001) and 0.98 (P < 0.0001), respectively. In addition, we found that the salivary miRNA data showed a good correlation with the TCGA miRNA data, thereby providing an independent validation.
Conclusions
We show that we have developed a reliable method to isolate miRNAs from small volumes of saliva, and that the saliva-derived miRNAs miR-9, miR-134 and miR-191 may serve as novel biomarkers to reliably detect HNSCC.




Similar content being viewed by others
References
D. Chin, G.M. Boyle, S. Porceddu, D.R. Theile, P.G. Parsons, W.B. Coman, Head and neck cancer: past, present and future. Expert. Rev. Anticancer. Ther. 6, 1111–1118 (2006)
D. Weiss, C. Stockmann, K. Schrodter, C. Rudack, Protein expression and promoter methylation of the candidate biomarker TCF21 in head and neck squamous cell carcinoma. Cell. Oncol. 36, 213–224 (2013)
T. Nakaoka, A. Ota, T. Ono, S. Karnan, H. Konishi, A. Furuhashi, Y. Ohmura, Y. Yamada, Y. Hosokawa, Y. Kazaoka, Combined arsenic trioxide-cisplatin treatment enhances apoptosis in oral squamous cell carcinoma cells. Cell. Oncol. 37, 119–129 (2014)
P.A. Wingo, T. Tong, S. Bolden, Cancer statistics for African Americans. CA Cancer J. Clin. 45, 8–30 (1995)
D.G. Haller, L.D. Wagma, K.A. Camphausen and W.J. Hoskins, Cancer Management Handbook, 12th Edition. (Cancer Network, 2013), http://www.cancernetwork.com/cancer-management-12
T. Pfaffe, J. Cooper-White, P. Beyerlein, K. Kostner, C. Punyadeera, Diagnostic potential of saliva: current state and future applications. Clin. Chem. 57, 675–687 (2011)
R. Nagadia, P. Pandit, W.B. Coman, J. Cooper-White, C. Punyadeera, miRNAs in head and neck cancer revisited. Cell. Oncol. 36, 1–7 (2013)
U.M. Bailey, C. Punyadeera, J.J. Cooper-White, B.L. Schulz, Analysis of the extreme diversity of salivary alpha-amylase isoforms generated by physiological proteolysis using liquid chromatography-tandem mass spectrometry. J. Chromatogr. B Analyt. Technol. Biomed. Life Sci. 911, 21–26 (2012)
B.L. Schulz, J. Cooper-White, C.K. Punyadeera, Saliva proteome research: current status and future outlook. Crit. Rev. Biotechnol. 33, 246–59 (2012)
C. Punyadeera, P.D. Slowey, in Nanobiomaterials in clinical dentistry, ed. by K. Subramani, W. Ahmed, J.K. Hartsfield (William Andrew Publishing, Waltham, 2013), pp. 453–473
R.J. Genco, Salivary diagnostic tests. J. Am. Dent. Assoc. 143, 3S–5S (2012)
H. Fabryova, P. Celec, On the origin and diagnostic use of salivary RNA. Oral Dis. 20, 146–52 (2013)
Y. Li, X. Zhou, M.A. St John, D.T. Wong, RNA profiling of cell-free saliva using microarray technology. J. Dent. Res. 83, 199–203 (2004)
N.J. Park, H. Zhou, D. Elashoff, B.S. Henson, D.A. Kastratovic, E. Abemayor, D.T. Wong, Salivary microRNA: discovery, characterization, and clinical utility for oral cancer detection. Clin. Cancer Res. 15, 5473–5477 (2009)
R.S. Patel, A. Jakymiw, B. Yao, B.A. Pauley, W.C. Carcamo, J. Katz, J.Q. Cheng, E.K. Chan, High resolution of microRNA signatures in human whole saliva. Arch. Oral Biol. 56, 1506–1513 (2011)
D.T. Wong, Salivaomics. J. Am. Dent. Assoc. 143(10 Suppl), 19S–24S (2012)
S. Babashah, M. Sadeghizadeh, M.R. Tavirani, S. Farivar, M. Soleimani, Aberrant microRNA expression and its implications in the pathogenesis of leukemias. Cell. Oncol. 35, 317–334 (2012)
E. Yiannakopoulou, Targeting epigentic mechanisms and microRNAs by aspirin and other non steroidal anti-inflammatory agents-implications for cancer treatment and chemoprevention. Cell. Oncol. 37, 167–178 (2014)
D.P. Bartel, MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297 (2004)
G.A. Calin, C.D. Dumitru, M. Shimizu, R. Bichi, S. Zupo, E. Noch, H. Aldler, S. Rattan, M. Keating, K. Rai, L. Rassenti, T. Kipps, M. Negrini, F. Bullrich, C.M. Croce, Frequent deletions and down-regulation of micro-RNA genes miR15 and miR16 at 13q14 in chronic lympocytic leukemmia. Proc. Natl. Acad. Sci. U. S. A. 99, 15524–15529 (2002)
D. Xiao, J. Ohlendorf, Y. Chen, D.D. Taylor, S.N. Rai, S. Waigel, W. Zacharias, H. Hao, K.M. McMasters, Identifying mRNA, microRNA and protein profiles of melanoma exosomes. PLoS One 7, e46874 (2012)
R. Mohamed, J.L. Campbell, J. Cooper-White, G. Dimeski, C. Punyadeera, The impact of saliva collection and processing methods on CRP, IgE, and Myoglobin immunoassays. Clin. Transl. Med. 1, 19 (2012)
Y. Choi, H.P. Dienes, K. Krawczynski, Kinetics of miR-122 expression in the liver during acute HCV infection. PLoS One 8, e76501 (2013)
S.C. Sahu, microRNAs in: Toxicology and Medicine, 1st edn. (Wiley, 2013), pg 486
S.A. Bustin, G.L. Shipley, J. Vandesompele, C.T. Wittwer, V. Benes, J.A. Garson, J. Hellemans, J. Huggett, M. Kubista, R. Mueller, T. Nolan, M.W. Pfaffl, The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin. Chem. 55, 611–622 (2009)
A.L. Carvalho, W.-W. Jiang, Q. Claybourne, Y. Tokumaru, J. Lee, D. Goldenberg, E. Garrett-Mayer, S. Goodman, C.-s. Moon, W. Koch, W.H. Westra, C. Jeronimo, D. Sidransky, J.A. Califano, M.M. Kim, R. Henrique, Z. Zhang, M.O. Hoque, S. Chang, M. Brait, C.S. Nayak, Evaluation of promoter hypermethylation detection in body fluid as a screening/diagnosis tool for head and neck squamous cell carcinoma. Clin. Cancer Res. 14, 97–107 (2008)
A.K. El-Naggar, K. Hurr, J.G. Batsakis, M.A. Luna, H. Goepfert, V. Huff, Sequential loss of heterozygosity at microsatellite motifs in preinvasive and invasive head and neck squamous carcinoma. Cancer Res. 55, 2656–2659 (1995)
M.F. Huang, Y.C. Chang, P.S. Liao, T.H. Huang, C.H. Tsay, M.Y. Chou, Loss of heterozygosity of p53 gene of oral cancer detected by exfoliative cytology. Oral Oncol. 35, 296–301 (1999)
A.K. El-Naggar, L. Mao, G. Staerkel, M.M. Coombes, S.L. Tucker, M.A. Luna, G.L. Clayman, S. Lippman, H. Goepfert, Genetic heterogeneity in saliva from patients with oral squamous carcinomas: implications in molecular diagnosis and screening. J. Mol. Diagn. 3, 164–170 (2001)
K.J. Livak, T.D. Schmittgen, Analysis of relative gene expression data using real-time quantitative PCR and the 2 (−Delta Delta C(T)) Method. Methods 25, 402–408 (2001)
L. Zhu, H. Chen, D. Zhou, D. Li, R. Bai, S. Zheng, W. Ge, MicroRNA-9 up-regulation is involved in colorectal cancer metastasis via promoting cell motility. Med. Oncol. 29, 1037–1043 (2012)
L. Ma, F. Reinhardt, E. Pan, J. Soutschek, B. Bhat, E.G. Marcusson, J. Teruya-Feldstein, G.W. Bell, R.A. Weinberg, Therapeutic silencing of miR-10b inhibits metastasis in a mouse mammary tumor model. Nat. Biotechnol. 28, 341–347 (2010)
A.B. Hui, A. Lin, W. Xu, L. Waldron, B. Perez-Ordonez, I. Weinreb, W. Shi, J. Bruce, S.H. Huang, B. O’Sullivan, J. Waldron, P. Gullane, J.C. Irish, K. Chan, F.F. Liu, Potentially prognostic miRNAs in HPV-associated oropharyngeal carcinoma. Clin. Cancer Res. 19, 2154–2162 (2013)
K.-i. Kozaki, I. Imoto, S. Mogi, K. Omura, J. Inazawa, Exploration of tumor-suppressive microRNAs silenced by DNA hypermethylation in oral cancer. Cancer Res. 68, 2094–2105 (2008)
Y. Khew-Goodall, G.J. Goodall, Myc-modulated miR-9 makes more metastases. Nat. Cell Biol. 12, 209–211 (2010)
S.C. Lin, W.G. Shen, C.J. Liu, MiR-134 expression is oncogenic for oral carcinoma. Oral Oncol. 47, S121–S121 (2011)
C.-J. Liu, W.G. Shen, S.-Y. Peng, H.-W. Cheng, S.-Y. Kao, S.-C. Lin, K.-W. Chang, miR-134 induces oncogenicity and metastasis in head and neck carcinoma through targeting WWOX gene. Int. J. Cancer 134, 811–821 (2014)
E. Elyakim, E. Sitbon, A. Faerman, S. Tabak, E. Montia, L. Belanis, A. Dov, E.G. Marcusson, C.F. Bennett, A. Chajut, D. Cohen, N. Yerushalmi, hsa-miR-191 is a candidate oncogene target for hepatocellular carcinoma therapy. Cancer Res. 70, 8077–8087 (2010)
S. Volinia, G.A. Calin, C.G. Liu, S. Ambs, A. Cimmino, F. Petrocca, R. Visone, M. Iorio, C. Roldo, M. Ferracin, R.L. Prueitt, N. Yanaihara, G. Lanza, A. Scarpa, A. Vecchione, M. Negrini, C.C. Harris, C.M. Croce, A microRNA expression signature of human solid tumors defines cancer gene targets. Proc. Natl. Acad. Sci. U. S. A. 103, 2257–2261 (2006)
C. Hebert, K. Norris, M.A. Scheper, N. Nikitakis, J.J. Sauk, High moility group A2 is a target for miRNA-98 in head and neck squamous cell carcinoma. Mol. Cancer 6, 5 (2007)
E.V. Barker, N.K. Cervigne, P.P. Reis, R.S. Goswami, W. Xu, I. Weinreb, J.C. Irish, S. Kamel-Reid, microRNA evaluation of unknown primary lesions in the head and neck. Mol. Cancer 8, 127 (2009)
Acknowledgments
The authors would like to acknowledge the financial support of the Queensland Government Smart Futures Fellowship Programme (QGSFF), Queensland Centre for Head and Neck cancer and University of Queensland Diamantina Institute Central Funds. In addition, the authors wish to acknowledge the on-going clinical support from the ENT Department at the Princess Alexandra Hospital in Woolloongabba, Australia. Special thanks goes to Ms Dana Middleton, A/Prof Chris Perry and A/Prof Ben Panizza.
Financial disclosure
None.
Conflicts of interest
None.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Table 1
(DOCX 21 kb)
Supplementary Table 2
(DOCX 19 kb)
Supplementary Table 3
(DOCX 29 kb)
Supplementary Table 4
(DOCX 12 kb)
Rights and permissions
About this article
Cite this article
Salazar, C., Nagadia, R., Pandit, P. et al. A novel saliva-based microRNA biomarker panel to detect head and neck cancers. Cell Oncol. 37, 331–338 (2014). https://doi.org/10.1007/s13402-014-0188-2
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13402-014-0188-2