Abstract
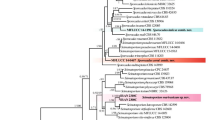
In the late 1990s, a novel Botryosphaeria-like fungal pathogen was observed causing a disease on wheat in Queensland, characterised as white grain disorder (WGD). In recent years, this disease has sporadically appeared across the eastern states of Australia. In this study, internal transcribed spacer (ITS) region sequences were used to compare these fungi to other Botryosphaeriaceae spp. to show that they should be reclassified as members of the Eutiarosporella genus. Using a small population of WGD isolates, we built a three-loci maximum likelihood tree, using ITS, β-tubulin, and Elongation Factor1-α sequences to show that there are three separate Eutiarosporella spp. found in infected grain. This multigene tree, with the support of phenotypic differences between clades observed in vitro, show that that the causal agents of WGD should be delimited into three divergent species; Eutiarosporella tritici-australis, Eutiarosporella darliae, and Eutiarosporella pseudodarliae.






Similar content being viewed by others
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Alvarez-Loayza P, White JF Jr, Torres MS, Balsev H, Kristiansen T, Svenning JC, Gil N (2011) Light converts endosymbiotic fungus to pathogen, influencing seedling survival and niche-space filling of a common tropical tree, Iriartea deltoidea. PLoS One 6, e16386
Aoki T, Vaughan MM, McCormick SP, Busman M, Ward T, Kelly A, O’Donnell K, Johnston P, Geiser DM (2014) Fusarium dactylidis sp. nov., a novel nivalenol toxin-producing species sister to F. pseudograminearum isolated from orchard grass (Dactylis glomerata) in Oregon and New Zealand. Mycologia: doi:10.3852/14-213
Bankevich A, Nurk S, Antipov D, Gurevich AA, Dvorkin M, Kulikov AS, Lesin VM, Nikolenko SI, Pham S, Prjibelski AD (2012) SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol 19:455–477
Barber PA, Burgess TJ, Hardy GE, Slippers B, Keane PJ, Wingfield MJ (2005) Botryosphaeria species from Eucalyptus in Australia are pleoanamorphic, producing dichomera synamorphs in culture. Mycol Res 109:1347–1363
Bihon W, Wingfield MJ, Slippers B, Duong TA, Wingfield BD (2014) MAT gene idiomorphs suggest a heterothallic sexual cycle in a predominantly asexual and important pine pathogen. Fungal Genet Biol 62:55–61
Blanco-Ulate B, Rolshausen P, Cantu D (2013) Draft genome sequence of Neofusicoccum parvum isolate UCR-NP2, a fungal vascular pathogen associated with grapevine cankers. Genome Announcements 1:e00339–13
Crous PW, Slippers B, Wingfield MJ, Rheeder J, Marasas WF, Philips AJ, Alves A, Burgess T, Barber P, Groenewald JZ (2006) Phylogenetic lineages in the Botryosphaeriaceae. Stud Mycol 55:235–253
Crous PW, Muller MM, Sanchez R, Giordano L, Bianchinotti MV, Anderson FE, Groenewald JZ (2015) Resolving Tiarosporella spp. allied to Botryosphaeriaceae and Phacidiaceae. Phytotaxa 202:73–93
De Leon C, Jeffers D, Maize I, Center WI (2004) Maize diseases: a guide for field identification, 4th edn. International Maize and Wheat Improvement Center (CIMMYT), Mexico, 119 p
Dettman JR, Jacobson DJ, Taylor JW (2003) A multilocus genealogical approach to phylogenetic species recognition in the model eukaryote Neurospora. Evolution 57:2703–2720
Evans M (2013) White grain rejection spreading. Ground Cover Supplement 102:4
Gardiner DM, McDonald MC, Covarelli L, Solomon PS, Rusu AG, Marshall M, Kazan K, Chakraborty S, McDonald BA, Manners JM (2012) Comparative pathogenomics reveals horizontally acquired novel virulence genes in fungi infecting cereal hosts. PLoS Pathog 8, e1002952
Hyde KD (1993) Tropical Australian freshwater fungi VI. Tiarosporella paludosa and Clohesyomyces aquaticus gen. et sp. Nov. (Coelomycetes). Aust Syst Bot 6:169–173
Hyde KD, Chomnunti P, Crous PW, Groenewald JZ, Damm U, Ko TWK, Shivas RG, Summerell BA, Tan YP (2010) A case for re-inventory of Australia’s plant pathogens. Persoonia: Mol Phylogeny Evol Fungi 25:50–60
Islam MS, Haque M, Emdad EM, Halim A, Hossen QMM, Hossain MZ, Ahmed B, Rahim S, Rahman MS (2012) Tools to kill: genomes of one of the most destructive plant pathogenic fungi Macrophomina phaseolina. BMC Genomics 13:493
Jami F, Slippers B, Wingfield MJ, Gryzenhout M (2012) Five new species of the Botryosphaeriaceae from Acacia Karroo in South Africa. Cryptogam Mycol 33:245–266
Jami F, Slippers B, Wingfield MJ, Gryzenhout M (2013) Greater Botryosphaeriaceae diversity in healthy than associated diseased Acacia karroo tree tissues. Australas Plant Pathol 42:421–430
Jami F, Slippers B, Wingfield MJ, Gryzenhout M (2014) Botryosphaeriaceae species overlap on four unrelated, native South African hosts. Fungal Biol 118:168–179
Klosterman SJ, Subbarao KV, Kang S, Veronese P, Gold SE, Thomma BP, Chen Z, Henrissat B, Lee YH, Pk J (2011) Comparative genomics yields insights into niche adaptation of plant vascular wilt pathogens. PLoS Pathog 7, e1002137
Kopinski JS, Blaney BJ (2010) Nutritive value and non-toxicity of Botryosphaeria zeae-infected wheat for weaner pigs. J Anim Physiol Anim Nutr 94:44–54
Leigh JW, Susko E, Baumgartner M, Roger AJ (2008) Testing congruence in phylogenomic analysis. Syst Biol 57:104–115
Lohse M, Bolger AM, Nagel A, Fernie AR, Lunn JE, Stitt M, Usadel B (2012) RobiNA: a user-friendly, integrated software solution for RNA-Seq-based transcriptomics. Nucleic Acids Res 40:W622–W627
Luque J, Martos S, Phillips AJL (2005) Botryosphaeria viticola sp. nov. on grapevines: a new species with a Dothiorella anamorph. Mycologia 97:1111–1121
McDonald MC, Razavi M, Friesen TL, Brunner PC, McDonald BA (2012) Phylogenetic and population genetic analyses of Phaeosphaeria nodorum and its close relatives indicate cryptic species and an origin in the Fertile Crescent. Fungal Genet Biol 49:882–895
Mead O, Thynne E, Winterberg B, Solomon PS (2013) Characterising the Role of GABA and Its Metabolism in the Wheat Pathogen Stagonospora nodorum. PLoS One 8, e78368
Murray GM, Brennan JP (2009) Estimating disease losses to the Australian wheat industry. Australas Plant Pathol 38:558–570
Penfold K (2011) White grains may be blight disease. GRDC Ground Cover 92
Phillips A, Alves A, Abdollahzadeh J, Slippers B, Wingfield MJ, Groenewald J, Crous P (2013) The Botryosphaeriaceae: genera and species known from culture. Stud Mycol 76:51–167
Platz G (2011) Wheat and barley disease management in 2011. Yellow spot and head diseases in wheat. Strategies and products for barley leaf rust. GRDC Update Papers
Sakalidis M, Hardy GE, Burgess TI (2011a) Endophytes as potential pathogens of the baobab species Adansonia gregorii: a focus on the Botryosphaeriaceae. Fungal Ecol 4:1–14
Sakalidis M, Hardy GE, Burgess TI (2011b) Use of the Genealogical Sorting Index (GSI) to delineate species boundaries in the Neofusicoccum parvum–Neofusiccocum ribis species complex. Mol Phylogenet Evol 60:333–344
Sakalidis M, Slippers B, Wingfield B, Hardy GSJ, BURGESS T (2013) The challenge of understanding the origin, pathways and extent of fungal invasions: global populations of the Neofusicoccum parvum–N. ribis species complex. Divers Distrib 19:873–883
Slippers B, Wingfield MJ (2007) Botryosphaeriaceae as endophytes and latent pathogens of woody plants: diversity, ecology and impact. Fungal Biol Rev 21:90–106
Slippers B, Crous PW, Denman S, Coutinho TA, Wingfield BD, Wingfield MJ (2004a) Combined multiple gene genealogies and phenotypic characters differentiate several species previously identified as Botryosphaeria dothidea. Mycologia 96:83–101
Slippers B, Fourie G, Crous PW, Coutinho TA, Wingfield BD, Carnegie AJ, Wingfield MJ (2004b) Speciation and distribution of Botryosphaeria spp. on native and introduced Eucalyptus trees in Australia and South Africa. Stud Mycol 50:343–358
Slippers B, Fourie G, Crous PW, Coutinho TA, Wingfield BD, Wingfield MJ (2004c) Multiple gene sequences delimit Botryosphaeria australis sp. nov. from B. lutea. Mycologia 96:1030–1041
Slippers B, Summerell BA, Crous PW, Coutinho TA, Wingfield BD, Wingfield MJ (2005) Preliminary studies on Botryosphaeria species from Southern Hemisphere conifers in Australasia and South Africa. Australas Plant Pathol 34:213–220
Slippers B, Burgess T, Pavlic D, Ahumada R, Maleme S, Rodas C, Wingfield MJ (2009) A diverse assemblage of Botryosphaeriaceae infect Eucalyptus in native and non-native environments. South For 71:101–110
Slippers B, Boissin E, Phillips A, Groenewald J, Lombard L, Wingfield M, Postma A, Burgess T, Crous P (2013) Phylogenetic lineages in the Botryosphaeriales: a systematic and evolutionary framework. Stud Mycol 76:31–49
Stamatakis A (2006) RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22:2688–2690
Sutton BC, Marasas WFO (1976) Observations on Neottiosporina and Tiarosporella. Trans Br Mycol Soc 67:69–76
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Taylor A, Hardy GESJ, Wood P, Burgess T (2005) Identification and pathogenicity of Botryosphaeria species associated with grapevine decline in Western Australia. Australas Plant Pathol 34:187–195
Thynne E, McDonald MC, Solomon PS (2015) Phytopathogen emergence in the genomics era. Trends Plant Sci. doi:10.1016/j.tplants.2015.01.009
Vasconcelos AF, Dekker RH, Barbosa A, Paccola-Meirelles L (2001) Comparison of the laccases, molecular marker proteins, and induction of pycnidia by three species of Botryosphaeriaceous fungi. Mycoscience 42:543–548
White TJ, Bruns T, Lee SJWT, Taylor JW (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. PCR Protoc: Guide Methods Appl 18:315–322
Wildermuth G, Williamson P, Shivas R, McNamara R (2001) Premature head blight and white grain – a new disease of wheat. Proceedings of 13th Biennial Australasian Plant Pathology Conference, Queensland Department of Primary Industries
Zhou YJ, Zhang J, Wang XD, Yang L, Jang DH, Li GQ, Hsiang T, Zhuang WY (2014) Morphological and phylogenetic identification of Botryotis sinoviticola, a novel cryptic species causing grey mold disease of table grapes (Vitis vinifera) in China. Mycologia 106:43–56
Acknowledgments
This study was supported by the Grains Research and Development Corporation (GRDC) (ANU00019). Peter Solomon is an Australian Research Council Future Fellow and Elisha Thynne is a recipient of an Australian Postgraduate Award and a GRDC Grains Industry Research Scholarship.
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Supplementary Table 1
(DOCX 37 kb)
Supplementary Fig 1
Phylogenetic analyses separate isolates of WGD fungi into three distinct clades. Tree is rooted to Botryosphaeria agaves. (JPEG 553 kb)
Supplementary Fig 2
Phylogenetic analyses comparing isolates of WGD fungi with a range of species from the former genus of Tiarosporella spp.. Tree is rooted to Marasasiomyces karoo. (JPEG 1739 kb)
Supplementary Fig 3
Alignment of EF1-α sequences of the isolates of wheat-infecting Tiarosporella spp. focusing on the region of base-deletion in B. zeae and Clade 2 B. zeae. (JPEG 473 kb)
Supplementary Fig 4
Growth of each of the clades of wheat-infecting Tiarosporella spp. at different temperatures. (JPEG 2495 kb)
Supplementary Fig 5
Histogram displaying the comparative conidial dimensions of Tiarosporella spp. in relation to WGD fungi. (JPEG 7644 kb)
Supplementary Fig 6
Image of Clade 1 pigment production on liquid minimal media (with sucrose and glutamine). (JPEG 1334 kb)
Supplementary Fig 7
Growth of each of the three clades on OMA, GABA-OMA,V8-OMA. (JPEG 979 kb)
Rights and permissions
About this article
Cite this article
Thynne, E., McDonald, M.C., Evans, M. et al. Re-classification of the causal agent of white grain disorder on wheat as three separate species of Eutiarosporella . Australasian Plant Pathol. 44, 527–539 (2015). https://doi.org/10.1007/s13313-015-0367-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13313-015-0367-2