Abstract
Retinoblastoma is a rare type of eye cancer of the retina that commonly occurs in early childhood and mostly affects the children before the age of 5. It occurs due to the mutations in the retinoblastoma gene (RB1) which inactivates both alleles of the RB1. RB1 was first identified as a tumor suppressor gene, which regulates cell cycle components and associated with retinoblastoma. Previously, genetic alteration was known as the major cause of its occurrence, but later, it is revealed that besides genetic changes, epigenetic changes also play a significant role in the disease. Initiation and progression of retinoblastoma could be due to independent or combined genetic and epigenetic events. Remarkable work has been done in understanding retinoblastoma pathogenesis in terms of genetic alterations, but not much in the context of epigenetic modification. Epigenetic modifications that silence tumor suppressor genes and activate oncogenes include DNA methylation, chromatin remodeling, histone modification and noncoding RNA-mediated gene silencing. Epigenetic changes can lead to altered gene function and transform normal cell into tumor cells. This review focuses on important epigenetic alteration which occurs in retinoblastoma and its current state of knowledge. The critical role of epigenetic regulation in retinoblastoma is now an emerging area, and better understanding of epigenetic changes in retinoblastoma will open the door for future therapy and diagnosis.
Similar content being viewed by others
References
Bishop JO, Madsen EC. Retinoblastoma: review of the current status. Surv Ophthalmol. 1975;19:342–66.
Knudson Jr AG. Mutation and cancer: statistical study of retinoblastoma. Proc Natl Acad Sci U S A. 1971;68:820–3.
Berger SL, Kouzarides T, Shiekhattar R, Shilatifard A. An operational definition of epigenetics. Genes Dev. 2009;23:781–3.
Dolinoy DC, Weidman JR, Jirtle RA. Epigenetic gene regulation: linking early developmental environment to adult disease. Reprod Toxicol. 2007;23:297–307.
Rodenhiser D, Mann M. Epigenetics and human disease: translating basic biology into clinical applications. CMAJ. 2006;174:341–8.
Gonzalo S. Epigenetic alterations in aging. J Appl Physiol. 2010;109:586–97.
Belinsky SA. Gene-promoter hypermethylation as a biomarker in lung cancer. Nat Rev Cancer. 2004;4:707–17.
Franklin TB, Mansuy IM. Epigenetic inheritance in mammals: evidence for the impact of adverse environmental effects. Neurobiol Dis. 2010;39:61–5.
Baylin SB, Herman JG, Graff JR, Vertino PM, Issa JP. Alterations in DNA methylation: a fundamental aspect of neoplasia. Adv Cancer Res. 1998;72:141–96.
Greger V, Passarge E, Höpping W, Messmer E, Horsthemke B. Epigenetic changes may contribute to the formation and spontaneous regression of retinoblastoma. Hum Genet. 1989;83:155–8.
Babenko OV, Zemliakova VV, Saakian SV, Brovkina AF, Strel’nikov VV, Zaletaev DV, et al. RB1 and CDKN2A functional defects resulting in retinoblastoma. Mol Biol (Mosk). 2002;36:777–83.
Livide G, Epistolato MC, Amenduni M, Disciglio V, Marozza A, Mencarelli MA, et al. Epigenetic and copy number variation analysis in retinoblastoma by MS-MLPA. Pathol Oncol Res. 2012;18:703–12.
Choy KW, Pang CP, Fan DS, Lee TC, Wang JH, Abramson DH, et al. Microsatellite instability and MLH1 promoter methylation in human retinoblastoma. Invest Ophthalmol Vis Sci. 2004;45:3404–9.
Choy KW, Pang CP, To KF, Yu CB, Ng JS, Lam DS. Impaired expression and promoter hypermethylation of O6-methylguanine-DNA methyltransferase in retinoblastoma tissues. Invest Ophthalmol Vis Sci. 2002;43:1344–9.
Harada K, Toyooka S, Maitra A, Maruyama R, Toyooka KO, Timmons CF, et al. Aberrant promoter methylation and silencing of the RASSF1A gene in pediatric tumors and cell lines. Oncogene. 2002;21:4345–9.
Maekawa M, Inomata M, Sasaki MS, Kaneko A, Ushiama M, Sugano K, et al. Electrophoretic variant of a lactate dehydrogenase isoenzyme and selective promoter methylation of the LDHA gene in a human retinoblastoma cell line. Clin Chem. 2002;48:1938–45.
Tosi GM, Trimarchi C, Macaluso M, La Sala D, Ciccodicola A, Lazzi S, et al. Genetic and epigenetic alterations of RB2/p130 tumor suppressor gene in human sporadic retinoblastoma: implications for pathogenesis and therapeutic approach. Oncogene. 2005;24:5827–36.
Cohen Y, Merhavi-Shoham E, Avraham RB, Frenkel S. Pe’er J, Goldenberg-Cohen N. Hypermethylation of CpG island loci of multiple tumor suppressor genes in retinoblastoma. Exp Eye Res. 2008;86:201–6.
Zhang J, Benavente CA, McEvoy J, Flores-Otero J, Ding L, Chen X, et al. A novel retinoblastoma therapy from genomic and epigenetic analyses. Nature. 2012;481:329–34.
Livide G, Epistolato MC, Amenduni M, Disciglio V, Marozza A, Mencarelli MA, et al. Epigenetic and copy number variation analysis in retinoblastoma by MS-MLPA. Pathol Oncol Res. 2012.
Beta M, Chitipothu S, Khetan V, Biswas J, Krishnakumar S. Hypermethylation of adenomatosis polyposis coli-2 and its tumor suppressor role in retinoblastoma. Curr Eye Res. 2015;40:719–28.
Philippeit C, Busch M, Dünker N. Epigenetic control of trefoil factor family (TFF) peptide expression in human retinoblastoma cell lines. Cell Physiol Biochem. 2014;34:1001–14.
Sakai T, Toguchida J, Ohtani N, Yandell DW, Rapaport JM, Dryja TP. Allele-specific hypermethylation of the retinoblastoma tumor-suppressor gene. Am J Hum Genet. 1991;48:880–8.
Ohtani-Fujita N, Fujita T, Aoike A, Osifchin NE, Robbins PD, Sakai T. CpG methylation inactivates the promoter activity of the human retinoblastoma tumor-suppressor gene. Oncogene. 1993;8:1063–7.
Greger V, Debus N, Lohmann D, Höpping W, Passarge E, Horsthemke B. Frequency and parental origin of hypermethylated RB1 alleles in retinoblastoma. Hum Genet. 1994;94:491–6.
Stirzaker C, Millar DS, Paul CL, Warnecke PM, Harrison J, Vincent PC, et al. Extensive DNA methylation spanning the Rb promoter in retinoblastoma tumors. Cancer Res. 1997;57:2229–37.
Joseph B, Mamatha G, Raman G, Shanmugam MP, Kumaramanickavel G. Methylation status of RB1 promoter in Indian retinoblastoma patients. Cancer Biol Ther. 2004;3:184–7.
Gonzalo S, García-Cao M, Fraga MF, Schotta G, Peters AH, Cotter SE, et al. Role of the RB1 family in stabilizing histone methylation at constitutive heterochromatin. Nat Cell Biol. 2005;7:420–8.
De La Rosa-Velázquez IA, Rincón-Arano H, Benítez-Bribiesca L, Recillas-Targa F. Epigenetic regulation of the human retinoblastoma tumor suppressor gene promoter by CTCF. Cancer Res. 2007;67:2577–85.
Choy KW, Lee TC, Cheung KF, Fan DS, Lo KW, Beaverson KL, et al. Clinical implications of promoter hypermethylation in RASSF1A and MGMT in retinoblastoma. Neoplasia. 2005;7:200–6.
Shivakumar L, Minna J, Sakamaki T, Pestell R, White MA. The RASSF1A tumor suppressor blocks cell cycle progression and inhibits cyclin D1 accumulation. Mol Cell Biol. 2002;22:4309–18.
Gozuacik D, Kimchi A. DAPK protein family and cancer. Autophagy. 2006;2:74–9.
Liu R, Gao L, GX L, Tang LS, Zhu XH, Wang J. Methylation status of RASSF1A and DAPK promoter in retinoblastoma. Zhonghua Yan Ke Za Zhi. 2009;45:631–5.
Indovina P, Acquaviva A, De Falco G, Rizzo V, Onnis A, Luzzi A, et al. Downregulation and aberrant promoter methylation of p16INK4A: a possible novel heritable susceptibility marker to retinoblastoma. J Cell Physiol. 2010;223:143–50.
Qu Y, Mu G, Wu Y, Dai X, Zhou F, Xu X, et al. Overexpression of DNA methyltransferases 1, 3a, and 3b significantly correlates with retinoblastoma tumorigenesis. Am J Clin Pathol. 2010;134:826–34.
Zhao JJ, Yang J, Lin J, Yao N, Zhu Y, Zheng J, et al. Identification of miRNAs associated with tumorigenesis of retinoblastoma by miRNA microarray analysis. Childs Nerv Syst. 2009;25:13–20.
Mu G, Liu H, Zhou F, Xu X, Jiang H, Wang Y, et al. Correlation of overexpression of HMGA1 and HMGA2 with poor tumor differentiation, invasion, and proliferation associated with let-7 down-regulation in retinoblastomas. Hum Pathol. 2010;41:493–502.
Xu X, Jia R, Zhou Y, Song X, Wang J, Qian G, et al. Microarray-based analysis: identification of hypoxia-regulated microRNAs in retinoblastoma cells. Int J Oncol. 2011;38:1385–93.
Conkrite K, Sundby M, Mukai S, Thomson JM, Mu D, Hammond SM, et al. MiR-17 ∼ 92 cooperates with RB pathway mutations to promote retinoblastoma. Genes Dev. 2011;25:1734–45.
To KH, Pajovic S, Gallie BL, Thériault BL. Regulation of p14ARF expression by miR-24: a potential mechanism compromising the p53 response during retinoblastoma development. BMC Cancer. 2012;12:69.
Martin J, Bryar P, Mets M, Weinstein J, Jones A, Martin A, et al. Differentially expressed miRNAs in retinoblastoma. Gene. 2013;(512):294–9.
Beta M, Venkatesan N, Vasudevan M, Vetrivel U, Khetan V, Krishnakumar S. Identification and insilico analysis of retinoblastoma serum microRNA profile and gene targets towards prediction of novel serum biomarkers. Bioinform Biol Insights. 2013;7:21–34.
Martin A, Jones A, Bryar PJ, Mets M, Weinstein J, Zhang G, et al. MicroRNAs-449a and -449b exhibit tumor suppressive effects in retinoblastoma. Biochem Biophys Res Commun. 2013;440:599–603.
Wang J, Wang X, Li Z, Liu H, Teng Y. MicroRNA-183 suppresses retinoblastoma cell growth, invasion and migration by targeting LRP6. FEBS J. 2014;281:1355–65.
Liu SS, Wang YS, Sun YF, Miao LX, Wang J, Li YS, et al. Plasma microRNA-320, microRNA-let-7e and microRNA-21 as novel potential biomarkers for the detection of retinoblastoma. Biomed Rep. 2014;2:424–8.
Lei Q, Shen F, Wu J, Zhang W, Wang J, Zhang L. MiR-101, downregulated in retinoblastoma, functions as a tumor suppressor in human retinoblastoma cells by targeting EZH2. Oncol Rep. 2014;32:261–9.
Beta M, Khetan V, Chatterjee N, Suganeswari G, Rishi P, Biswas J, et al. EpCAM knockdown alters microRNA expression in retinoblastoma-functional implication of EpCAM regulated miRNA in tumor progression. PLoS One. 2014;(9):e114800.
Wu X, Zeng Y, Wu S, Zhong J, Wang Y, Xu J. MiR-204, down-regulated in retinoblastoma, regulates proliferation and invasion of human retinoblastoma cells by targeting CyclinD2 and MMP-9. FEBS Lett. 2015;589:645–50.
Montoya V, Fan H, Bryar PJ, Weinstein JL, Mets MB, Feng G, et al. Novel miRNA-31 and miRNA-200a-mediated regulation of retinoblastoma proliferation. PLoS One. 2015;10:e0138366.
Shen F, Mo MH, Chen L, An S, Tan X, Fu Y, et al. MicroRNA-21 down-regulates Rb1 expression by targeting PDCD4 in retinoblastoma. J Cancer. 2014;5:804–12.
Lodygin D, Tarasov V, Epanchintsev A, Berking C, Knyazeva T, Körner H, et al. Inactivation of miR-34a by aberrant CpG methylation in multiple types of cancer. Cell Cycle. 2008;7:2591–600.
Jo DH, Kim JH, Park WY, Kim KW, Yu YS, Kim JH. Differential profiles of microRNAs in retinoblastoma cell lines of different proliferation and adherence patterns. J Ped. Hematol Oncol. 2011;33:529–53.
Kandalam MM, Beta M, Maheswari UK, Swaminathan S, Krishnakumar S. Oncogenic microRNA 17-92 cluster is regulated by epithelial cell adhesion molecule and could be a potential therapeutic target in retinoblastoma. Mol Vis. 2012;18:2279–87.
Jordan T, Hanson I, Zaletayev D, Hodgson S, Prosser J, Seawright A, et al. The human PAX6 gene is mutated in two patients with aniridia. Nat Genet. 1992;1:328–32.
Liu DK, Huang J, Xie M, Yu Y, Zhu S, Kang R, et al. MIR34A regulates autophagy and apoptosis by targeting HMGB1 in the retinoblastoma cell. Autophagy. 2014;10:442–52.
Jo DH, Kim JH, Cho CS, Cho YL, Jun HO, Yu YS, et al. STAT3 inhibition suppresses proliferation of retinoblastoma through down-regulation of positive feedback loop of STAT3/miR-17-92 clusters. Oncotarget. 2014;5:11513–25.
Esquela-Kerscher A, Slack FJ. Oncomirs: microRNAs with a role in cancer. Nat Rev Cancer. 2006;6:259–69.
Kent OA, Mendell JTA. Small piece in the cancer puzzle: microRNAs as tumor suppressors and oncogenes. Oncogene. 2006;25:6188–96.
Zhang B, Pan X, Cobb GP, Anderson TA. MicroRNAs as oncogenes and tumor suppressors. Dev Biol. 2007;302:1–12.
Dalgard CL, Gonzalez M, deNiro JE, O’Brien JM. Differential microRNA-34a expression and tumor suppressor function in retinoblastoma cells. Invest Ophthalmol Vis Sci. 2009;50:4542–51.
Thanos D, Maniatis T. The high mobility group protein HMG I (Y) is required for NF-kappa B-dependent virus induction of the human IFN-beta gene. Cell. 1992;7:777–89.
Fusco A, Fedele M. Roles of HMGA proteins in cancer. Nat Rev Cancer. 2007;7:899–910.
Ryan DG, Oliveira-Fernandes M, Lavker RM. MicroRNAs of the mammalian eye display distinct and overlapping tissue specificity. Mol Vis. 2006;12:1175–84.
Nittner D, Lambertz I, Clermont F, Mestdagh P, Köhler C, Nielsen SJ, et al. Synthetic lethality between Rb, p53 and Dicer or miR-17-92 in retinal progenitors suppresses retinoblastoma formation. Nat Cell Biol. 2012;14:958–65.
Wu L, Chen Z, Zhang J, Xing Y. Effect of miR-513a-5p on etoposide-stimulating B7-H1 expression in retinoblastoma cells. J Huazhong Univ Sci Technol Med Sci. 2012;32:601–6.
Zhang Y, Wu JH, Han F, Huang JM, Shi SY, Gu RD, et al. Arsenic trioxide induced apoptosis in retinoblastoma cells by abnormal expression of microRNA-376a. Neoplasma. 2013;60:247–53.
Wang J, Wang X, Wu G, Hou D, Hu Q. MiR-365b-3p, down-regulated in retinoblastoma, regulates cell cycle progression and apoptosis of human retinoblastoma cells by targeting PAX6. FEBS Lett. 2013;587:1779–86.
Ding Y, Wu M, Liu J, Wu C, Huang R, Zhu R, et al. Seed-targeting anti-miR-21 inhibiting malignant progression of retinoblastoma and analysis of their phosphorylation signaling pathways. Exp Eye Res. 2014;122:1–8.
Subramanian N, Kanwar JR, Kanwar RK, Krishnakumar S. Blocking the maturation of OncomiRNAs using pri-miRNA-17∼92 aptamer in retinoblastoma. Nucleic Acid Ther. 2015;25:47–52.
Cress WD, Seto E. Histone deacetylases, transcriptional control, and cancer. J Cell Physiol. 2000;184:1–16.
Gray SG, Teh BT. Histone acetylation/deacetylation and cancer: an “open” and “shut” case? Curr Mol Med. 2001;1:401–29.
Siddiqui H, Fox SR, Gunawardena RW, Knudsen ES. Loss of RB compromises specific heterochromatin modifications and modulates HP1alpha dynamics. J Cell Physiol. 2007;211:131–7.
Sherr CJ, McCormick F. The RB and p53 pathways in cancer. Cancer Cell. 2002;2:103–12.
Inoue Y, Kitagawa M, Taya Y. Phosphorylation of pRB at Ser612 by Chk1/2 leads to a complex between pRB and E2F-1 after DNA damage. EMBO J. 2007;26:2083–93.
Saddic LA, West LE, Aslanian A, Yates JR, Rubin SM, Gozani O, et al. Methylation of the retinoblastoma tumor suppressor by SMYD2. J Biol Chem. 2010;285:37733–40.
Chan HM, Krstic-Demonacos M, Smith L, Demonacos C, La Thangue NB. Acetylation control of the retinoblastoma tumour-suppressor protein. Nat Cell Biol. 2001;3:667–74.
Munro S, Khaire N, Inche A, Carr S, La Thangue NB. Lysine methylation regulates the pRb tumour suppressor protein. Oncogene. 2010;29:2357–67.
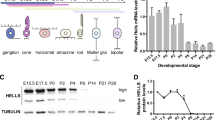
Benavente CA, Finkelstein D, Johnson DA, Marine JC, Ashery-Padan R, Dyer MA. Chromatin remodelers HELLS and UHRF1 mediate the epigenetic deregulation of genes that drive retinoblastoma tumor progression. Oncotarget. 2014;5:9594–608.
Khan M, Walters LL, Li Q, Thomas DG, Miller JM, Zhang Q, et al. Characterization and pharmacologic targeting of EZH2, a fetal retinal protein and epigenetic regulator, in human retinoblastoma. Lab Investig. 2015;95:1278–90.
Chakraborty S, Khare S, Dorairaj SK, et al. Identification of genes associated with tumorigenesis of retinoblastoma by microarray analysis. Genomics. 2007;90:344–53.
Akbari MT, Naderi A, Saremi L, Sayad A, Irani S, Ahani A. Methionine synthase A2756G variation is associated with the risk of retinoblastoma in Iranian children. Cancer Epidemiol. 2015;39:1023–5.
Dunaief JL, Strober BE, Guha S, Khavari PA, Alin K, Luban J, et al. The retinoblastoma protein and BRG1 form a complex and cooperate to induce cell cycle arrest. Cell. 1994;79:119–30.
Aldiri I, Ajioka I, Xu B, Zhang J, Chen X, Benavente C, et al. Brg1 coordinates multiple processes during retinogenesis and is a tumor suppressor in retinoblastoma. Development. 2015;142:4092–106.
Acknowledgments
We sincerely acknowledge the support of the Department of Biotechnology, Govt. of India.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Singh, U., Malik, M.A., Goswami, S. et al. Epigenetic regulation of human retinoblastoma. Tumor Biol. 37, 14427–14441 (2016). https://doi.org/10.1007/s13277-016-5308-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-016-5308-3