Abstract
Sorghum grain traits are important agronomic traits directly related to yield and are key factors affecting the brewing process of distill liquor. Exploring the genes controlling those traits is of great significance for understanding the genetic mechanism of sorghum grain development. In this study, we conducted genotyping using Super-GBS technology on a recombinant inbred lines (RILs) population derived from the cross between “BTx623” and “Hongyingzi,” consisting of 205 lines. The grain-related traits of the RIL population were investigated in Guiyang, Anshun in Guizhou, and Ledong in Hainan in China. By inclusive composite interval mapping (ICIM) method, a total of 47 quantitative trait locus (QTL) related to four grain traits (thousand grain weight, grain length, grain width, and length–width ratio) were identified across 10 chromosomes. Among them, 20 important QTL were repeatedly detected in multiple traits or environments and distributed on chromosomes 1 (1), 2 (2), 3 (5), 4 (5), 5 (1), 6 (2), 7 (2), 8 (1), and 9 (1). Six candidate genes were identified within the confidence interval of these QTL, and they are homologous to genes controlling rice grain development (OsMADS1, RGG2, OsNST1, SMG1, OsGRF8, and OsAP2-39). The results provide a basis for further cloning and functional verification of these candidate genes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Sorghum (Sorghum bicolor (L.)Moench, 2n = 2x = 20) is the world’s fifth-largest cereal crop and has a long history of cultivation in China (Ding et al. 2019; Morris et al. 2013; Lu 1999). It is recorded that sorghum has been utilized for brewing liquor in China since the Yuan Dynasty (1270–1368CE) (Shinoda 1958; Zhang et al. 2023a, b). Currently, sorghum is mainly grown in the southwest, north, and northeast regions of China, with over 80% of sorghum grains used to produce distill liquor (Chen et al. 2019; Li et al. 2021; Shinoda 1958; Tang and Xie 2019). In the southwest region, almost all sorghum grains are predominantly utilized for liquor brewing, with a historical emphasis on the production of renowned Maotai-flavored liquor. Sorghum grain traits are not only closely related to yield but also play a critical role in the brewing process (Gao et al. 2022), which comprises over 30 steps and requires small and round grains to obtain good resistance to cooking and stirring (Feng et al. 2022). The increasing scale of Maotai-flavored liquor production in recent years has led to a growing demand for sorghum cultivation. However, there are issues such as severe degradation and low yield in sorghum varieties for liquor, which affect the supply of raw materials for liquor production (Ding et al. 2020). Therefore, QTL mapping and gene cloning for grain related traits, as well as developing tightly linked molecular markers, are significant for improving the efficiency of breeding high-yielding sorghum varieties for liquor brewing purposes.
Grain-related traits are complex quantitative traits influenced by both genotype and environment (Baye et al. 2022; Tao et al. 2018). With the continuous improvement of sorghum genome, and the development of high-throughput sequencing technology, numerous QTL controlling grain related traits have been identified in sorghum (Han and Cai 2019). Currently, utilizing parental populations and natural populations, 29 studies have reported 186 QTLs affecting grain related traits on the 10 chromosomes of sorghum (aussorgm.org.au). Among these, grain weight has received the most attention as it is one of the three essential components contributing to yield, with over 80% of the QTLs related to thousand grain weight (TGW). However, there have been few reports on QTL for other grain related traits, with only 18 QTL detected for grain length (GL), 4 for grain width (GW), 4 for grain length–width ratio (GLWR), and 8 for grain roundness (GR). QTL for GL and GW are located on all chromosomes, except for chromosome 8, whereas QTL for GLWR and GR are found on chromosomes 1, 3, 6, 7 and 9 (Guindo et al. 2019; Liu et al. 2019; Zhang et al. 2015).
Despite numerous studies on QTL mapping of sorghum grain-related traits, there is a few report of fine mapping or gene cloning of QTL, which lags behind the research in major cereal crops such as rice, maize, and wheat (Boyles et al. 2016; Han et al. 2015; Tao et al. 2020; Zou et al. 2020). Han et al. finely mapped of a major QTL (qGW1) for GW within a range of approximately 101 kb on the short arm of chromosome 1, and Sobic.001G038300 was preliminarily determined as the candidate gene for qGW1 (Han et al. 2015). Boyles et al. discovered a novel QTL for TGW in the 5 M interval of chromosome 5, with the peak marker located on the gene Sobic.005G188400 encoding an unknown function remorid protein, which is mainly expressed in early floral tissues and developing embryos (Boyles et al. 2016). Based on resequencing of a RIL population of 244 lines, Zuo et al. detected a major QTL (qTGW1a) on chromosome 1 and employed map-based cloning to identify Sobic.001G341700 as the candidate gene, which is homologous to the gene GS3 controlling grain size in rice. Functional analysis revealed that qTGW1a acts as a negative regulator of grain size in sorghum (Zou et al. 2020). Using genome-wide association analysis (GWAS) and nested association mapping, Tao et al. identified an important QTL (qFHGS2.5) on chromosome 2. They further determined the candidate gene (Sobic.002G216600) which is homologous to DEP1 in rice, a positive regulator of grain number related to grain weight (Tao et al. 2020).
To date, the sorghum accessions used for QTL mapping and GWAS analysis mainly originate from foreign resources (https://aussorgm.org.au/sorghum-qtlatlas/). However, significant differences exist between Chinese sorghum and foreign sorghum, and there may be new important QTL loci controlling grain traits. Therefore, in this study, utilizing the high-density linkage map, QTL related to sorghum grain traits was detected in the RIL constructed from Chinese and foreign sorghum (Ding et al. 2023), and candidate genes for the significant QTL were identified, which would provide a basis for further gene cloning, functional analysis, and molecular breeding of sorghum grain traits.
Materials and methods
Plant materials
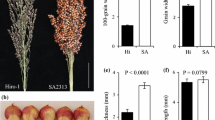
A set of 205 RILs derived from a cross between sorghum inbred lines BTx623 and Hongyingzi (HYZ) was developed and used in this study. There were significant differences in panicle type and grain traits between parents. The Inbred line BTx623 had a compact panicle with large and round seeds, and HYZ had a large and loose panicle with small and oval seeds. Each RIL was derived from a signal F2 plant by single seed descent until the F12 generation.
In field trials, the 205 lines and their parents were planted for investigation of grain traits in Guiyang in 2020 (20GY) and 2021(21GY), Guizhou Province [26.67°N, 106.62°E, altitude 1175 m]; in 2021 in Anshun (21AS), Guizhou Province [25.21°N, 105.13°E, altitude 1280 m]; and in 2021 in Ledong (21HN), Hainan Province [18.74°N, 109.17°E, altitude 28 m]. Planting seasons in different environments (sowing in April and harvesting in August in Guizhou Province, sowing in November and harvesting in March the following year in Hainan Province) have the following total rainfall and average temperature for 20GY, 21GY, 21AS, and 21HN respectively: 933.94 mm/20.84 °C, 1089.44 mm/20.58 °C, 918.04 mm/22.35 °C, and 155.6 mm/19.59 °C (for specific information, please see Table S1). We fertilized 750 kg compound fertilizer (N + P2O5 + K2O ≥ 45%) per hectare in the field before planting Sorghum. The ratio of nitrogen, phosphorus, and potassium is 15–15-15 in the compound fertilizer.
Phenotyping
Two hundreds grams of with intact and undamaged seeds were selected to measure the thousand grain weight (TGW), grain length (GL), grain width (GW), and length–width ratio (GLW), using the Wanshen SCG automatic seed analyzer (Hangzhou Wanshen Testing Technology Co., Ltd.). TGW is recorded in grams, and the measurements are considered valid if the difference between three weight measurements is less than 0.1 g (within 5%), and then the average value is then taken. GL and GW are measured in millimeters, with each line repeated three times, and the average value is calculated. The GLW is obtained by dividing GL by GW (for specific information, please see Table S2). Descriptive statistical analyses, including mean, standard deviation, and frequency distribution, as well as correlation analyses, were conducted on the grain traits across different environments using SPSS 21.0 software. Graphs were generated using R4.03 software.
QTL mapping and candidate gene prediction
Based on the constructed genetic linkage map (Ding et al. 2023), QTL analysis was performed using the inclusive composite interval mapping (ICIM) method in QTL Ici-Mapping 4.2 software. The parameters used were a stepwise regression probability of P < 0.001, chromosome step length of 0.1 cM, and the logarithm of the odds (LOD) threshold of 2.5. QTL which were named according to the “QTL + trait + chromosome + order on the chromosome” principle. Previously reported sorghum QTL information was searched on the Sorghum QTL Atlas (http://aussorgm.org.au/sorghum-qtl-atlas/). Candidate genes in QTL interval were searched in the BTx623 reference genome, and their putative functions were annotated in the available databases: phytozome (https://phytozome-next.jgi.doe.gov/) and rice (https:// www. riced ata. cn/ gene/).
RNA extraction and transcriptome expression analysis
Differences of samples were collected from the parents BTx623 and HYZ at four different stages of panicle/seed development (5 days after flowering in young panicles, 10 days after flowering in seeds, 15 days after flowering in seeds, 20 days after flowering in seeds). Total RNA was extracted from the samples. The NEBNextⓇ UltraTM RNA Library Prep Kit (New England BioLabs, USA) was used for transcriptome library construction, and the libraries that passed quality control were sequenced on the NovaSeq 6000 platform (Illumina, USA). After sequencing, the Filter-FQ software was used to remove adapters and low-quality sequences. The clean data were then aligned to the reference genome BTx623 using the HISAT2 software. The expression levels of genes were quantified using the cufflinks software, which calculated the gene FPKM (Fragments Per Kilobase of transcript per Million mapped reads) values. Three biological replicates were performed to calculate the mean ± SD expression levels for each sample.
Results
Phenotypic variation of the parents and RILs
From 2020 to 2021, the phenotype of the parents and RILs were surveyed in four environments, including Guiyang and Anshun, as well as Ledong (Table 1). The mean TGW, GL, GW, and GLWR of HYZ were 21.61 g, 4.11 mm, 3.35 mm, and 1.23, and the respective values of BTx623 were 24.77 g, 4.08 mm, 3.67 mm and 1.11. The distributions of four traits in the RIL population were continuous with coefficients of variation ranging from 0.04% to 0.54%, indicating the quantitative inheritance of these characteristics. The average values of these traits fell within the range of the two parental phenotypic values, and both the minimum and maximum values exceeded the parental phenotypic values. In addition, except for GLWR, the skewness and kurtosis absolute values of the RIL population’s TGW, GL, and GW were all less than 1, indicating that these traits followed a normal distribution.
In order to understand the relationship between various traits under different environments, Pearson correlation analysis was conducted for four traits (Table 2). The results showed that TGW was significantly negatively correlated with GLWR in all four environments (P < 0.01), with a range of R values from − 0.18 to − 0.32. TGW was significantly positively correlated with GL and GW in all four environments (P < 0.01), with a range of R values from 0.69 to 0.91.
QTL mapping for four grain traits
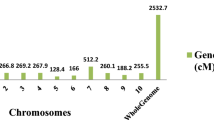
A total of 47 QTL related to the four grain traits were detected across all 10 chromosomes using the ICIM method across four environments (Table 3, Fig. 1). Of these, 8 QTLs were associated with TGW, 6 QTLs with GL, 15 QTLs with GW, and 18 QTLs with GLWR, involving a total of 35 unique loci.
Eight QTLs for TGW were located on chromosomes 2 (1), 3 (1), 4 (2), 7 (2), 8 (1), and 9 (1), respectively. The qTGW8.1 on chromosome 8 was detected in all four environments, with a maximum LOD score of 4.90 and a phenotypic contribution rate of 8.79%. An important QTL on chromosome 4 was detected in three environments, with a maximum LOD score of 6.83 and a phenotypic contribution rate of 9.45%. Except for qTGW7.1, qTGW7.2, and qTGW9.1, the rest of 5 QTLs with additive alleles from the parental line BTx623.
Six QTLs for GL were located on chromosomes 3 (1), 4 (2), 6 (1), and 9 (2), respectively. The qGL4.1 on chromosome 4 was detected in all four environments, with an average LOD score of 5.52 and a range of 3.18 ~ 6.88, explaining an average phenotypic variation of 9.98% and a range of 6.41 ~ 13.01%. One QTL on chromosome 6 was detected in three environments, with a maximum LOD score of 6.52 and a phenotypic contribution rate of 9.88%. Except for qGL3.1 and qGL9.2, the additive alleles of the other four QTLs originated from the parental line BTx623.
Fifteen QTLs for GW were located on chromosomes 1 (1), 2 (1), 3 (1), 4 (4), 5 (1), 6 (3), 7 (2), 8 (1), and 9 (1), respectively. Two important QTLs (qGW6.3 and qGW9.1) detected in three environments were located on chromosome 6 and chromosome 9, with LOD score ranges of 3.32 ~ 4.56 and 3.57 ~ 10.06, and explained phenotypic variation ranges of 3.57 ~ 8.29% and 3.82 ~ 16.35%, respectively. Five QTLs (qGW1.1, qGW1.2, qGW6.3, qGW7.2, and qGW9.1) had additive alleles from HYZ, while the other 10 QTLs had additive alleles from BTx623.
Eighteen QTLs for GLWR were located on chromosomes 1 (1), 2 (1), 3 (7), 4 (2), 5 (1), 6 (1), 7 (1), 8 (2), 9 (1), and 10 (1). On chromosome 3, two QTLs (qGLW2.1 and qGLW3.3) were detected in two environments, with LOD score ranges of 5.70 ~ 12.99 and 3.25 ~ 5.98, and explained phenotypic variation ranges of 8.31 ~ 10.68% and 2.38 ~ 7.94%, respectively. Apart from qGLW2.1, qGLW2.2, qGLW3.2, qGLW3.3, qGLW3.4, qGLW3.7, qGLW4.1, qGLW5.1, and qGLW8.1, with additive alleles from the parental line HYZ, the additive alleles for the other 9 QTLs originated from BTx623.
In summary, among the 35 unique QTLs associated with the four grain traits, 20 loci were repeatedly detected in multienvironments or multitraits (≥ 2) (Fig. 1). The three loci with the most clustered QTLs were located on chromosomes 4, 6, and 9, respectively. The six QTLs for GL and GLWR consistently colocalized on the 58.59 ~ 59.35 Mb region of chromosome 4. The five QTLs for GW and GLWR overlapped on the 54.59 ~ 57.26 Mb region of chromosome 6. The seven QTLs for GL, GW, and TGW overlapped on the 57.67 ~ 57.85 Mb region of chromosome 9.
Candidate genes predicting and expression analysis
To identify candidate genes for 20 important QTLs detected in multienvironments or multitraits, we examined the genomic regions covered by each QTL in the reference BTx623 genome version on Phytozome 13. Among the 20 major QTLs, 6 candidate genes were identified from 6 of these QTLs. The six genes Sobic.001G455900, Sobic.004G035200, Sobic.004G214100, Sobic.004G323600, Sobic.005G150900, and Sobic.006G210300 correspond to the rice homologous genes OsMADS1, RGG2, OsNST1, OsMKK4, OsGRF8, and OsAP2-39, respectively (Table 4).
The expression levels of the six candidate genes showed significant differences in the four seed developmental stages between the parents (Fig. 2, Table S3). The expression trends of Sobic.001G455900 and Sobic.004G035200 were consistent in both parents, with expression levels gradually decreasing as panicle development progressed. For Sobic.004G035200, the expression levels in HYZ were higher than those in BTx623 across all four stages, with expression levels in the first and fourth stages being more than twice those in BTx623. The expression trend of Sobic.004G214100 was similar in both parents, decreasing from S1 to S3 and slightly increasing at S4, with the highest expression levels occurring at S1. In the first two stages of HYZ, the Sobic.004G323600 gene showed almost no change in expression levels gradually decreasing from the third stage, while expression levels in BTx623 remained stable across all four stages. The expression trends of Sobic.005G150900 were completely opposite in the two varieties, with HYZ showing an increase followed by a decrease, reaching the highest level at S2, whereas BTx623 showed a decrease followed by an increase, reaching the lowest level at S2, with HYZ expression levels being more than four times those of BTx623 at this stage. Sobic.006G210300 showed a high-high-low–high expression trend across the four stages in HYZ, whereas in BTx623, the trend was high-low-low–high, with both parents showing the lowest expression levels at S3 stage.
Discussion
Since its introduction to China during the Yuan Dynasty, Sorghum has been an important ingredient for brewing famous distill liquor (Ding et al. 2023). In recent years, there has been a rapid increase in demand for the Maotai-flavor liquor, leading to a continuously expanding supply demand gap for brewing sorghum (Zhang et al. 2023a, b). Sorghum grain traits play a vital role in determining yield and brewing process. The utilization of molecular markers in modern breeding offers a unique opportunity to investigate the genetic control of numerous target traits and monitor the process of incorporating favorable alleles of genes involved in their variation. This study conducted QTL mining for grain-related traits, aiming to accelerate the breeding of high-yield sorghum varieties that meet the requirements of Maotai-flavor liquor production.
QTL mapping for grain-related traits
Currently, extensive research has been conducted on grain-related traits in sorghum, and more than 180 relevant QTLs have been identified (Han and Cai 2019). Using the RIL population of BTx623/HYZ, this study repeatedly detected 20 important QTLs that affect grain size and shape in multi-environments or multi-traits, of which 14 loci are consistent with the reported QTL in previous research.
The QTL on chromosome 1 (qGW1.1, 66.30–73.61 Mb) overlaps with reported grain weight QTLs using four genetic mapping populations, BTx623/S.virgatum (Liu et al. 2019), CK60/China17 (Gelli et al. 2016), CK60/PI229828 (Pereira et al. 1995), and HYP/DTP (Phuong and Stützel 2013). Two adjacent QTLs on chromosome 2 (qGW2.1/qTGW2.1 and qGLW2.1) overlap with the positions of grain roundness QTL reported using the RIL population P118 (Guindo et al. 2019). Two QTLs for GLWR were identified on chromosome 3. qGLW3.2 is consistent with GL QTL using the RILs of P118 and grain weight QTL using the SA2313/Hiro-1 population (Han et al. 2015), while qGLW3.3 is consistent with grain weight QTL mapping with the M35-1/B35 population (Nagaraja et al. 2013). On chromosome 4, two important QTLs (qTGW4.2/qGL4.3 and qGLW4.2/qGL4.1) overlap with grain weight QTL detected using the R931945-2–2*2 / S.bicolor subsp.verticilliflorum population (Tao et al. 2018). The QTL on chromosome 5 (qGLW5.1/qGW5.1) is consistent with grain weight QTL obtained using the Red Kafir/ Takakibi (F2:3) population (Shehzad and Okuno 2015). The QTL on chromosome 6 (qGLW6.1/qGW6.3, 54.59–57.26 Mb) corresponds to the GW QTL obtained from the E-Tian/Ji2731 population (Mocoeur et al. 2015), while another QTL (qGL6.1) on chromosome 6 is consistent with grain weight QTL identified using GWAS analysis (Zhang et al. 2015). There are 2 QTLs on chromosome 7, with qGLW7.1 overlapping with the position of grain weight QTL reported in the P114 population, and qTGW7.2/qGW7.2 overlapping with grain weight QTL positions identified in two RIL populations (BTx623/ S.propinquum and SA2313/ Hiro-1) (Feltus et al. 2006; Han et al. 2015). The QTL affecting TGW and GW (qTGW8.1/qGW8.1) on chromosome 8 has the same position as grain weight QTL identified in the RIL population BTx6 (Boyles et al. 2017). On chromosome 9, the QTL (qTGW9.1/qGL9.2/qGW9.1) is consistent with grain weight QTL reported in the M35-1/B35 population (Nagaraja et al. 2013).
However, there are 6 important QTLs in this study (qTGW3.1/qGW3.1, qTGW3.7/qGL3.1, qGL4.2/qGW4.4, qTGW4.1/qGW4.1, qGLW4.1, and qGL4.2/qGW4.4) that have not been reported in pervious sorghum studies, which may be due to two reasons: Firstly, it could be influenced by differences in the genetic background, population type and size, marker type, and other factors of the mapping population. Currently, the parents of mapping populations for grain trait QTL mapping mostly come from American sweet cultivar, while one of the parents in this study is Chinese brewing glutinous variety “HYZ” and may carry different genes related to grain traits, resulting in differences in the number and position of QTLs (Tao et al. 2021). Additionally, this study used the Sup-GBS technology to construct a high-density map, enabling more precise mapping of QTLs for grain traits. Most of previous QTLs mapping on sorghum have used SSR markers (Han and Cai 2019).
Candidate genes for grain-related traits
Among 20 important QTLs, 6 QTLs were identified candidate genes within its confidence interval. Which four QTLs have been reported in previous studies. The gene Sobic.001G455900 in the interval of qGW1.1 is homologous to OsMADS1 in rice, which encodes a MADS domain transcription factor that regulates grain size and shape, improving the quality and yield of rice (Hu et al. 2015; Liu et al. 2018). The gene Sobic.004G214100 within the qTGW4.2/qGL4.3 interval is homologous to the BC14 gene in rice, which encodes a nucleotide sugar transporter in the Golgi apparatus, regulating the biosynthesis of seed cell walls and plant growth with pleiotropic effects (Song et al. 2011; Zhang et al. 2011). The gene Sobic.005G150900 within qGLW5.1/qGW5.1 interval is homologous to OsGRF8, a growth-regulating factor in rice by regulating flavonoid biosynthesis pathway, which increases rice yield by regulating GL and thousand grain weight (Dai et al. 2019; Yang et al. 2021). The gene Sobic.006G210300 in qGLW6.1/qGW6.3 is homologous to OsAP2-39 in rice, which controls rice growth and seed production through the ABA/GA balance mechanism (Wan et al. 2011; Yaish et al. 2010).
Although two QTLs on chromosome 4 have not been previously reported, the gene Sobic.004G035200 within the (qGLW4.1) interval is homologous to RGG2, which negatively regulates grain size, organ size, and yield of rice by participating in gibberellin biosynthesis (Miao et al. 2019). Another gene (Sobic.004G323600) in the interval of qGL4.2/qGW4.4 is homologous to OsMKK4/SMG1, a key pleiotropic gene controlling rice yield, which regulates cell proliferation through the serine/threonine protein kinase OsMKK4, affecting grain size and environmental stress resistance in rice (Duan et al. 2014; Guo et al. 2020).
Currently, the study of functional genes influencing grain-related traits in sorghum lags significantly behind major cereal crops such as rice (Zhu et al. 2023), maize (Feng et al. 2023), and wheat (Zhang et al. 2023a, b). Only two genes have been cloned and functionally verified in sorghum until now (Tao et al. 2021; Zou et al. 2020). In order to further identify potential candidate genes in this study, subsequent analyses will include sequence divergence analysis among parents to determine genetic polymorphism, development of functional markers, and functional validation through gene editing techniques.
Conclusions
Based on the genetic map, this study identified 6 QTLs for GL, 15 for GW, 8 for TGW, and 18 for GLWR across four environments. In addition, 20 significant QTLs were detected in multi-environments or multi-traits. Among them, six candidate genes potentially related to grain traits were identified, which are homologous to genes controlling grain size in rice. Our findings are critical to selecting target QTL/genes for improving grain-related traits in Chinese brewing sorghum in breeding program.
Data availability
The resequencing data generated in this study can be found in online repositories. The name of the repository and accession number can be found below: China National GeneBank (CNGB); CNP0005779. The website links are as follows: https://db.cngb.org/search/project/CNP0005779/.
References
Baye W, Xie Q, Xie P (2022) Genetic architecture of grain yield-related traits in sorghum and maize. Int J Mol Sci 23:2405
Boyles RE, Cooper EA, Myers MT, Brenton Z, Rauh BL, Morris GP, Kresovich S (2016) Genome-Wide association studies of grain yield components in diverse sorghum germplasm. Plant Genome 9:1–17
Boyles RE, Pfeiffer BK, Cooper EA, Zielinski KJ, Myers MT, Rooney WL, Kresovich S (2017) Quantitative trait loci mapping of agronomic and yield traits in two grain sorghum biparental families. Crop Sci 57:2443–2456
Chen BR, Wang CY, Wang P, Zhu ZX, Xu N, Shi GS, Yu M, Wang N, Li JD, Hou JM, Li SJ, Zhou YF, Gao SJ, Lu XC, Huang RD (2019) Genome-wide association study for starch content and constitution in sorghum (Sorghum bicolor (L.) Moench). J Integra Agric 18:2446–2456
Dai ZY, Tan J, Zhou C, Yang XF, Yang F, Zhang SJ, Sun SC, Miao XX, Shi ZY (2019) The OsmiR396–OsGRF8–OsF3H-flavonoid pathway mediates resistance to the brown planthopper in rice (Oryza sativa). Plant Biotechnol J 17:1657–1669
Ding YQ, Zhou LB, Wang C, Cao N, Cheng B, Gao X, Peng Q, Shao MB, Zhang LY (2019) Research advance in glutinous sorghum for making sauce-flavor liquor in China. Biotechnol Bull 35:28–34
Ding YQ, Cao N, Zhou LB, Cheng B, Gao X, Wang C, Zou GH, Zhang LY (2020) Construction of EMS-induced mutant library and mutant screening in liquor-making waxy sorghum. J South Agric 51:2884–2891
Ding YQ, Xu JX, Wang C, Zhou LB, Zhang GB, Zhao Q, Shao MB, Zhang LY (2023) QTL mapping of grain traits related to brewing in sorghum based on super-GBS technology. J Nucl Agric Sci 37:0241–0250
Duan PG, Rao YC, Zeng DL, Yang YL, Xu R, Zhang BL, Dong GJ, Qian Q, Li YH (2014) SMALL GRAIN 1, which encodes a mitogen-activated protein kinase kinase 4, influences grain size in rice. Plant J 77:547–557
Feltus FA, Hart GE, Schertz KF, Casa AM, Kresovich S, Abraham S, Klein PE, Brown PJ, Paterson AH (2006) Alignment of genetic maps and QTLs between inter- and intra-specific sorghum populations. Theor Appl Genet 112:1295–1305
Feng Z, Cao N, Ding YQ, Xu JX, Gao X, Cheng B, Zou GH, Zhang LY (2022) Mapping QTL for sorghum physical properties using ultra-high-density bin map. J Plant Genet Resour 23:1746–1755
Feng J, Liu PF, Tu L, Gao Y, Guo XY, Wang AG, Zhu YF, Wu X, Chen ZH (2023) QTL Mapping and candidation gend analysis of maize kernel ralated traits. Seed 42:18–24
Gao X, Xu JX, Ding YQ, Wang L, Feng Z, Cao N, Cheng B, Zhang LY (2022) Analysis of physicochemical quality and cooking characteristics of brewer sorghum grains. Food Sci Technol 48:159–166
Gelli M, Mitchell SE, Liu K, Clemente TE, Weeks DP, Zhang C, Holding DR, Dweikat IM (2016) Mapping QTLs and association of differentially expressed gene transcripts for multiple agronomic traits under different nitrogen levels in sorghum. BMC Plant Biol 16:16
Guindo D, Teme N, Vaksmann M, Doumbia M, Vilmus I, Guitton B, Sissoko A, Mestres C, Davrieux F, Fliedel G, Kouressy M, Courtois B, Rami JF (2019) Quantitative trait loci for sorghum grain morphology and quality traits: toward breeding for a traditional food preparation of West-Africa. J Cereal Sci 85:256–272
Guo T, Lu ZQ, Shan JX, Ye WW, Dong NQ, Lin HX (2020) ERECTA1 acts upstream of the OsMKKK10-OsMKK4-OsMPK6 cascade to control spikelet number by regulating cytokinin metabolism in rice. Plant Cell 32:2763–2779
Han LJ, Cai HW (2019) Progress on genetic research of sorghum grain weight. Biotechnol Buiietin 35:15–27
Han LJ, Chen J, Mace ES, Liu YS, Zhu MJ, Yuyama N, Jordan DR, Cai HW (2015) Fine mapping of qGW1, a major QTL for grain weight in sorghum. Theor Appl Genet 128:1813–1825
Hu Y, Liang WQ, Yin CS, Yang XL, Ping BZ, Li AX, Jia R, Chen MJ, Luo ZJ, Cai Q, Zhao XX, Zhang DB, Yuan Z (2015) Interactions of OsMADS1 with floral homeotic genes in rice flower development. Mol Plant 8:1366–1384
Li SG, Liu M, Liu F, Zou JQ, Lu XC, Diao XM (2021) Current status and future prospective of sorghum production and seed industry in China. Sci Agric Sin 54:471–482
Liu Q, Han RX, Wu K, Zhang JQ, Ye YF, Wang SF, Chen JF, Pan YJ, Li Q, Xu XP, Zhou JW, Tao DY, Wu YJ, Fu XD (2018) G-protein βγ subunits determine grain size through interaction with MADS-domain transcription factors in rice. Nat Commun 9:852
Liu HH, Liu HQ, Zhou LN, Lin ZW (2019) Genetic architecture of domestication- and improvement-related traits using a population derived from sorghum virgatum and sorghum bicolor. Plant Sci 283:135–146
Lu QS (1999) Sorghum. China Agriculture Press, Beijing
Miao J, Yang ZF, Zhang DP, Wang YZ, Xu MB, Zhou LH, Wang J, Wu SJ, Yao YL, Du X, Gu FF, Gong ZY, Gu MH, Liang GH, Zhou Y (2019) Mutation of RGG2, which encodes a type B heterotrimeric G protein γ subunit, increases grain size and yield production in rice. Plant Biotechnol J 17:650–664
Mocoeur A, Zhang YM, Liu ZQ, Shen X, Zhang LM, Rasmussen SK, Jing HC (2015) Stability and genetic control of morphological, biomass and biofuel traits under temperate maritime and continental conditions in sweet sorghum (Sorghum bicolour). Theor Appl Genet 128:1685–1701
Morris GP, Ramu P, Deshpande SP, Hash CT, Shah T, Upadhyaya HD, Riera-Lizarazu O, Brown PJ, Acharya CB, Mitchell SE, Harriman J, Glaubitz JC, Buckler ES, Kresovich S (2013) Population genomic and genome-wide association studies of agroclimatic traits in sorghum. Proc Natl Acad Sci USA 110:453–458
Nagaraja RR, Madhusudhana R, Murali MS, Chakravarthi DV, Mehtre SP, Seetharama N, Patil JV (2013) Mapping QTL for grain yield and other agronomic traits in post-rainy sorghum [Sorghum bicolor (L.) Moench]. Theor Appl Genet 126:1921–1939
Pereira MG, Ahnert D, Lee M, Klier K (1995) Genetic mapping of quantitative trait loci panicle characteristics and seed weight in sorghum. Brazil J Genet 18:249–257
Phuong NH, Stützel UR (2013) Quantitative trait loci associated to agronomic traits and yield components in a Sorghum bicolor L.Moench RIL population cultivated under pre-flowering drought and well-watered conditions. Agric Sci 4:781–791
Shehzad T, Okuno K (2015) QTL mapping for yield and yield-contributing traits in sorghum (Sorghum bicolor (L.) Moench) with genomebased SSR marker. Euphytica 203:17–31
Shinoda (1958) Baijiu – the introduction of sorghum. Taiwan Elite Publishing Co Ltd, Taipei
Song XQ, Zhang BC, Zhou YH (2011) Golgi-localized UDP-glucose transporter is required for cell wall integrity in rice. Plant Signal Behav 6:1097–1100
Tang SY, Xie Q (2019) Sorghum-biguse of small crops. Biotechnol. Bulletin 35:1
Tao YF, Mace E, George-Jaeggli B, Hunt C, Cruickshank A, Henzell R, Jordan D (2018) Novel grain weight loci revealed in a cross between cultivated and wild sorghum. Plant Genome 11:1921–1939
Tao YF, Zhao XR, Wang XM, Hathorn A, Hunt C, Cruickshank AW, van Oosterom EJ, Godwin ID, Mace ES, Jordan DR (2020) Large-scale GWAS in sorghum reveals common genetic control of grain size among cereals. Plant Biotechnol J 18:1093–1105
Tao YF, Luo H, Xu JB, Cruickshank A, ZhaoXR TF, Hathorn A, Wu XY, Liu YM, Shatte T, Jordan D, Jing HC, Mace E (2021) Extensive variation within the pan-genome of cultivated and wild sorghum. Nat Plants 7:766–773
Wan LY, Zhang JF, Zhang HW, Zhang ZJ, Quan RD, Zhou SR, Huang RF (2011) Transcriptional activation of OsDERF1 in OsERF3 and OsAP2-39 negatively modulates ethylene synthesis and drought tolerance in rice. PLoS ONE 6:e25216
Yaish MW, El-Kereamy A, Zhu T, Beatty PH, Good AG, Bi YM, Rothstein SJ (2010) The APETALA-2-like transcription factor OsAP2-39 controls key interactions between abscisic acid and gibberellin in rice. PLoS Genet 6:e1001098
Yang XF, Zhao XL, Dai ZY, Ma FL, Miao XX, Shi ZY (2021) OsmiR396/growth regulating factor modulate rice grain size through direct regulation of embryo-specific miR408. Plant Physiol 186:519–533
Zhang BC, Liu XL, Qian Q, Liu LF, Dong GJ, Xiong GY, Zeng DL, Zhou YH (2011) Golgi nucleotide sugar transporter modulates cell wall biosynthesis and plant growth in rice. Proc Natl Acad Sci USA 108:5110–5115
Zhang D, Li JP, Compton RO, Robertson J, Goff VH, Epps E, Kong W, Kim CS, Paterson AH (2015) Comparative genetics of seed size traits in divergent cereal lineages represented by sorghum (Panicoidae) and rice (Oryzoidae). G3 (Bethesda) 5:1117–1128
Zhang LY, Xu JX, Ding YQ, Cao N, Gao X, Feng Z, Li KY, Cheng B, Zhou LB, Ren MJ, Tao YZ, Zou GH (2023a) GWAS of grain color and tannin content in Chinese sorghum based on whole-genome sequencing. Theor Appl Genet 136:77
Zhang ZH, Cheng YK, Ren Y, Wang R, Dong YF, Geng HW (2023b) Genome-wide association analysis of wheat grain related traits. J Plant Genet Resour 24:993–1006
Zhu XY, Gou YJ, Heng YQ, Ding WY, Li YJ, Zhou DG, Li XQ, Liang CR, Wu CY, Wang HY, Shen RX (2023) Targeted manipulation of grain shape genes effectively improves outcrossing rate and hybrid seed production in rice. Plant Biotechnol J 21:381–390
Zou GH, Zhai GW, Yan S, Li SJ, Zhou LB, Ding YQ, Liu HQ, Zhang ZP, Zou JQ, Zhang LY, Chen JP, Xin ZG, Tao YZ (2020) Sorghum qTGW1a encodes a G-protein subunit and acts as a negative regulator of grain size. J Exp Bot 71:5389–5401
Funding
This work was funded by the Guizhou Provincial Basic Research Program (Natural Sciences) (QKHJC[2023]YB169), the Innovation capacity building project of Guizhou scientific institutions (QKFQ[2022]007]), the National Natural Science Foundation of China (32160459), and the Guizhou Academy of Agricultural Sciences Project (Guizhou Agricultural Germplasm Resources (2023) 06).
Author information
Authors and Affiliations
Contributions
NC analyzed data and wrote the manuscript. BC, XG, and JX planted for investigation of grain. YD and WL analyzed data. LZ and GZ conceived, designed the experiments, and revised the manuscript. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Communicated by: Izabela Pawłowicz
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
I declare on behalf of my co-authors that the work described is original, previously unpublished research, and not under consideration for publication elsewhere. The experiments in this study comply with the current laws of China.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cao, N., Ding, Y., Xu, J. et al. QTL analysis of sorghum grain traits based on high-density genetic map. J Appl Genetics (2024). https://doi.org/10.1007/s13353-024-00904-w
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13353-024-00904-w