Abstract
Many studies use strategies that allow for the identification of a large number of genes expressed in response to different stress conditions to which the plant is subjected throughout its cycle. In order to obtain accurate and reliable results in gene expression studies, it is necessary to use reference genes, which must have uniform expression in the majority of cells in the organism studied. RNA isolation of leaves and expression analysis in real-time quantitative polymerase chain reaction (RT-qPCR) were carried out. In this study, nine candidate reference genes were tested, actin 11 (ACT11), ubiquitin conjugated to E2 enzyme (UBC-E2), glyceraldehyde-3-phosphate dehydrogenase (GAPDH), beta tubulin (β-tubulin), eukaryotic initiation factor 4α (eIF-4α), ubiquitin 10 (UBQ10), ubiquitin 5 (UBQ5), aquaporin TIP41 (TIP41-Like) and cyclophilin, in two genotypes of rice, AN Cambará and BRS Querência, with different levels of soil moisture (20%, 10% and recovery) in the vegetative (V5) and reproductive stages (period preceding flowering). Currently, there are different softwares that perform stability analyses and define the most suitable reference genes for a particular study. In this study, we used five different methods: geNorm, BestKeeper, ΔCt method, NormFinder and RefFinder. The results indicate that UBC-E2 and UBQ5 can be used as reference genes in all samples and softwares evaluated. The genes β-tubulin and eIF-4α, traditionally used as reference genes, along with GAPDH, presented lower stability values. The gene expression of basic leucine zipper (bZIP23 and bZIP72) was used to validate the selected reference genes, demonstrating that the use of an inappropriate reference can induce erroneous results.












Similar content being viewed by others
References
Alves MS, Dadalto SP, Gonçalves AB et al (2013) Plant bZIP transcription factors responsive to pathogens: a review. Int J Mol Sci 14:7815–7828. doi:10.3390/ijms14047815
Andersen CL, Jensen JL, Ørntoft TF (2004) Normalization of real-time quantitative reverse transcription-PCR data: a model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res 64:5245–5250. doi:10.1158/0008-5472.CAN-04-0496
Borowski JM, Galli V, da Silva Messias R et al (2014) Selection of candidate reference genes for real-time PCR studies in lettuce under abiotic stresses. Planta 239:1187–1200. doi:10.1007/s00425-014-2041-2
Brunner AM, Yakovlev IA, Strauss SH (2004) Validating internal controls for quantitative plant gene expression studies. BMC Plant Biol 4:14. doi:10.1186/1471-2229-4-14
Bustin SA (2002) Quantification of mRNA using real-time reverse transcription PCR (RT-PCR): trends and problems. J Mol Endocrinol 29:23–39. doi:10.1677/jme.0.0290023
Caldana C, Scheible W-R, Mueller-Roeber B, Ruzicic S (2007) A quantitative RT-PCR platform for high-throughput expression profiling of 2500 rice transcription factors. Plant Methods 3(1):7. doi:10.1186/1746-4811-3-7
Cotsaftis O, Plett D, Johnson AAT et al (2011) Root-specific transcript profiling of contrasting rice genotypes in response to salinity stress. Mol Plant 4:25–41. doi:10.1093/mp/ssq056
Cruz-Rus E, Amaya I, Sánchez-Sevilla JF et al (2011) Regulation of L-ascorbic acid content in strawberry fruits. J Exp Bot 62:4191–4201. doi:10.1093/jxb/err122
Czechowski T, Stitt M, Altmann T et al (2005) Genome-wide identification and testing of superior reference genes for transcript normalization. Plant Physiol 139:5–17. doi:10.1104/pp.105.063743
de Oliveira LA, Breton MC, Bastolla FM et al (2012) Reference genes for the normalization of gene expression in eucalyptus species. Plant Cell Physiol 53:405–422. doi:10.1093/pcp/pcr187
Derveaux S, Vandesompele J, Hellemans J (2010) How to do successful gene expression analysis using real-time PCR. Methods 50:227–230. doi:10.1016/j.ymeth.2009.11.001
Dreher K, Callis J (2007) Ubiquitin, hormones and biotic stress in plants. Ann Bot 99:787–822. doi:10.1093/aob/mcl255
Expósito-Rodríguez M, Borges AA, Borges-Pérez A et al (2008) Selection of internal control genes for quantitative real-time RT-PCR studies during tomato development process. BMC Plant Biol 8:131. doi:10.1186/1471-2229-8-131
Fleige S, Pfaffl MW (2006) RNA integrity and the effect on the real-time qRT-PCR performance. Mol Aspects Med 27:126–139. doi:10.1016/j.mam.2005.12.003
Foley JA, Ramankutty N, Brauman KA et al (2011) Solutions for a cultivated planet. Nature 478:337–342. doi:10.1038/nature10452
Guénin S, Mauriat M, Pelloux J et al (2009) Normalization of qRT-PCR data: the necessity of adopting a systematic, experimental conditions-specific, validation of references. J Exp Bot 60:487–493. doi:10.1093/jxb/ern305
Gutierrez L, Mauriat M, Guénin S et al (2008) The lack of a systematic validation of reference genes: a serious pitfall undervalued in reverse transcription-polymerase chain reaction (RT-PCR) analysis in plants. Plant Biotechnol J 6:609–618. doi:10.1111/j.1467-7652.2008.00346.x
Hernandez-Garcia CM, Martinelli AP, Bouchard RA et al (2009) A soybean (Glycine max) polyubiquitin promoter gives strong constitutive expression in transgenic soybean. Plant Cell Rep 28:837–849. doi:10.1007/s00299-009-0681-7
Jain M, Nijhawan A, Tyagi AK et al (2006) Validation of housekeeping genes as internal control for studying gene expression in rice by quantitative real-time PCR. Biochem Biophys Res Commun 345:646–651. doi:10.1016/j.bbrc.2006.04.140
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(−Delta Delta C(T)) Method. Methods 25:402–408. doi:10.1006/meth.2001.1262
Maroufi A, Van Bockstaele E, De Loose M (2010) Validation of reference genes for gene expression analysis in chicory (Cichorium intybus) using quantitative real-time PCR. BMC Mol Biol 11:15. doi:10.1186/1471-2199-11-15
Moraes GP, Benitez LC, do Amaral MN et al (2015) Evaluation of reference genes for RT-qPCR studies in the leaves of rice seedlings under salt stress. Genet Mol Res 14:2384–2398. doi:10.4238/2015.March.27.24
Muñoz-Bertomeu J, Cascales-Miñana B, Mulet JM et al (2009) Plastidial glyceraldehyde-3-phosphate dehydrogenase deficiency leads to altered root development and affects the sugar and amino acid balance in Arabidopsis. Plant Physiol 151:541–558. doi:10.1104/pp.109.143701
Pfaffl MW, Tichopad A, Prgomet C et al (2004) Determination of stable housekeeping genes, differentially regulated target genes and sample integrity: BestKeeper—Excel-based tool using pair-wise correlations. Biotechnol Lett 26:509–515. doi:10.1023/B:BILE.0000019559.84305.47
Radonić A, Thulke S, Mackay IM et al (2004) Guideline to reference gene selection for quantitative real-time PCR. Biochem Biophys Res Commun 313:856–862. doi:10.1016/j.bbrc.2003.11.177
Rasmussen R (2001) Quantification on the LightCycler. In: Meuer S, Wittwer C, Nakagawara KI (eds) Rapid cycle real-time PCR. Methods and applications. Springer, Berlin, pp 21–34. doi: 10.1007/978-3-642-59524-0_3
Serraj R, McNally KL, Slamet-Loedin I et al (2011) Drought resistance improvement in rice: an integrated genetic and resource management strategy. Plant Prod Sci 14:1–14. doi:10.1626/pps.14.1
Shivhare R, Lata C (2016) Selection of suitable reference genes for assessing gene expression in pearl millet under different abiotic stresses and their combinations. Sci Rep 6:23036. doi:10.1038/srep23036
Silver N, Best S, Jiang J et al (2006) Selection of housekeeping genes for gene expression studies in human reticulocytes using real-time PCR. BMC Mol Biol 7:33. doi:10.1186/1471-2199-7-33
Sun CW, Griffen S, Callis J (1997) A model for the evolution of polyubiquitin genes from the study of Arabidopsis thaliana ecotypes. Plant Mol Biol 34:745–758. doi:10.1023/A:1005848828368
Vandesompele J, De Preter K, Pattyn F et al (2002) Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol 3:research0034.1. doi:10.1186/gb-2002-3-7-research0034
Walia H, Wilson C, Condamine P et al (2005) Comparative transcriptional profiling of two contrasting rice genotypes under salinity stress during the vegetative growth stage. Plant Physiol 139:822–835. doi:10.1104/pp.105.065961
Wan H, Zhao Z, Qian C et al (2010) Selection of appropriate reference genes for gene expression studies by quantitative real-time polymerase chain reaction in cucumber. Anal Biochem 399:257–261. doi:10.1016/j.ab.2009.12.008
Zhang Z, Hu J (2007) Development and validation of endogenous reference genes for expression profiling of Medaka (Oryzias latipes) exposed to endocrine disrupting chemicals by quantitative real-time RT-PCR. Toxicol Sci 95(2):356–368. doi:10.1093/toxsci/kfl161
Acknowledgements
The authors thank the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES) and the Research Foundation of Rio Grande do Sul (FAPERGS) for supporting the research, as well as the companies Agro Norte, MT and Embrapa Temperate Climate for providing seeds.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Author contributions
Conceived and designed the experiments: PAA, LCB, MNA and EJBB. Performed the experiments: PAA, MNA, ILV and GSR. Analysed the data: PAA, MNA, LCM and EJBB. Wrote the paper: PAA. Corrected the manuscript: LCM and EJBB.
Additional information
Communicated by: Andrzej Górny
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Material 1
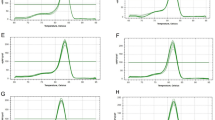
Specificity of candidate reference genes for rice genotypes AN Cambará and BRS Querência in vegetative and reproductive stages (GIF 58 kb)
Rights and permissions
About this article
Cite this article
Auler, P.A., Benitez, L.C., do Amaral, M.N. et al. Evaluation of stability and validation of reference genes for RT-qPCR expression studies in rice plants under water deficit. J Appl Genetics 58, 163–177 (2017). https://doi.org/10.1007/s13353-016-0374-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13353-016-0374-1