Abstract
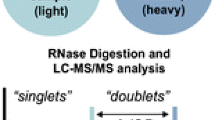
Mapping, sequencing, and quantifying individual noncoding ribonucleic acids (ncRNAs), including post-transcriptionally modified nucleosides, by mass spectrometry is a challenge that often requires rigorous sample preparation prior to analysis. Previously, we have described a simplified method for the comparative analysis of RNA digests (CARD) that is applicable to relatively complex mixtures of ncRNAs. In the CARD approach for transfer RNA (tRNA) analysis, two complete sets of digestion products from total tRNA are compared using the enzymatic incorporation of 16O/18O isotopic labels. This approach allows one to rapidly screen total tRNAs from gene deletion mutants or comparatively sequence total tRNA from two related bacterial organisms. However, data analysis can be challenging because of convoluted mass spectra arising from the natural 13C and 15 N isotopes present in the ribonuclease-digested tRNA samples. Here, we demonstrate that culturing in 12C-enriched/13C-depleted media significantly reduces the isotope patterns that must be interpreted during the CARD experiment. Improvements in data quality yield a 35 % improvement in detection of tRNA digestion products that can be uniquely assigned to particular tRNAs. These mass spectral improvements lead to a significant reduction in data processing attributable to the ease of spectral identification of labeled digestion products and will enable improvements in the relative quantification of modified RNAs by the 16O/18O differential labeling approach.

ᅟ





Similar content being viewed by others
References
Phizicky, E.M., Hopper, A.K.: tRNA biology charges to the front. Genes Dev. 24, 1832–1860 (2010)
Novoa, E.M., Ribas de Pouplana, L.: Speeding with control: codon usage, tRNAs, and ribosomes. Trends Genet. 28, 574–581 (2012)
Gollnick, P., Babitzke, P.: Transcription attenuation. Biochem. Biophys. Acta 1577, 240–250 (2002)
Dittmar, K.A., Mobley, E.M., Radek, A.J., Pan, T.: Exploring the regulation of tRNA distribution on the genomic scale. J. Mol. Biol. 337, 31–47 (2004)
Wohlgemuth, S.E., Gorochowski, T.E., Robous, J.A.: Translational sensitivity of the Escherichia coli genome to fluctuating tRNA availability. Nucleic Acids Res. 41, 8021–8033 (2013)
Hershberg, R., Petrov, D.A.: Selection on codon bias. Annu. Rev. Genet. 42, 287–299 (2008)
Cantara, W.A., Crain, P.F., Rozenski, J., McCloskey, J.A., Harris, K.A., Zhang, X., Vendeix, F.A.P., Fabris, D., Agris, P.F.: The RNA modification database, RNAMDB: 2011 update. Nucleic Acids Res. 39, D195–D201 (2011)
Machnicka, M., Milanowska, K., Osman, O., Purta, E., Kurkowska, M., Olchowik, A., Januszewski, W., Kalinowski, S., Dunin-Horkawicz, S., Rother, K., Helm, M., Bujnicki, J., Grosjean, H. MODOMICS: a database of RNA modification pathways—2013 update. Nucleic Acids Res. D262–267 (2012)
Graeber, M.B., Muller, U.: Recent developments in the molecular genetics of mitochondrial disorders. J. Neurol. Sci. 153, 251–263 (1998)
Saikia, M., Fu, Y., Pavon-Eternod, M., He, C., Pan, T.: Genome-wide analysis of N1-methyl-adenosine modification in human tRNAs. RNA 16, 1317–1327 (2010)
Maynard, N.D., Macklin, D.N., Kirkegaard, K., Covert, M.W.: Competing pathways control host resistance to virus via tRNA modification and programmed ribosomal frameshifting. Mol. Syst. Biol. 8, 567 (2012)
Florentz, C.: Molecular investigations on tRNAs involved in human mitochondrial disorders. Biosci. Rep. 22, 81–98 (2002)
Gebetsberger, J., Zywicki, M., Kunzi, A., Polacek, N.: tRNA-derived fragments target the ribosome and function as regulatory non-coding RNA in Haloferax volcanii. Archaea 2012, 11 (2012)
Gingold, H., Dahan, O., Pilpel, Y.: Dynamic changes in translational efficiency are deduced from codon usage of the transcriptome. Nucleic Acids Res. 40, 10053–10063 (2012)
Chang, D.-E., Smalley, D., Conway, T.: Gene expression profiling of Escherichia coli growth transitions: an expanded stringent response model. Mol. Microbiol. 45, 289–306 (2003)
Moukadiri, I., Garzon, M.J., Bjork, G.R., Armengod, M.E.: The output of the tRNA modification pathways controlled by the Escherichia coli MnmEG and MnmC enzymes depends on the growth conditions and the tRNA species. Nucleic Acids Res. 42, 2602–2623 (2014)
Chan, C.T., Dyavaiah, M., DeMott, M.S., Taghizadeh, K., Dedon, P.C., Begley, T.J.: A quantitative systems approach reveals dynamic control of tRNA modifications during cellular stress. PLoS Genet. 6, e1001247 (2010)
Dumelin, C., Chen, Y., Leconte, A., Chen, Y., Liu, D.: Discovery and biological characterization of geranylated RNA in bacteria. Nat. Chem. Biol. 8, 913–919 (2012)
Pavon-Eternod, M., Gomes, S., Geslain, R., Dai, Q., Rosner, M.R., Pan, T.: tRNA over-expression in breast cancer and functional consequences. Nucleic Acids Res. 37, 7268–7280 (2009)
Dong, H., Nilsson, L., Kurland, C.G.: Co-variation of tRNA abundance and codon usage in Escherichia coli at different growth rates. J. Mol. Biol. 260, 649–663 (1996)
Liu C, B.T., Hou YM.: Fluorophore labeling to monitor tRNA dynamics. Methods Enzymol. 469, 69–93 (2009)
Yokogawa, T., Kumazawa, Y., Miura, K., Watanabe, K.: Purification and characterization of two serine isoacceptor tRNAs from bovine mitochondria by using a hybridization assay method. Nucleic Acids Res. 17, 2623–2638 (1989)
Mir, K.U., Southern, E.M.: Determining the influence of structure on hybridization using oligonucleotide arrays. Nat. Biotechnol. 17, 788–792 (1999)
McCloskey, J.A., Nishimura, S.: Modified nucleosides in transfer RNA. Acc. Chem. Res. 10, 403–409 (1977)
Kowalak, J.A., Pomerantz, S.C., Crain, P.F., McCloskey, J.A.: A novel method for the determination of post-transcriptional modification in RNA by mass spectrometry. Nucleic Acids Res. 21, 4577–4585 (1993)
Douthwaite, S., Kirpekar, F.: Identifying modifications in RNA by MALDI mass spectrometry. Methods Enzymol. 425, 1–20 (2007)
Huang, T.Y., Liu, J., McLuckey, S.A.: Top-down tandem mass spectrometry of tRNA via ion trap collision-induced dissociation. J. Am. Soc. Mass Spectrom. 21, 890–898 (2010)
Matthiesen, R., Kirpekar, F.: Identification of RNA molecules by specific enzyme digestion and mass spectrometry: software for and implementation of RNA mass mapping. Nucleic Acids Res. 37, e48 (2009)
Suzuki, T., Ikeuchi, Y., Noma, A., Sakaguchi, Y.: Mass spectrometric identification and characterization of RNA-modifying enzymes. Methods Enzymol. 425, 211–229 (2007)
Taucher, M., Breuker, K.: Characterization of modified RNA by top-down mass spectrometry. Angew. Chem. Int. Ed. Engl. 51, 11289–11292 (2012)
Buvoli, A., Buvoli, M., Leinwand, L.A.: Enhanced detection of tRNA isoacceptors by combinatorial oligonucleotide hybridization. RNA 6, 912–918 (2000)
Hossain, M., Limbach, P.A.: Mass spectrometry-based detection of transfer RNAs by their signature endonuclease digestion products. RNA 13, 295–303 (2007)
Wetzel, C., Limbach, P.: The global identification of tRNA isoacceptors by targeted tandem mass spectrometry. Analyst 138, 6063–6072 (2013)
Li, S., Limbach, P.: Method for Comparative Analysis of Ribonucleic Acids Using Isotope Labeling and Mass Spectrometry. Anal. Chem. 84, 8607–8613 (2012)
Castleberry, C.M., Limbach, P.: Relative quantitation of transfer RNAs using liquid chromatography mass spectrometry and signature digestion products. Nucleic Acids Res. 38, e162 (2010)
Wetzel, C., Limbach, P.: Global identification of transfer RNAs by liquid chromatography-mass spectrometry (LC-MS). J. Proteome 75, 3450–3464 (2011)
Li, S., Limbach, P.: Mass spectrometry sequencing of transfer ribonucleic acids by the comparative analysis of RNA digests (CARD) approach. Analyst 138, 1386–1394 (2013)
Meng, Z., Limbach, P.: Quantitation of Ribonucleic Acids using 18O labeling and mass spectrometry. Anal. Chem. 77, 1891–1895 (2005)
Castleberry, C.M., Lilleness, K., Baldauff, R., Limbach, P.A.: Minimizing 18O/16O back-exchange in the relative quantification of ribonucleic acids. J. Mass Spectrom. 44, 1195–1202 (2009)
Murakami, S., Fujishima, K., Tomita, M., Kanai, A.: Metatranscriptomic analysis of microbes in an oceanfront deep-subsurface hot spring reveals novel small RNAs and type-specific tRNA degradation. Appl. Environ. Microbiol. 78, 1015–1022 (2012)
Acknowledgments
Financial support of this work was provided by the National Science Foundation (CHE1212625) and a University of Cincinnati Department of Chemistry Doctoral Enhancement Award to C.W.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 137 kb)
Rights and permissions
About this article
Cite this article
Wetzel, C., Li, S. & Limbach, P.A. Metabolic De-Isotoping for Improved LC-MS Characterization of Modified RNAs. J. Am. Soc. Mass Spectrom. 25, 1114–1123 (2014). https://doi.org/10.1007/s13361-014-0889-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13361-014-0889-9