Abstract
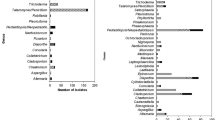
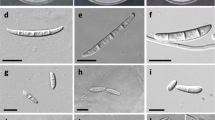
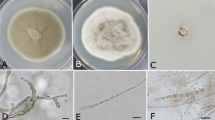
A rapid increase in the number of Gossypium hirsutum L. plants suddenly wilting and dying in a commercial cotton field in Central Queensland initiated this study. The aim was to characterise fungal species recovered from discoloured vascular tissue of dead plants and determine if they contributed to their death. Isolations were consistently dominated by one fungus based on culture morphology. Identification was established on sequences of the internal transcribed spacer (ITS) region of ribosomal DNA and revealed that the isolates all belong to Diatrypaceae, with high homology to Eutypella scoparia. Further analyses showed that there were two distinct Eutypella species present in the isolates, which are quite different from E. scoparia from the sequence dissimilarity due to the presence of ITS1 and ITS2 insertions. In diseased root samples, community profiling showed two operational taxonomic units (OTUs) related to E. scoparia were the most abundant fungi accounting for 45 to 99% of all sequences. Pathogenicity tests showed that a Eutypella isolate when inoculated into the stem of healthy G. hirsutum using two inoculation methods caused cankerous growth and necrosis of vascular tissue, typical of trunk disease. The fungus caused a red—brown streaking of the vascular tissue like that observed in diseased field plants. This study shows that the fungal isolates, which form a distinct group within the Eutypella, are associated with the root and stem of dying G. hirsutum and were the dominant fungi of diseased roots. This is the first known case of Eutypella affecting cotton worldwide and is considered an expansion of this genus’ host range.








Similar content being viewed by others
References
Allen SJ, Nehl DB (1999) Maximising mycorrhizal infection in cotton. Final report prepared for the CRC for Sustainable Cotton Production. NSW Agriculture
Bakkeren G, Kronstad JW, Levesque CA (2000) Comparison of AFLP fingerprints and ITS sequences as phylogenetic markers in Ustilaginomycetes. Mycologia 92:510–521. https://doi.org/10.1080/00275514.2000.12061187
Carter MV (1957) Eutypa armeniacae Hansf. & Carter, sp. nov., an airborne vascular pathogen of Prunus armeniaca L. in Southern Australia. Aust J Bot 5:21–35. https://doi.org/10.1071/BT9570021
Carter MV (1991) The status of Eutypa lata as a pathogen. Monograph, Phytopathological Paper No 32. Commonwealth Agricultural Bureau, International Mycological Institute, UK
Correia KC, Câmara MPS, Barbosa MAG, Sales Jr R, Agustí-Brisach C, Gramaje D, León M, García-Jiménez J, Abad-Campos P, Armengol J, Michereff SJ (2013) Fungal trunk pathogens associated with table grape decline in Northeastern Brazil. Phytopathol Mediterr 52(2):380–387. https://doi.org/10.14601/Phytopathol_Mediterr-11377
Cotton Australia (2014) Australian Grown Cotton Sustainability Report, 2014 https://www.cottonaustralia.com.au/industry. Accessed 15 April 2021
Cotton Australia (2021) Cotton Australia Economics https://cottonaustralia.com.au/economics. Accessed 15 April 2021
Crespo M, Lawrence DP, Nouri MT, Doll DA, Trouillas FP (2019) Characterization of Fusarium and Neocosmospora species associated with crown rot and stem canker of pistachio rootstocks in California. Plant Dis 103:1931–1939. https://doi.org/10.1094/pdis-11-18-2012-re
Davidson RW, Lorenz RC (1938) Species of Eutypella and Schizoxylon associated with cankers of maple. Phytopathol 28:733–745
Davis RM, Colyer PD, Rothrock CS, Kochman JK (2006) Fusarium wilt of cotton: Population diversity and implications for management. Plant Dis 90(6):692–703. https://doi.org/10.1094/PD-90-0692
Deshpande V, Wang Q, Greenfield P, Charleston M, Porras-Alfaro A, Kuske CR, Cole JR, Midgley DJ, Nai T-D (2016) Fungal identification using a Bayesian classifier and the Warcup training set of internal transcribed spacer sequences. Mycologia 108:1–5. https://doi.org/10.3852/14-293
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32(5):1792–1797. https://doi.org/10.1093/nar/gkh340
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinform 26(19):2460–2461. https://doi.org/10.1093/bioinformatics/btq461
Eskandari S, Guppy CN, Knox OCG, Backhouse D, Haling RE (2017) Mycorrhizal symbioses of cotton grown on sodic soils: A review from an Australian Perspective. Pedosphere 27:1015–1026. https://doi.org/10.1016/S1002-0160(17)60491-0
Glawe DA, Rogers JD (1984) Diatrypaceae in the Pacific Northwest. Mycotaxon 20:401–460
Gottwald TR (2001) Diseases of Pecan. American Phytopathological Society. https://www.apsnet.org/edcenter/resources/commonnames/Pages/Pecan.aspx. Accessed 15 April 2021
Gramaje D, Urbez-Torres JR, Sosnowski MR (2018) Managing grapevine trunk diseases with respect to etiology and epidemiology: Current strategies and future prospects. Plant Dis 102:12–39. https://doi.org/10.1094/pdis-04-17-0512-fe
Gupta VVSR, Bramley RGV, Greenfield P, Yu J, Herderich MJ (2019) Vineyard soil microbiome composition related to rotundone concentration in Australian cool climate ‘peppery’ Shiraz grapes. Frontiers in Microbiol 10:1607. https://doi.org/10.3389/fmicb.2019.01607
Madeira F, Park YM, Lee J, Buso N, Gur T, Madhusoodanan N, Basutkar P, Tivey ARN, Potter S, Finn RD, Lopez R (2019) The EMBL-EBI search and sequence analysis tools APIs in 2019. Nucleic Acids Res 47(W1):W636–W641. https://doi.org/10.1093/nar/gkz268
Magarey PA, Carter MV (1986) New technology facilitates control of Eutypa dieback in apricots and grapevines. Plant Prot Q 1:156–159
Mayorquin JS, Wang DH, Twizeyimana M, Eskalen A (2016) Identification, distribution, and pathogenicity of Diatrypaceae and Botryosphaeriaceae associated with citrus branch canker in the southern California desert. Plant Dis 100:2402–2413. https://doi.org/10.1094/pdis-03-16-0362-re
Moller WJ, Kasimatis AN (1978) Dieback of grapevine caused by Eutypa armeniacae. Plant Dis Rep 62:254–258
Moyo P, Mostert L, Spies CFJ, Damm U, Halleen F (2018) Diversity of Diatrypaceae species associated with dieback of grapevines in South Africa, with the description of Eutypa cremea sp. nov. (2018) Plant Dis 102(1):220–230. https://doi.org/10.1094/pdis-05-17-0738-re
Ophel-Keller K, McKay A, Hartley D, Curran H, Curran J (2008) Development of a routine DNA-based testing service for soilborne diseases in Australia. Australas Plant Pathol 37:243–253. https://doi.org/10.1071/AP08029
Paolinelli-Alfonso M, Serrano-Gomez C, Hernandez-Martinez R (2015) Occurrence of Eutypella microtheca in grapevine cankers in Mexico. Phytopathol Mediterr 54(1):86–93. http://www.fupress.net/index.php/pm/article/view/14998/15062
Pearson RC, Burr TJ (1981) Eutypa dieback. Disease Identification Sheet No. 1. NY State Agric Exp Stn Geneva, NY. http://hdl.handle.net/1813/43093
Pitt WM, Huang R, Trouillas FP, Savocchia S, Steel CC (2010) Evidence that Eutypa lata and other Diatrypaceous species occur in New South Wales vineyards. Australas Plant Pathol 39:97–106. https://doi.org/10.1071/AP09051
Pitt WM, Trouillas FP, Gubler WD, Savocchia S, Sosnowski MR (2013) Pathogenicity of diatrypaceous fungi on grapevines in Australia. Plant Dis 97:749–756. https://doi.org/10.1094/PDIS-10-12-0954-RE
Rambaut A (2018) FigTree v1.4.4. Institute of Evolutionary Biology, University of Edinburgh, Edinburgh. https://github.com/rambaut/figtree
Rolshausen PE, Sosnowski MR, Trouillas FP, Gubler WD (2015) Eutypa dieback. In: Wilcox WF, Gubler WD, Uyemoto KJ (eds) Compendium of Grape Diseases, Disorders, and Pests, 2nd edn. American Phytopathological Society Press, pp 57–61
Sas A (2020) Rural R&D for Profit Program, Digital technologies for dynamic management of disease, stress and yield, Final Report Wine Australia June 2016 – August 2020. https://www.wineaustralia.com/getmedia/f5851743-0583-463c-8aed-ac3b1476338c/RnD-1601-RnD4Profit-15-02-018-Digital-technologies-FINAL-REPORT.pdf
Sosnowski M (2018) Best practice management guide Eutypa dieback. (Version 1.1, Feb 2018) Wine Australia. https://www.wineaustralia.com/getmedia/850410cd-bcea-4b5c-b593-f4d42ae33fdd/RD_BPMG_EutypaDieback_Feb2018_Final
Sosnowski M, Lardner R, Scott E (2006) Diagnosis and management of eutypa dieback. Final Report, Grape and Wine Research and Development Corporation, Project S2.2.4 (CRV 03/06S), July 2006, 106 pp. https://www.wineaustralia.com/getmedia/ea804245-1b8d-482e-a914-d14a6177ebb9/CRV-03-06S-Final-Report
Sosnowski MR, Lardner R, Wicks TJ, Scott ES (2007) The influence of grapevine cultivar and isolate of Eutypa lata on wood and foliar symptoms. Plant Dis 91:924–931. https://doi.org/10.1094/pdis-91-8-0924
Sosnowski MR, Luque J, Loschiavo AP, Martos S, Garcia-Figueres F, Wicks T, Scott ES (2011) Studies on the effect of water and temperature stress on grapevines inoculated with Eutypa lata. Phytopathol Mediterr 50:S127–S138. https://doi.org/10.14601/Phytopathol_Mediterr-8959
Stiller WN, Wilson IW (2014) Australian cotton germplasm resources. In: Abdurakhmonov IY (ed) World Cotton Germplasm Resources. IntechOpen: London, UK, pp 1–34
Taieb S, Triki M, Hammami I, Rhouma A (2014) First report of dieback of olive trees caused by Phoma fungicola in Tunisia. 96(4)S117. https://doi.org/10.4454/JPP.V96I4.013
Toju H, Tanabe AS, Yamamoto S, Sato H (2012) High-Coverage ITS Primers for the DNA-Based Identification of Ascomycetes and Basidiomycetes in Environmental Samples. PLoS ONE 7(7):e40863. https://doi.org/10.1371/journal.pone.0040863
Trouillas FP, Gubler WD (2010) Host range, biological variation, and phylogenetic diversity of Eutypa lata in California. Phytopathol 100:1048–1056. https://doi.org/10.1094/PHYTO-02-10-0040
Trouillas F, Pitt W, Sosnowski M, Huang R, Fr P, Loschiavo A, Savocchia S, Scott E, Gubler W (2011) Taxonomy and DNA phylogeny of Diatrypaceae associated with Vitis vinifera and other woody plants in Australia. Fungal Divers 49:203–223. https://doi.org/10.1007/s13225-011-0094-0
Úrbez-Torres JR, Adams P, Kamas J, Gubler WD (2009) Identification, incidence, and pathogenicity of fungal species associated with grapevine dieback in Texas. Vitis 4:497–507
Waterhouse AM, Procter JB, Martin DMA, Clamp M, Barton GJ (2009) Jalview Version 2 - a multiple sequence alignment editor and analysis workbench. Bioinform 25(9):1189–1191. https://doi.org/10.1093/bioinformatics/btp033
Yuan ZQ (1996) Fungi and Associated Tree Diseases in Melville Island. Northern Territory, Australia Aust Systematic Bot 9:337–360. https://doi.org/10.1071/SB9960337
Acknowledgements
This work was supported by the Cotton Research and Development Corporation (funding for DAQ2002). Funding for the individual participation was provided by Department of Agriculture and Fisheries in Queensland (LJS, DK, LS, JC) and Commonwealth Scientific and Industrial Research Organisation (PG, VVSRG). The authors wish to thank Juliane Henderson and Elizabeth Czislowski from Biosecurity Queensland for their contribution in determining the cause of disease was not a known exotic pathogen. Sincere thank you to Mark Sosnowski (South Australian Research and Development Institute) for helpful advice and discussions on Diatrypaceae species. Many thanks to Roger Mitchell, Sarah Dodd and Andrew Manners from GrowHelp for their assistance with characterisation of fungal isolates. Thank you to Stephen Neate and Murray Sharman for reviewing this document and providing feedback. We thank cotton growers for helpful discussions on their observations in field and allowing us to collect samples.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare that they have no conflict of interest.
Supplementary information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Smith, L., Greenfield, P., Kafle, D. et al. First report of Eutypella species associated with dying cotton (Gossypium hirsutum L.). Australasian Plant Pathol. 51, 101–116 (2022). https://doi.org/10.1007/s13313-021-00843-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13313-021-00843-8