Abstract
Gas-phase hydrogen/deuterium exchange measured by mass spectrometry (gas-phase HDX-MS) is a fast method to probe the conformation of protein ions. The use of gas-phase HDX-MS to investigate the structure and interactions of protein complexes is however mostly unharnessed. Ionizing proteins under conditions that maximize preservation of their native structure (native MS) enables the study of solution-like conformation for milliseconds after electrospray ionization (ESI), which enables the use of ND3-gas inside the mass spectrometer to rapidly deuterate heteroatom-bound non-amide hydrogens. Here, we explored the utility of gas-phase HDX-MS to examine protein-protein complexes and inform on their binding surface and the structural consequences of gas-phase dissociation. Protein complexes ranging from 24 kDa dimers to 395 kDa 24mers were analyzed by gas-phase HDX-MS with subsequent collision-induced dissociation (CID). The number of exchangeable sites involved in complex formation could, therefore, be estimated. For instance, dimers of cytochrome c or α-lactalbumin incorporated less deuterium/subunit than their unbound monomer counterparts, providing a measure of the number of heteroatom-bound side-chain hydrogens involved in complex formation. We furthermore studied if asymmetric charge-partitioning upon dissociation of protein complexes caused intermolecular H/D migration. In larger multimeric protein complexes, the dissociated monomer showed a significant increase in deuterium. This indicates that intermolecular H/D migration occurs as part of the asymmetric partitioning of charge during CID. We discuss several models that may explain this increase deuterium content and find that a model where only deuterium involved in migrating charge can account for most of the deuterium enrichment observed on the ejected monomer. In summary, the deuterium content of the ejected subunit can be used to estimate that of the intact complex with deviations observed for large complexes accounted for by charge migration.

ᅟ
Similar content being viewed by others
Introduction
The interactions of proteins dictate their biological activity, and the ability to characterize these on a molecular level is critical to understanding and deciphering the mechanisms of cellular processes, and to aid drug discovery and development [1,2,3,4]. Native MS has become a popular and powerful means to inform on the composition, stoichiometry, interaction, and subunit connectivity of protein complexes [5,6,7]. Using this methodology, proteins are ionized gently from solution, and the resulting native-like conformational states can be preserved in the gas-phase on a timescale ranging from milliseconds to seconds, enabling closer analysis of these conformations by MS [8,9,10,11]. Inside the mass spectrometer, gaseous biomolecules can, in addition to the mass measurement, be probed by an array of gas-phase techniques [12, 13]. These include ion mobility spectrometry (IMS) [9, 14, 15], and techniques that use gas-phase reactions such as proton-transfer reactions [16, 17], radical-based reactions [18, 19], or hydrogen/deuterium exchange (HDX).
Gas-phase HDX reactions inside a mass spectrometer have traditionally been performed by infusion of a deuterated gas, such as ND3, D2O, or MeOD, into selected parts of the mass spectrometer [20,21,22,23,24,25]. Heteroatom-bound hydrogens (hydrogens bound to O, N, or S) exchange with deuterium from the labeling gas over time and the resulting increase in mass can be measured using MS. To inform on conformational states pertinent to the solution phase, it is essential to complete the labeling reaction within tens of milliseconds after ionization [26, 27]. It has been shown that gas-phase HDX can be performed on a millisecond timescale shortly after ESI by use of ND3 to allow efficient labeling of heteroatom-bound side-chain hydrogens of native-like protein states [26,27,28,29]. Less reactive sites such as backbone amides do not exchange significantly in this time frame [24, 26, 30, 31], as the extent and rate of HDX depends on the proton affinity of the donor (ND3) and the acceptor site (-XH) [20, 30]. Gas-phase HDX-MS can detect and report on conformational differences present in solution, and has been used to study peptides, native and denatured proteins, protein-ligand complexes, peptide fibrillization, and oligosaccharides [26,27,28,29, 31,32,33]. Gas-phase HDX-MS thus reports on the surface chemistry as well as hydrogen bonding of side chains, and represents a complementary technique to other MS approaches, such as IMS [26, 29, 31, 33].
Here, we examine a series of protein complexes, in the range 24–395 kDa, to show that gas-phase HDX-MS in combination with native MS can be used to study multimeric protein-protein complexes, including cytochrome c and α-lactalbumin dimers, transthyretin tetramers, HSP16.9 12mers, and HSP16.5 24mers. Gas-phase labeling followed by subsequent collision-induced dissociation (CID) of the complexes made it possible to measure the deuterium content of both intact complex and dissociated subunits in the same experiment, and thereby approximate the number of heteroatom-bound non-amide hydrogens participating in the binding interfaces. In addition, upon analysis of larger multimeric protein complexes (12–24mers), a significant amount of intermolecular H/D migration was observed as part of the redistribution of charge during asymmetric partitioning, consistent with charge migration and the partial unfolding of the leaving subunit during activation.
Experimental
Materials
Deuterated ammonia gas (ND3-gas) was purchased from either AGA Gas (Pullach, Germany) in a lecture bottle containing 10 l 98% ND3-gas fitted with a Z14850 regulator (Sigma-Aldrich, Schnelldorf, Germany), or in a lecture bottle containing 10 l 99% ND3-gas purchased from CK Isotopes (Desford, UK) fitted with a CGA 180 CONCOA 422 series regulator (CK isotopes). HSP16.9 (wheat) was expressed and purified as described previously [34], and HSP16.5 (Methanocaldococcus jannaschii) was purified using the following method. In brief, the protein was expressed in BL21(DE3) cells for 3 h at 37 °C. Cells were lysed using a microfluidiser in 20 mM tris (pH 8), 50 mM NaCl, 0.1 mM EDTA. Protein was first purified by anion exchange chromatography performed by two 5 ml Hi-Trap Q columns (GE Healthcare) using 20 mM tris (pH 8) for protein binding and eluted with a linear gradient to 1 M NaCl. Fractions were pooled and concentrated to which an equal volume of 2 M ammonium sulphate was added before filtration. The filtrate was then loaded onto a 5 ml hydrophobic interaction chromatography (HIC) phenyl sepharose column (GE Healthcare) in 20 mM tris, 50 mM NaCl, 1 M ammonium sulphate (pH 8.0), and eluted using a linear gradient in the absence of ammonium sulphate. Final purification and buffer exchange were performed using size exclusion chromatography (SEC) on a Superdex 200 Increase 10/300 column (GE Healthcare) with a 200 mM ammonium acetate solution (pH 8).
All other samples and reagents were purchased from Sigma-Aldrich including lyophilized protein samples: Transthyretin/prealbumin (human plasma), α-lactalbumin (bovine milk), and cytochrome c (equine heart). The lyophilized proteins were unless stated otherwise, dissolved in LC-MS grade water and diluted into a suitable concentration in 200 mM ammonium acetate. Additional desalting steps (1–3) were performed utilizing Micro Bio-Spin 6 columns as per the manufacturer’s instructions (Bio-Rad Laboratories, Hercules, CA, USA). The Bio-Spin columns were, before use, buffer exchanged five times into 200 mM ammonium acetate in LC-MS grade water. Cytochrome c and α-lactalbumin dimer experiments were performed at 75–100 μM (dimer), and further diluted to 10 μM (monomer) for monomer control experiments. Transthyretin, HSP16.9, and HSP16.5 samples were prepared and analyzed at 0.5–5 μM (oligomer).
Native Mass Spectrometry of Protein-Protein Complexes
Protein samples were ionized in positive ionization mode from native-like conditions (200 mM ammonium acetate). Samples were prepared and loaded in borosilicate capillary emitters (either: Proxeon/Thermo or made in-house), and ionized using a nanoflow Z-spray ESI source (Waters, Wilmslow, UK) mounted on either a Synapt G1 or a Synapt G2 HDMS mass spectrometer (Waters, Wilmslow, UK) operating in TOF mode, according to published protocols [35]. The source and spray settings were tuned to obtain the best possible signal-to-noise and HDX efficiency: capillary 1.4 kV, sampling cone 10–120 V (120 V for in-source CID experiments), extraction cone 3 V, accelerating potential into the trap collision cell 8–40 V, and source block temperature 20 °C.
Gas-Phase HDX-MS and Gas-Phase Dissociation of Protein-Protein Complexes
Gas-phase HDX-MS was performed by leaking purified ND3-gas into a trap traveling wave ion guide (TWIG) of the mass spectrometer, and thus leading to labeling of the ions as a function of the ND3-gas pressure in the TWIG and the time the ions traversed the TWIG (governed to a large extent by the trap T-Wave velocity). Gas pressures, T-Wave heights, and T-Wave velocities of the TWIGs were adjusted for optimal ion transmission and labeling: trap ND3 pressure 12–14 μbar, trap T-Wave height 7.0 V, and trap T-Wave velocity 150 m/s (for experiments with α-lactalbumin, cytochrome c, and transthyretin). Alternatively, trap ND3 pressures of 22.2 μbar (HSP16.9) or 42 μbar (HSP16.5), with trap T-Wave height 0.1–0.2 V, and trap T-Wave velocity 200 m/s. By manually setting the gas regulator metering valve to obtain similar TWIG pressures of ND3, with an average deviation in the m/z of deuterated species at ~ 0.01%, between replicates. These minor differences in uptake probably reflect day-to-day thermal errors inherent to the Pirani gauge. We also note that all pressures reported in the mass spectrometer software are always “nitrogen-equivalent,” and conversion to absolute pressure values for ND3 would require a correction factor depending on the thermal conductivity of ND3 versus N2. The protein complexes were dissociated by post-source CID in the transfer TWIG to measure the labeling of individual components of the protein complex, at suitable activation energy for the complex. It would additionally be possible to perform gas-phase HDX in the transfer TWIG after CID in the trap TWIG, this was however not the focus of this study. Control experiments, in which no labeling occur, were performed with either Ar-gas or NH3-gas. For experiments with or without HDX, the gas pressure and T-Wave velocity setting are kept in the standard operating range for performing native mass spectrometry of protein complexes [9]. The T-Wave height setting is increased to ensure no ion “roll-over” is occurring in the T-Wave [36, 37]. See Figure S-1 for an overview of modifications performed on the mass spectrometers to enable gas-phase HDX-MS. For additional information regarding modifications on the low-pressure regions of the Synapt G1 and G2 instruments to enable gas-phase HDX-MS, we refer to previous papers [27, 29, 31, 32, 37, 38].
Data Analysis
Processing of mass spectra was carried out using MassLynx V4.1 software (Waters, Wilmslow, UK) with a suitable mean filter smoothing and subsequent Gaussian fit using an in-house developed script using Excel 2013 (Microsoft, Redmond, WA, USA). The deuterium content of the deuterium-labeled proteins and protein-protein complexes was determined by subtracting the apex of the fitted Gaussian peak (corresponding to the m/z), with the non-deuterated control sample in the absence of deuteration (either NH3- or Ar-gas) and multiplied by the charge state of the peak.
Proton Migration Model Estimations
To determine the fraction of deuteriums from the total number of labile sidechain heteroatom-bound hydrogens, the number of surface accessible labile side-chain sites had to be estimated. This was performed through the generation of the solvent-accessible surface through PyMOL visualization software (Schrödinger, NY, USA). The X-ray crystal structures used for determination were 1GME and 1SHS, respectively for HSP16.9 and HSP16.5. The N-terminal residues in each oligomer are buried in the center of the complex with only a small portion of the N-terminal structure visible. As such, structures were reduced to the α-crystallin domain and the C-terminal region, residues 44 and 43 onwards for HSP16.9 and HSP16.5, respectively. Spheres of 1 van der Waals radii were added to each of the side-chain hetero-atoms for the following residues: Lys, Arg, His, Tyr, Asp, Glu, Thr, Ser, and Cys. If the sphere was present at, or protruding from, the solvent-accessible surface, the site for that residue was deemed exposed and included in the calculation. Salt bridges were also determined using the polar contact feature in the PyMOL software with a distance cut-off set at 3.6 Å. If a salt-bridge interaction was present, the labile hydrogen count was reduced by 1. It should be noted that the true shielding/restrictive effect of a salt bridge may be greater than this and could result in up to three hydrogens being removed from the mobile pool. Our calculation does not account for protonated side chains solvated by the backbone carbonyl group picoseconds after ionization [10]. Backbone carbonyl group solvation in the gas-phase does not appear to render these hydrogens exchange incompatible, as peptides have shown to exchange most heteroatom-bound non-amide hydrogens during gas-phase HDX-MS [26].
Results and Discussion
Setup for Gas-Phase HDX-MS and CID
Gas-phase HDX-MS combined with CID were performed on a modified hybrid Q-TOF mass spectrometer (Waters Synapt), which allowed selective labeling of the fast-exchanging heteroatom-bound non-amide hydrogens of both intact protein complexes and their subunits (Figure 1a, b). Gas-phase HDX was performed in the trap TWIG which normally functions as a collision cell. By exchanging the standard Ar-gas to ND3-gas, it was possible to perform gas-phase HDX on a millisecond timescale as ions were pushed through the ion guide by the traveling wave. The degree of labeling could be controlled by varying either the partial pressure of the ND3-gas or the wave velocity in the trap TWIG, which to a large extent governs the ions residence time in the TWIG cell (0.1–10 ms) [26, 39]. By labeling in the trap TWIG with prior quadrupole selection, the possibility of subsequent complex dissociation with post-source CID in the transfer TWIG (referred to as CID) was enabled (Figure 1b). Experiments were performed by infusing ND3-gas (labeled sample), NH3-gas, or Ar-gas (reference/unlabeled sample) to the Trap TWIG, Figure 1c, d, respectively. The difference in the measured m/z of each ion from the labeled and the unlabeled sample could be used to calculate its deuterium content. No significant difference was observed for the relative distribution of monomer to dimer, charge state envelope, or adduct formation, between NH3-gas and ND3-gas in the trap TWIG.
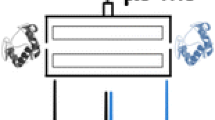
Gas-phase HDX-MS for studying protein-protein complexes. (a) Heteroatom-bound non-amide hydrogens present on side chains (red) are labeled by fast gas-phase HDX-MS using ND3-gas. (b) Schematic of the mass spectrometer setup for performing gas-phase HDX-MS with subsequent complex dissociation by CID on a Q-TOF mass spectrometer equipped with traveling wave ion guides (TWIG). Selected charge states of the complexes were isolated in the quadrupole, and gas-phase HDX-MS labeling was performed downstream in the trap TWIG by infusing purified ND3-gas into the chamber. Lastly, before mass analysis, dissociation of the complex was performed in the transfer TWIG by CID. Mass spectra are shown for the protein cytochrome c as full scan, quadrupole-selected complex, and dissociated complex without (c) and with HDX (d). An increase in mass as observed for the HDX experiments is a result of H/D exchange in the protein. The difference in m/z of labeled and unlabeled samples was used to calculate deuterium uptake for both precursor multimer and dissociated subunits, as illustrated by the mass spectra of cytochrome c
Gas-Phase HDX-MS and CID of Protein-Protein Dimers: Symmetrical or Asymmetrical Charge Partitioning
As a first step, we sought to investigate the ability of gas-phase HDX-MS in combination with CID to provide conformational information on protein-protein dimer complexes and their binding interface. The proteins cytochrome c (CYTC) and α-lactalbumin (ALAC) have a similar mass (12 kDa, 14 kDa), and form non-specific dimers at high concentration [40, 41]. Gas-phase HDX-MS and CID were performed on the 9+ dimer of each protein. The 9+ dimer of ALAC dissociated primarily symmetrically into monomers of 5+/4+ (Figures 2a and S-2a), whereas the 9+ dimer of CYTC dissociated both symmetrically into 5+/4+ monomers or more asymmetrically into 6+/3+ monomers (Figures 2b and S-2b). Gas-phase HDX-MS was performed on the different dimers in the trap TWIG with increasing collision energy in the transfer TWIG, ranging 10–60 V for 9+ ALAC and 10–90 V for 9+ CYTC. At the lowest energy settings (10 V), the proteins were present primarily as dimers (> 95%), and dissociation into monomers increased as a function of collision energy. At the highest energy settings, little dimer was observed in the mass spectrum (< 0.5% dimer). The increase in m/z upon labeling by gas-phase HDX-MS is shown for ALAC (Figure 2c) and CYTC (Figure 2d).
Symmetric or asymmetric charge partitioning of noncovalent protein-protein dimers by gas-phase HDX-MS and CID. Mass spectra of α-lactalbumin (ALAC) (a) or cytochrome c (CYTC) (b) dimers. The quadrupole-selected 9+ dimer underwent gas-phase HDX-MS labeling in trap TWIG followed by dissociation in the transfer TWIG. Crystal structures of dimeric ALAC (PDB: 1HFY) and CYTC (PDB: 1BBH) are additionally shown in (a) and (b), respectively. Mass spectra of 9+ and dissociated monomers are shown for ALAC (c) and CYTC (d), as black and colored for without and with gas-phase HDX-MS, respectively. Deuterium content per subunit is shown for the 9+ dimer (gray) and dissociated monomers of different charge states originating from the dimer (various colors), for ALAC (e) and CYTC (f). The deuterium content is shown as ribbon graphs where the midpoint and width correspond to the average deuterium uptake and 1× SD, respectively. The experiments were performed in triplicate
The measured deuterium contents for dimer and the dissociated monomers are shown for ALAC (Figure 2e) and CYTC (Figure 2f) as deuterium content per subunit, for experiments performed at different collision energy settings in the transfer TWIG. Dimers of ALAC and CYTC showed a deuterium content of 35.0–39.8 (17.5–19.9 D/subunit) and 51.4–55.2 D (25.7–27.6 D/subunit), respectively. Interestingly, for dimers of both proteins, the dissociated monomers contained approximately half the deuterium content of the precursor dimer (Figure 2e, f), with 16.8–20.1 D/subunit (ALAC) and 25.1–28.8 D/subunit (CYTC). This indicated that the labeling was retained equally in each component of the complex upon dissociation, irrespective of whether dissociation gave rise to symmetric (4+/5+) or asymmetric partitioning (i.e., 3+/6+) of charge. Similar results were obtained for 11+ ALAC and 11+ CYTC, which similarly dissociated mostly symmetrically (5+/6+) and more asymmetrically (4+/7+). Comparable deuterium content was measured for dissociated monomers of different charge. Thus, asymmetric charge-partitioning upon CID does not appear to incur measurable intermolecular H/D migration for the small homodimers studied. Interestingly, in all monomer pairs, the deuterium content was higher for the monomer with the lowest charge.
Gas-phase HDX-MS was also performed on unbound monomers of these proteins, which showed a deuterium content of 27.5 D for 5+ ALAC monomer and 28.7 D for CYTC monomer. This showed that the deuterium content of monomers dissociated from protein dimers was lower compared to unbound monomers (e.g., by approx. 9 D for 5+ ALAC). This finding indicates that complex formation can protect sites from HDX and thus gas-phase HDX-MS can be used to study the binding interface of such protein-protein complexes. We note that the difference in deuterium uptake upon complex formation could be explained partially by other factors, such as differences in the collision cross sections (CCS) or changed inertia of the larger dimers. The 5+ ALAC monomer exchanged ~ 9 D less upon complex formation, whereas the 5+ CYTC only show a smaller ~ 3 D decrease in deuterium uptake upon dimerization. This can be explained by a difference in the number of heteroatom-bound non-amide hydrogens partaking in the binding interface, as was previously shown for protein-ligand complexes [28]. Alternatively, ALAC could adopt a more compact conformation upon complex formation than CYTC, resulting in fewer surface-accessible sites as has been indicated in earlier work employing IMS and gas-phase HDX using D2O on a longer (s) timescale [42]. No significant difference in the measured deuterium content was observed for either the dimer or dissociated monomers at increasing dissociation energy for CYTC, which indicates that the collision energy does not significantly alter the gas-phase HDX-MS results and only affects the ratio of dimer:monomer present in the mass spectrum. ALAC, however, showed a smaller increase in deuterium content upon collision activation.
Gas-Phase HDX-MS and CID to Inform on Protein-Protein Multimers
Transthyretin (TTR) is a 56 kDa homotetrameric protein formed as a dimer of dimers, with a prominent β-sheet structure symmetry [43]. TTR was ionized from native conditions and was observed primarily as a tetramer of charge states 13–15+. Gas-phase dissociation of the 15+ tetramer by CID (40 V) resulted in asymmetric partitioning of charge per subunit. CID resulted primarily in monomers (6–9+) and trimers (6–9+), Figure 3a. Performing the experiments in the presence of ND3-gas instead of Ar-gas decreased the peak widths, indicating a higher amount of solvent and salt adducts present for the experiments performed with ND3-gas. Reference/unlabeled experiments were therefore performed with NH3-gas in the trap TWIG to allow accurate measurement of deuterium uptake during gas-phase HDX-MS. No difference between ND3-gas and NH3-gas was observed in the ratio of monomer:trimer, charge state distribution, or salt- and solvent adduct formation. NH3-gas was therefore found to be a suitable reference gas for the unlabeled control measurements during gas-phase HDX-MS analysis of larger noncovalent protein-protein complexes (Figure S-3). The deuterium content was measured at 27.6 D for the intact tetrameric complex, which corresponded to 6.9 D per subunit. A comparable deuterium content was measured for the dissociated monomers (6.8–8.0 D/subunit) and trimers (5.9–7.5 D/subunit) of different charge, Figure 3c. Dissociated TTR species (monomers and trimers) generally showed a small increase in deuterium content (~ 0.4 D/subunit) per charge. Furthermore, dissociated monomers showed a small average increase in deuterium of 0.5 D/subunit relative to dissociated trimers of the same charge. Together, this could indicate a minor degree of intermolecular H/D migration during dissociation of the protein complex by CID. Overall, deuterium content/subunit was similar in both intact complex and dissociated subunits upon dissociation suggesting that even dissociation with a highly asymmetric partitioning of charge does not cause a large redistribution of deuterium (due to intermolecular H/D migration) between subunits. It, therefore, appears feasible to accurately dissect the labeling of multimeric protein-protein complexes by gas-phase HDX-MS and subsequent CID and thus gain information on both the intact complex and individual subunits.
Dissociation of tetrameric transthyretin studied by gas-phase HDX-MS. (a) Mass spectrum of quadrupole-selected 15+ tetramer of transthyretin following gas-phase HDX-MS and complex dissociation (magnified tenfold for m/z > 2500). Zoom-ins are of mass spectra of selected peaks, without (black) and with (colored) gas-phase HDX-MS (b) mass spectrum of transthyretin with activation in the source (170 V) before gas-phase HDX-MS and without quadrupole selection. Zoom-ins are of mass spectra of selected peaks, without (black) and with (colored) gas-phase HDX-MS. Deuterium content per subunit is shown for precursor 15+ tetramer (dashed gray line) and dissociated monomers (orange) and trimers (blue) of different charge state, in experiments performed with gas-phase HDX-MS performed before (c) or after collisional activation (d). Experiments were done in at least three replicates, and the error bars denote the 1× standard deviation
For comparison, TTR was also dissociated before gas-phase HDX (CID-HDX). The TTR tetramer was activated in the source by increasing the cone voltage to 120 V, which gave rise to a mass spectrum with tetramers (10–16+), trimers (5–9+), and monomers (5–10+) of different charge state (Figure 3b). The deuterium content of the intact tetramer (16.0 D/subunit), monomer (19.8–44.0 D/subunit), and trimer (3.0–3.3 D/subunit) of the same charge states as for the HDX-CID experiments were calculated, Figure 3d. The deuterium content per subunit observed for all TTR species during this CID-HDX experiment was significantly different from the deuterium content per subunit of corresponding TTR species of the same charge measured in the prior conventional experiments where CID was performed after HDX (HDX-CID). Specifically, the intact tetramer and monomers showed an increased deuterium content, and trimers showed a decreased deuterium content. Compared to HDX-CID experiments, the unbound monomers of TTR showed a much larger increase in deuterium content with charge state, exchanging 4.8 D (5+), 9.5 D (6+), 19.8 D (7+), 39.2 D (8+), and 44.1 D (9+) (Figure 3d). It is therefore likely that the small increase in deuterium content for monomers of increasing charge observed in the HDX-CID experiments is not caused by differences in the exchange rate of the individual subunits, but rather a small degree of intermolecular H/D migration upon CID. It is, however, difficult to compare the deuterium content of ions of different charge state and relate this to structural or conformational differences. Increasing the number of charges can influence several factors, apart from the structure, that could impact deuterium exchange kinetics including (I) increased internal energy of the ion during ionization and transmission, (II) unfolding or changes in collision cross-section due to increased Coulumbic repulsion, or (III) a different ability to form exchange-competent ion-ND3 complexes. We therefore only directly compare proteins or protein complexes of similar charge and subunit count. As described earlier, the deuterium content of both the tetrameric complex and dissociated products were similar for TTR ionized under soft conditions (Figure 3c), though a small degree of intermolecular H/D migration was observed for asymmetric dissociation. Labeling by gas-phase HDX-MS can thus inform on the folding of individual subunits of protein-protein complexes, and the labeling of individual subunits can be dissected by gas-phase dissociation. This enables information to be gained for the conformation and chemical surface composition of the intact protein complex and subunits using the same workflow.
Larger Protein-Protein Multimers
Small heat-shock proteins (sHSP) are expressed in organisms across all kingdoms of life. They are diverse molecular chaperones that are important to a variety of biological processes, including the cell cycle, cell differentiation, cell death, and adaptation to stressful conditions [44]. The sHSP are characterized by the presence of the highly conserved α-crystallin domain and are typically composed of low molecular mass monomers (15–40 kDa). These monomers assemble into larger multimeric complexes that can contain as many as 50 subunits [45]. These proteins have proven to be excellent systems for studying the mechanism of protein complex dissociation in the gas phase owing to their significant asymmetric charge partitioning [46, 47]. Here, we have explored the gas-phase HDX-MS and CID methodology to study HSP16.9, which forms a homomeric 12mer (201 kDa) [48], as well as HSP16.5, a 395 kDa sHSP 24mer complex [49].
Collision-induced dissociation of HSP16.9 gave rise to the highly asymmetric partitioning of charge through the sequential ejection of monomeric units. Initially, at collision voltages of 40–140 V, only singly stripped species were generated with the unfolded monomer chain leaving with more than half of the precursor charge. Further activation (160–180 V) resulted in the additional loss of a monomeric unit from the already stripped, N-1, species taking almost half of the remaining charge. For gas-phase labeling experiments, the 32+ intact 12mer was isolated in the quadrupole and activated (160 V) to give both N-1 and N-2 species centred around the 18+ and 10+ charge state, respectively (Figure 4a). The observed monomers, therefore, originated from both the intact 12mer and the N-1 precursors with the charge states following a bimodal distribution centred on 13+ and 9+, with the higher charge state series coming from the first dissociation event (12mer). HSP16.5, 24mer, was also studied owing to its ability to dissociate monomers asymmetrically over two sequential dissociation events. The intact 48+ 24mer was quadrupole isolated, and the CID spectra were recorded over 70–180 V of activation. At 70 V, HSP16.5 dissociated into N-1 and N-2 stripped oligomers with a significant amount of intact precursor still populated (~30%), in addition to the corresponding monomers (Figure 4b). The partitioning of charge was less asymmetric than for HSP16.9 with the N-1 and N-2 monomers leaving with around a quarter and a third of the total charge, respectively. The monomers arose from all dissociation events and resulted in similarily charged species leading to a single charge state distribution for stripped monomers of HSP16.5.
Gas-phase HDX and subsequent CID of HSP16.9 (12mer) and HSP16.5 (24mer). Mass spectra of quadrupole-selected, gas-phase labeled, and collisionally activated HSP16.9 32+ (a) and HSP16.5 48+ (b). The spectral intensity has been magnified fivefold for m/z > 6000). The most abundant charge state for each of the dissociation products is shown in the presence of ND3 (blue) relative to the NH3 control (black). The average deuterium content per subunit is shown for each of the dissociation products, for HSP16.9 (c) and HSP16.5 (d). Symmetric partitioning of the intact uptake (dashed gray lines) is displayed for reference. The amount of D enrichment on the ejected monomer unit is shown for each observed charge state
To determine how deuterium from gas-phase HDX labeling partitioned on such asymmetric CID systems, HSP16.9 and HSP16.5 were labeled as described for other complexes following activation in the range 50–180 V (HDX-CID), Figure S-4a,b. The most abundant charge state was highlighted, and the shift in m/z upon gas-phase labeling is shown in Figure 4a, b. The monomeric region provides high-quality data with little deviation between repeats, in part due to the instrument sensitivity over the m/z range but also as the stripped monomers are less prone to influence from salt adduct shifts while more variation in peak shape and the determined centroid were seen for the higher m/z species (Figures S-4c, d and S-5).
The uptake for HSP16.9 and HSP16.5 was determined for each dissociation product (Figure 4a, b) and was averaged over all charges states across three repeats. The uptake for intact HSP16.9 was 138 D or 11.5 D per subunit and was determined from the spectrum at a collision voltage (80 V) where the intact precursor remained with high signal-to-noise. This uptake value also represents the theoretical uptake of the dissociated monomers should the deuterium content of the complex partition perfectly symmetrically (i.e., the ejected subunit leaves with 1/N of the total deuterium uptake of the intact complex). The average deuterium content per subunit for the N-2 oligomer, N-1 oligomer, and the monomeric species was found to be 9, 11.5, and 14 D/subunit (Figure 4c). The deuterium content for symmetric partitioning of D is shown as a reference in dashed gray lines. The deuterium content was similarly determined for HSP16.5 with the intact 24mer incorporating 12 D per subunit (288 D total), the monomers 15 D/subunit, N-1 oligomers 11 D/subunit, and the N-2 oligomers 10.5 D/subunit (Figure 4d). The amount of deuterium on the monomeric species in each case was higher than if uptake was partitioned symmetrically, the difference between these two values was then defined as the monomeric enrichment. The enrichment is shown in Figure 4c, d for each monomer charge state and loosely increases with charge.
Deuterium Enrichment of Highly Asymmetric Dissociated Subunits from Protein Complexes
For both HSP16.9 and HSP16.5, the dissociated monomers were ejected with more deuterium (D) than if each subunit left with a symmetric share. The result was therefore that the monomer became slightly enriched in deuterium and the stripped oligomers become slightly depleted. One likely explanation lies in the assymetric redistribution of the net charge, with respect to mass. The charge per subunit was significantly lower for the intact complex compared to the stripped monomers. For example, the average charge localized to each subunit in the 48+ 24mer of HSP16.5 was 2+, and the average charge state of dissociated monomers was 13+, representing an enrichment of charge of 11+ (Δz = 11). One likely method of charge enrichment during CID is the net migration of surface protons moving from the folded oligomer to the monomer, during unfolding and ejection of the monomer from the oligomer (Figure 5a) [50]. When the complex is labeled, a proportion of the mobile charge will be D+. The fraction of deuterium from the pool of labile heteroatom-bound non-amide hydrogens on the surface of the intact complex will determine how much D enrichment occurs as charge migrates. If HSP16.5 was fully saturated with deuterium, the D enrichment would simply be the number of charges that have migrated, Δz = 11. In this work, the surface was not fully exchanged as ND3 pressures were kept low to prevent ammonia adducts from forming inside the spectrometer. The D enrichment is then the difference in charge, Δz, scaled by the fraction of deuterons from the pool of all heteroatom-bound non-amide hydrogens available (FD), Eq. 1. This assumes that the rate of proton or deuteron site “hopping” is comparable at these energies and that the flux of charge only occurs toward the ejecting subunit.
Migration of deuterium upon dissociation of protein complexes. (a) Schematic representing different dissociation mechanisms and how they may result in differing degrees of enrichment on the ejected subunit. A folded homo-oligomer will have symmetric deuterium uptake across each subunit, highlighted in light blue. Each site is represented by a block either occupied by hydrogen (blue) or deuterium (red), demonstrated for 50% deuterium saturation. Dissociation through salt bridge rearrangement (SaBRe) or charge mediated unfolding will result in different enrichment levels, with the latter dependent on the fraction of deuterium in the system (b). Greater enrichment may occur if complete scrambling occur across all sites, including newly exposed sites from unfolding. Charge is shown as white +/− signs and a net migration of 2+ is shown in each case. (b) A protein subunit is represented as a number of heteroatom-bound sites (squares) originating from different regions of the protein structure: surface accessible and buried non-amide hydrogens, and amide hydrogens associated with the backbone. Upon activation, a greater number of H/Ds may become mobile depending from where they originate. Illustrated are four cases; where only surface accessible side-chain H/Ds can migrate but not including hydrogens “trapped” in salt bridges (SA-SB), the case where only labile surface side-chain H/Ds can migrate (SA), where all labile Hs in the protein sequence are involved (seq.), and finally when all labile Hs from side chains and the backbone amides can migrate (Seq. + BB). Progressing from SA-SB to Seq. + BB significantly decreases the fraction of D in the pool of migrating H/Ds as many exchange incompetent hydrogens are introduced to the system. The different models featured in (a, b) are shown for comparison with the observed monomer enrichment observed for HSP16.9 (c) and HSP16.5 (d) and discussed further in the text: Seq. + BB (orange), Seq. (blue), SA (red), and SA-SB (purple)
To estimate the degree of enrichment expected in these experiments, several scenarios were considered to provide an upper and lower bound (Figure 5b, Tables S-1 and S-2. Summing all the heteroatom-bound hydrogens associated with the side chains and backbone from the sequence provides an upper limit of the number of mobile Hs in the system, making FD equal to 0.04 and 0.05 (Seq. + BB model), for HSP16.9 and HSP16.5, respectively (based on the measured uptake of 138 D and 288 D for the two intact proteins). If we then assume that amide hydrogens are not sufficiently mobile at the energies supplied for complex dissociation they can be removed from the calculation, the FD increases to 0.09 and 0.12 (Seq. model). The observed enrichment lies above these lower enrichment limits (Seq. and Seq. + BB models Figure 5c, d) for both HSP16.9 and HSP16.5. This suggests that assuming all heteroatom-bound non-amide hydrogens, including those on the interior of the protein or at interfaces, are mobile is a poor approximation.
A more realistic model includes only labile Hs on the surface of the protein (surface accessible (SA) model); this was estimated for both proteins from the atomic coordinates of each protein and gives an FD of 0.24 for both HSP16.9 and HSP16.5. The observed enrichment for HSP16.5 broadly falls within both this surface-accessible model (SA) and the sequence model (Seq.) (Figure 5d). The deuterium enrichment levels for HSP16.9 do not quite fall within these limits. The surface-accessible model predicts an FD of 0.24 while a linear fit of the observed data would suggest an FD of 0.36 (Figure 5c). While this is greater than expected, the enrichment remains well below the maximum allowed, FD = 1. This would suggest that fewer Hs are mobile than the crystal structure would predict, implying that a greater number of residues are either buried in the gas-phase or involved in strong interactions preventing migration away from that site. Other factors that could affect H/D migration are differences in proton affinity of the sites, and solvation of side chains with carbonyl groups of the backbone. The number of salt bridges present in the crystal structure was determined and for each ion pairing interaction, the proton count reduced accordingly, surface accessible minus salt bridge (SA-SB) model. A total of 3 and 10 salt-bridge interactions were estimated for HSP16.9 and HSP16.5, respectively, giving FD values of 0.25 and 0.30.
One other scenario to consider is that subunits do not become asymmetrically charged due to charge migration, but instead interfacial salt bridges are asymmetrically broken in such a way that the ejected subunit leaves with more charge and the potential to subsequently unfold (SaBRe model) [51]. For a labeled system, HDX at the interface of the complex would be unlikely, and so this mechanism would not provide any means of subunit deuterium enrichment, instead resulting in perfectly symmetric D partitioning (i.e., no monomeric enrichment) (Figure 5a). This is inconsistent with the data for HSP16.9 and HSP16.5. However, it is conceivable that both proton migration and the SaBRe mechanisms operate in tandem which would act to decrease the value of Δz in Eq. 1.
An upper limit to the amount of deuterium enrichment can be imagined if migration occurs in addition to the net charge movement (Figure 5a). In this scenario, complete scrambling occurs across all possible labile sites on the complex surface with H/D flux both toward and from the unfolded, yet attached, subunit. As the ejecting subunit unfolds, it will reveal additional unlabeled, labile residues resulting in an excess of sites on the unfolded subunit. If complete equilibration of deuterium occurs over all sites, this will manifest in a deuterium enrichment of the ejected monomer. For HSP16.9, the unfolded subunit contains 123 non-amide labile H sites, and the remaining oligomer has 473, therefore after complete H/D migration, the monomer will be left with 20.6% of the total uptake, rather than 8.3% for symmetric dissociation. This means the monomer would be enriched by 12.3% of the total uptake, 17 D. Our observed enrichment falls well below this level (not shown in Figure 5c, d) suggesting complete H/D migration is not occurring and/or the ejected subunit is not entirely unfolded.
While the migration of surface charge, through the surface accessible and salt bridge models, does not completely explain the observed data for HSP 16.9, the models do provide a means of explaining the large majority of enrichment with a relatively simple scenario. Other contributing factors may certainly be at play such as perturbation of structure in the gas phase and the scale and impact of salt bridge and electrostatic interactions have on hindering H/D migration. Using a model where H/D flux is only toward the unfolding subunit appears valid as it accounts for most of the observed enrichment levels.
Conclusions
Here, we have shown that performing native MS and employing gas-phase HDX-MS in combination with CID can be used to study the conformational properties of multimeric protein complexes. We analyzed protein assemblies ranging from dimers to 24mers and showed that it was possible to label these by infusion of ND3-gas into the trap T-wave ion guide of the mass spectrometer. Dissociation downstream of gas-phase HDX allowed for the deuterium content to be determined at the subunit level and thus could be compared to that of subunits ionized from solution in an unbound state. This both reported on the exposed surface of free monomers and the intact complex, allowing inference of the protection from HDX provided by the interface. This gas-phase HDX-MS and CID methodology provided information on (a) the conformation of the individual subunits and the complex, (b) the size of the binding interface of the complex by approximation of heteroatom-bound non-amide hydrogens participating in the binding interface, and (c) provided information on the gas-phase dissociation of protein complexes.
Dimers of α-lactalbumin and cytochrome c were studied by gas-phase HDX-MS and dissociated at increasing CID energies. The deuterium content per subunit was comparable (± 2 D) for intact complex and dissociated monomers at various charge states. Increasing the activation energy for dissociation of the complexes produced more dissociated monomers to intact dimer, but only affected the deuterium content to a smaller degree (< 10%). This indicates that gas-phase HDX-MS of protein complexes can be performed at different activation energies for optimal dissociation of the complex. The dimers of α-lactalbumin and cytochrome c both additionally showed reduced gas-phase deuterium uptake compared to unbound monomers of the proteins. This indicated that the number of exchangeable heteroatom-bound non-amide hydrogens decreased upon dimerization, possibly due to the formation of hydrogen bonding at the binding interface and changes in the higher-order conformation of the proteins. Interestingly, α-lactalbumin showed a larger decrease (9 D) in HDX upon dimerization compared to cytochrome c (3 D). This could indicate that dimerization of α-lactalbumin results in a more compact formation or a larger binding interface, than for cytochrome c.
The gas-phase HDX-MS and CID methodology was also used to study larger multimeric protein complexes. Tetramers of transthyretin were first studied, which dissociated asymmetrically in terms of charge and mass into monomer and trimers. The monomers contained a higher amount of charges per subunit (6–9+) compared to trimers (2–3+). Comparable deuterium content per subunit were observed for both intact precursor and monomers and trimers originating from dissociation. Dissociated monomers showed a small increase in deuterium content (0.6 D) to dissociated trimers. Additionally, dissociated monomers and trimers of higher charge states both showed a limited increase in deuterium content (0.4–1.6 D), compared to lower charged species. Due to the relatively small migration of charges Δz = 3.75, only a small intermolecular H/D migration was expected, which is in line with the < 1 D enrichment of the dissociated monomers observed here. This indicated limited or localized intermolecular H/D migration upon asymmetric gas-phase dissociation of protein complexes, with increasing asymmetric charge migration leading to increasing H/D migration. This hypothesis was tested further on the larger 12mer HSP16.9 and 24mer HSP16.5 protein complexes which both dissociated very asymmetrically in terms of mass and charge. For dissociation of these large protein complexes, limited intermolecular H/D migration was clearly apparent. Dissociated monomers showed a relatively higher deuterium content than the intact complexes and the dissociated N-1 and N-2 complexes. The dissociated monomers additionally showed a small increase in deuterium content as a function of their charge state.
Intermolecular H/D Migration upon CID
The generally accepted explanation for asymmetric dissociation by CID is the electrostatic charge partitioning model, where the single dissociated subunit unravels and undergoes proton migration prior to ejection from the residual complex [50, 52,53,54]. The ejected subunit is therefore unfolded and carries a large percentage excess of charges, where the remaining subunits in the complex contain somewhat fewer charges.
Based on the electrostatic charge partitioning model, four different models were proposed for quantifying the amount of potential intermolecular migration of deuterium for dissociation of protein complexes that might be expected. (i) The sequence model (Seq. + BB), (ii) the sequence model not including backbone amide hydrogens (Seq.), (iii) the surface-accessible residue model (SA), and (iv) the surface-accessible model that accounts for salt-bridge interactions (SA-SB). For HSP16.5, the measured deuterium enrichment could best be explained by either the SA, SA-SB, or Seq. models. For HSP16.9, the measured deuterium enrichment was higher than the proposed methods, but the SA and SA-SB models showed close correlations to the experimental values. Of the four proposed models, the SA and SA-SB models seemed to be most consistent with deuterium enrichment measured.
An alternative and more recent model for explaining the dissociation of protein complexes is the salt bridge rearrangement (SaBRe) model that proposes that rearrangement and breaking of salt bridges at the protein complex interface are responsible for the asymmetric dissociation behavior [51]. Gas-phase labeling would not occur on the binding interface of protein complexes, as hydrogen is shielded from HDX due to either hydrogen bonding and/or being buried in the binding interface. No intermolecular enrichment of deuterium is therefore expected according to the SaBRe model upon complex dissociation as any charged sites at the interface will therefore not likely contain any deuterium. A clear indication of intermolecular H/D migration was observed for TTR tetramer and the larger sHSP 12mer and 24mer protein complexes. From these data, it is therefore not likely that the SaBRe model alone is responsible for dissociation of noncovalent protein complexes. However, it cannot be excluded that salt bridge rearrangements play a part in the dissociation of protein complexes.
The results presented here indicate that limited intermolecular movement of H/D is occurring upon CID of noncovalent protein complexes, with the deviation from the symmetric partitioning of deuterium depending heavily on the charge asymmetry. The amount of migration, however, is far below the level expected if all labile sites were sampled by the mobile H/D pool over the time of gas-phase dissociation event. These results are similar to previous studies on smaller peptide dimers labeled with deuterium at backbone amides sites in solution, which showed a smaller degree (5–10%) of intermolecular H/D migration upon CID [55]. In contrast, H/D levels at both backbone amides and side-chain sites are well known to migrate/scramble completely on an intramolecular level for both peptides and proteins upon CID [27, 56,57,58].
Our results demonstrate that gas-phase HDX-MS with CID can be a powerful tool to provide structural information on protein-protein complexes and mechanistic insight into their dissociation. The method can probe heterogeneous sample mixtures and proteins in a dynamic equilibrium, where solution HDX-MS and other biophysical techniques may not have adequate resolution or separation. Gas-phase HDX-MS allows analyses to be done by using only small amount of material, low picomole, and at low micromolar concentration requiring only little sample preparation (buffer exchange into an ESI compatible buffer that retains the protein in a native-like solution). Here, we show that the deuterium content of the gas-phase dissociated monomer can be used as a proxy for the intact uptake, as only limited intermolecular H/D migration is occurring upon CID even in cases of very asymmetric charge partitioning (Figure 6). This is of significant value for systems where the intact precursor cannot be cleanly isolated or carries significant adducts which make deuterium uptake determination difficult, and for which time-aligned experiments exploiting IMS is advantageous [59]. Further insight and development of the charge migration models used here could allow for a correction to be applied to intermolecular H/D migration upon complex dissociation. This could facilitate the further study of constituent parts of a heterogenous protein-protein complex within the mass spectrometer.
Deuterium levels in native protein complexes following gas-phase HDX are partitioned in a highly symmetrical manner during CID. The fraction of monomer uptake (Umonomer) relative to intact complex uptake (Uintact) is plotted against the inverse of the number of subunits (N) in the intact complex. For perfectly symmetric D partitioning points would fall on the diagonal dotted line
Change history
11 February 2019
In this issue, the citation information on the opening page of each article PDF is incorrect. It should read ?Journal of the American Society of Mass Spectrometry (2019)?,? not ?Journal of the American Society of Mass Spectrometry (2018)...?
11 February 2019
In this issue, the citation information on the opening page of each article PDF is incorrect. It should read ���Journal of the American Society of Mass Spectrometry (2019)���,��� not ���Journal of the American Society of Mass Spectrometry (2018)...���
References
Teilum, K., Olsen, J.G., Kragelund, B.B.: Functional aspects of protein flexibility. Cell. Mol. Life Sci. 66, 2231–2247 (2009)
Russel, D., Lasker, K., Phillips, J., Schneidman-Duhovny, D., Velázquez-Muriel, J.A., Sali, A.: The structural dynamics of macromolecular processes. Curr. Opin. Cell Biol. 21, 97–108 (2009)
Robinson, C.V., Sali, A., Baumeister, W.: The molecular sociology of the cell. Nature. 450, 973–982 (2007)
Frokjaer, S., Otzen, D.E.: Protein drug stability: a formulation challenge. Nat. Rev. Drug Discov. 4, 298–306 (2005)
Konijnenberg, A., Butterer, A., Sobott, F.: Native ion mobility-mass spectrometry and related methods in structural biology. Biochim. Biophys. Acta - Proteins Proteomics. 1834, 1239–1256 (2013)
Mehmood, S., Allison, T.M., Robinson, C.V.: Mass spectrometry of protein complexes: from origins to applications. Annu. Rev. Phys. Chem. 66, 453–474 (2015)
Loo, J.A.: Studying noncovalent protein complexes by electrospray ionization mass spectrometry. Mass Spectrom. Rev. 16, 1–23 (1997)
Sobott, F., Hernández, H., McCammon, M.G., Tito, M.A., Robinson, C.V.: A tandem mass spectrometer for improved transmission and analysis of large macromolecular assemblies. Anal. Chem. 74, 1402–1407 (2002)
Ruotolo, B.T., Benesch, J.L.P., Sandercock, A.M., Hyung, S.J., Robinson, C.V.: Ion mobility-mass spectrometry analysis of large protein complexes. Nat. Protoc. 3, 1139–1152 (2008)
Breuker, K., McLafferty, F.W.: Stepwise evolution of protein native structure with electrospray into the gas phase, 10−12 to 102 s. Proc. Natl. Acad. Sci. U. S. A. 105, 18145–18152 (2008)
Badman, E.R., Hoaglund-Hyzer, C.S., Clemmer, D.E.: Monitoring structural changes of proteins in an ion trap over ~10–200 ms: unfolding transitions in cytochrome c ions. Anal. Chem. 73, 6000–6007 (2001)
Green, M.K., Lebrilla, C.B.: Ion-molecule reactions as probes of gas-phase structures of peptides and proteins. Mass Spectrom. Rev. 16, 53–71 (1997)
Chandler, S.A., Benesch, J.L.: Mass spectrometry beyond the native state. Curr. Opin. Chem. Biol. 42, 130–137 (2018)
Bohrer, B.C., Mererbloom, S.I., Koeniger, S.L., Hilderbrand, A.E., Clemmer, D.E.: Biomolecule analysis by ion mobility spectrometry. Annu. Rev. Anal. Chem. 1, 293–297 (2008)
Jurneczko, E., Barran, P.E.: How useful is ion mobility mass spectrometry for structural biology? The relationship between protein crystal structures and their collision cross sections in the gas phase. Analyst. 136, 20–28 (2011)
Stephenson, J.L., McLuckey, S.A.: Ion/ion reactions in the gas phase: proton transfer reactions involving multiply-charged proteins. J. Am. Soc. Mass Spectrom. 118, 7390–7397 (1996)
Gross, D.S., Schnier, P.D., Rodriguez-Cruz, S.E., Clifton, K., Williams, E.R.: Conformations and folding of lysozyme ions in vacuo. Proc. Natl. Acad. Sci. U. S. A. 93, 3143–3148 (1996)
Cooper, H.J., Hakansson, K., Marshall, A.G.: The role of electron capture dissociation in biomolecular analysis. Mass Spectrom. Rev. 24, 201–222 (2005)
Zubarev, R.A., Horn, D.M., Fridriksson, E.K., Kelleher, N.L., Kruger, N.A., Lewis, M.A., Carpenter, B.K., McLafferty, F.W.: Electron capture dissociation for structural characterization of multiply charged protein cations. Anal. Chem. 72, 563–573 (2000)
Cheng, X., Fenselau, C.: Hydrogen/deuterium exchange of mass-selected peptide ions with ND3 in a tandem sector mass spectrometer. Int. J. Mass Spectrom. 122, 109–119 (1992)
Winger, B.E., Light-Wahl, K.J., Rockwood, A.L., Smith, R.D.: Probing qualitative conformation differences of multiply protonated gas-phase proteins via H/D isotopic exchange with D2O. J. Am. Chem. Soc. 114, 5897–5898 (1992)
Suckau, D., Shi, Y., Beu, S.C., Senko, M.W., Quinn, J.P., Wampler III, F.M., McLafferty, F.W.: Coexisting stable conformations of gaseous protein ions. Proc. Natl. Acad. Sci. U. S. A. 90, 790–793 (1993)
Heck, A.J.R., Jørgensen, T.J.D., O’Sullivan, M., von Raumer, M., Derrick, P.J.: Gas-phase noncovalent interactions between vancomycin-group antibiotics and bacterial cell-wall precursor peptides probed by hydrogen/deuterium exchange. J. Am. Soc. Mass Spectrom. 9, 1255–1266 (1998)
Pan, J., Heath, B.L., Jockusch, R.A., Konermann, L.: Structural interrogation of electrosprayed peptide ions by gas-phase H/D exchange and electron capture dissociation mass spectrometry. Anal. Chem. 84, 373–378 (2012)
Khakinejad, M., Kondalaji, S.G., Maleki, H., Arndt, J.R., Donohoe, G.C., Valentine, S.J.: Combining ion mobility spectrometry with hydrogen-deuterium exchange and top-down MS for peptide ion structure analysis. J. Am. Soc. Mass Spectrom. 25, 2103–2115 (2014)
Rand, K.D., Pringle, S.D., Murphy III, J.P., Fadgen, K.E., Brown, J., Engen, J.R.: Gas-phase hydrogen/deuterium exchange in a traveling wave ion guide for the examination of protein conformations. Anal. Chem. 81, 10019–10028 (2009)
Rand, K.D., Pringle, S.D., Morris, M., Brown, J.M.: Site-specific analysis of gas-phase hydrogen/deuterium exchange of peptides and proteins by electron transfer dissociation. Anal. Chem. 84, 1931–1940 (2012)
Mistarz, U.H., Brown, J.M., Haselmann, K.F., Rand, K.D.: Probing the binding interfaces of protein complexes using gas-phase H/D exchange mass spectrometry. Structure. 24, 310–318 (2016)
Beeston, H.S., Ault, J.R., Pringle, S.D., Brown, J.M., Ashcroft, A.E.: Changes in protein structure monitored by use of gas-phase hydrogen/deuterium exchange. Proteomics. 15, 2842–2850 (2015)
Campbell, S., Rodgers, M.T., Marzluff, E.M., Beauchamp, J.L.: Deuterium exchange reactions as a probe of biomolecule structure. Fundamental studies of gas phase H/D exchange reactions of protonated glycine oligomers with D2O, CD3OD, CD3CO2D, and ND3. J. Am. Chem. Soc. 117, 12840–12854 (1995)
Mistarz, U.H., Brown, J.M., Haselmann, K.F., Rand, K.D.: Simple setup for gas-phase H/D exchange mass spectrometry coupled to electron transfer dissociation and ion mobility for analysis of polypeptide structure on a liquid chromatographic time scale. Anal. Chem. 86, 11868–11876 (2014)
Uppal, S.S., Beasley, S.E., Scian, M., Guttman, M.: Gas-phase hydrogen/deuterium exchange for distinguishing isomeric carbohydrate ions. Anal. Chem. 89, 4737–4742 (2017)
Przygońska, K., Poznański, J., Mistarz, U.H., Rand, K.D., Dadlez, M.: Side-chain moieties from the N-terminal region of Aβ are involved in an oligomer-stabilizing network of interactions. PLoS One. 13, e0201761 (2018)
Lee, G.J., Pokala, N., Vierling, E.: Structure and in vitro molecular chaperone activity of cytosolic small heat shock proteins from pea. J. Biol. Chem. 270, 10432–10438 (1995)
Kondrat, F.D.L., Struwe, W.B., Benesch, J.L.P.: Native mass spectrometry: towards high-throughput structural proteomics. In: Owens, R.J. (ed.) Structural Proteomics, pp. 349–371. Springer, New York City (2014)
Giles, K., Pringle, S.D., Worthington, K.R., Little, D., Wildgoose, J.L., Bateman, R.H.: Applications of a travelling wave-based radio-frequency-only stacked ring ion guide. Rapid Commun. Mass Spectrom. 18, 2401–2414 (2004)
Mistarz, U.H., Rand, K.D.: Installation, validation, and application examples of two instrumental setups for gas-phase HDX-MS analysis of peptides and proteins. Methods. 144, 113–124 (2018)
Rand, K.D., Zehl, M., Jensen, O.N., Jørgensen, T.J.D.: Protein hydrogen exchange measured at single-residue resolution by Electron transfer dissociation mass spectrometry. Anal. Chem. 81, 5577–5584 (2009)
Pringle, S.D., Giles, K., Wildgoose, J.L., Williams, J.P., Slade, S.E., Thalassinos, K., Bateman, R.H., Bowers, M.T., Scrivens, J.H.: An investigation of the mobility separation of some peptide and protein ions using a new hybrid quadrupole/travelling wave IMS/oa-ToF instrument. Int. J. Mass Spectrom. 261, 1–12 (2007)
Jurchen, J.C., Williams, E.R.: Origin of asymmetric charge partitioning in the dissociation of gas-phase protein homodimers. J. Am. Chem. Soc. 125, 2817–2826 (2003)
Jurchen, J.C., Garcia, D.E., Williams, E.R.: Further studies on the origins of asymmetric charge partitioning in protein homodimers. J. Am. Soc. Mass Spectrom. 15, 1408–1415 (2004)
Wright, P.J., Douglas, D.J.: Gas-phase H/D exchange and collision cross sections of hemoglobin monomers, dimers, and tetramers. J. Am. Soc. Mass Spectrom. 20, 484–495 (2009)
Blake, C.C.F., Geisow, M.J., Oatley, S.J., Rérat, B., Rérat, C.: Structure of prealbumin: secondary, tertiary and quaternary interactions determined by Fourier refinement at 1.8 Å. J. Mol. Biol. 121, 339–356 (1978)
Carra, S., Alberti, S., Arrigo, P.A., Benesch, J.L., Benjamin, I.J., Boelens, W., Bartelt-Kirbach, B., Brundel, B.J.J.M., Buchner, J., Bukau, B., Carver, J.A., Ecroyd, H., Emanuelsson, C., Finet, S., Golenhofen, N., Goloubinoff, P., Gusev, N., Haslbeck, M., Hightower, L.E., Kampinga, H.H., Klevit, R.E., Liberek, K., Mchaourab, H.S., McMenimen, K.A., Poletti, A., Quinlan, R., Strelkov, S.V., Toth, M.E., Vierling, E., Tanguay, R.M.: The growing world of small heat shock proteins: from structure to functions. Cell Stress Chaperones. 22, 601–611 (2017)
Hilton, G.R., Lioe, H., Stengel, F., Baldwin, A.J., Benesch, J.L.P.: Small heat-shock proteins: paramedics of the cell. In: Jackson, S. (ed.) Molecular Chaperones, pp. 69–98. Springer Berlin Heidelberg, Berlin, Heidelberg (2013)
Benesch, J.L.P., Aquilina, J.A., Ruotolo, B.T., Sobott, F., Robinson, C.V.: Tandem mass spectrometry reveals the quaternary organization of macromolecular assemblies. Chem. Biol. 13, 597–605 (2006)
Benesch, J.L.P., Ruotolo, B.T., Sobott, F., Wildgoose, J., Gilbert, A., Bateman, R., Robinson, C.V.: Quadrupole-time-of-flight mass spectrometer modified for higher-energy dissociation reduces protein assemblies to peptide fragments. Anal. Chem. 81, 1270–1274 (2009)
van Montfort, R.L., Basha, E., Friedrich, K.L., Slingsby, C., Vierling, E.: Crystal structure and assembly of a eukaryotic small heat shock protein. Nat. Struct. Biol. 8, 1025–1030 (2001)
Kim, K.K., Kim, R., Kim, S.H.: Crystal structure of a small heat-shock protein. Nature. 394, 595–599 (1998)
Popa, V., Trecroce, D.A., McAllister, R.G., Konermann, L.: Collision-induced dissociation of electrosprayed protein complexes: an all-atom molecular dynamics model with mobile protons. J. Phys. Chem. B. 120, 5114–5124 (2016)
Loo, R.R.O., Loo, J.A.: Salt bridge rearrangement (SaBRe) explains the dissociation behavior of noncovalent complexes. J. Am. Soc. Mass Spectrom. 27, 975–990 (2016)
Light-Wahl, K.J., Schwartz, B.L., Smith, R.D.: Observation of the noncovalent quaternary associations of proteins by electrospray ionization mass spectrometry. J. Am. Chem. Soc. 116, 5271–5278 (1994)
Sinelnikov, I., Kitova, E.N., Klassen, J.S.: Influence of Coulombic repulsion on the dissociation pathways and energetics of multiprotein complexes in the gas phase. J. Am. Soc. Mass Spectrom. 18, 617–631 (2007)
Felitsyn, N., Kitova, E.N., Klassen, J.S.: Thermal decomposition of a gaseous multiprotein complex studied by blackbody infrared radiative dissociation. Investigating the origin of the asymmetric dissociation behavior. Anal. Chem. 73, 4647–4661 (2001)
Modzel, M., Stefanowicz, P., Szewczuk, Z.: Hydrogen scrambling in non-covalent complexes of peptides. Rapid Commun. Mass Spectrom. 26, 2739–2744 (2012)
Demmers, J.A.A., Rijkers, D.T.S., Haverkamp, J., Killian, J.A., Heck, A.J.R.: Factors affecting gas-phase deuterium scrambling in peptide ions and their implications for protein structure determination. J. Am. Chem. Soc. 124, 11191–11198 (2002)
Jørgensen, T.J.D., Gårdsvoll, H., Ploug, M., Roepstorff, P.: Intramolecular migration of amide hydrogens in protonated peptides upon collisional activation. J. Am. Chem. Soc. 127, 2785–2793 (2005)
Mistarz, U.H., Bellina, B., Jensen, P.F., Brown, J.M., Barran, P.E., Rand, K.D.: UV Photodissociation mass spectrometry accurately localize sites of backbone deuteration in peptides. Anal. Chem. 90, 1077–1080 (2018)
Shepherd, D.A., Marty, M.T., Giles, K., Baldwin, A.J., Benesch, J.L.P.: Combining tandem mass spectrometry with ion mobility separation to determine the architecture of polydisperse proteins. Int. J. Mass Spectrom. 377, 663–671 (2015)
Acknowledgements
K.D.R. acknowledges the generous financial support from the Danish Council for Independent Research | Natural Sciences (Steno Grant no. 11-104,058). U.H.M. gratefully acknowledges the COST Action BM1403 for an STSM grant and the Danish Ministry of Higher Education and Science for an Elite Research (EliteForsk) travel grant. S.A.C. and J.L.P.B. thank the Biotechnology and Biological Sciences Research Council (BB/L017067/1) and Waters Corp. for an iCASE studentship.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
ESM 1
(PDF 594 kb)
Rights and permissions
About this article
Cite this article
Mistarz, U.H., Chandler, S.A., Brown, J.M. et al. Probing the Dissociation of Protein Complexes by Means of Gas-Phase H/D Exchange Mass Spectrometry. J. Am. Soc. Mass Spectrom. 30, 45–57 (2019). https://doi.org/10.1007/s13361-018-2064-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13361-018-2064-1