Abstract
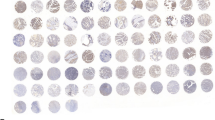
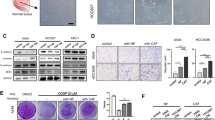
Chromosome 6 open reading frame 106 (C6orf106) is a newly discovered protein; its expression and clinical pathological significance in human tumors remains unclear. Immunohistochemistry, Western blot, and immunofluorescence were performed to assess C6orf106 expression in non-small cell lung cancer (NSCLC). In addition, the relationships between subcellular localization and clinical pathological factors were analyzed. Through C6orf106 overexpression and repression, respectively, in lung cancer cell lines, we explored the effect of this molecule on NSCLC invasion abilities. C6orf106 was highly expressed in the cytoplasm of lung cancer tissue cells (60.4 %, 75/124), compared with adjacent normal lung tissues (15.2 %, 7/46, p < 0.001). In addition, its expression was positively correlated with differentiation (p = 0.001), TNM stage (p = 0.011), lymph node metastasis (p = 0.018), and poor prognosis (p = 0.006). Furthermore, C6orf106 overexpression enhanced NSCLC cell invasion. Moreover, C6orf106 was shown to increase vimentin expression, while decreasing E-cadherin and P120ctn. C6orf106 is highly expressed in NSCLC and correlates with clinical and pathological factors, as well as poor prognosis. C6orf106 promotes invasion in NSCLC cells. Finally, C6orf106 upregulates vimentin, and downregulates E-cadherin and P120ctn.





Similar content being viewed by others
References
Mungall AJ, Palmer SA, Sims SK, Edwards CA, Ashurst JL, Wilming L, et al. The DNA sequence and analysis of human chromosome 6. Nature. 2003;425(6960):805–11.
Wang J, Huo K, Ma L, Tang L, Li D, Huang X, et al. Toward an understanding of the protein interaction network of the human liver. Mol Syst Biol. 2011;7:536.
Ménard S, Tagliabue E, Colnaghi MI. The 67 kDa laminin receptor as a prognostic factor in human cancer. Breast Cancer Res Treat. 1998;52(1–3):137–45.
Montuori N, Selleri C, Risitano AM, Raiola AM, Ragno P, Del Vecchio L, et al. Expression of the 67-kDa laminin receptor in acute myeloid leukemia cells mediates adhesion to laminin and is frequently associated with monocytic differentiation. Clin Cancer Res. 1999;5(6):1465–72.
Montuori N, Müller F, De Riu S, et al. Laminin receptors in differentiated thyroid tumors: restricted expression of the 67-kilodalton laminin receptor infollicular carcinoma cells. J Clin Endocrinol Metab. 1999;84(6):2086–92.
Zhang SC, Jin W, Liu H, Jin MJ, Chen ZX, Ding ZY, et al. Chen K.RPSA gene mutants associated with risk of colorectal cancer among the Chinese population. Asian Pac J Cancer Prev. 2013;14(12):7127–31.
Vande Broek I, Vanderkerken K, De Greef C, Asosingh K, Straetmans N, Van Camp B, et al. Laminin-1-induced migration of multiple myeloma cells involves the high-affinity 67 kD laminin receptor. Br J Cancer. 2001;85(9):1387–95.
Selleri C, Ragno P, Ricci P, Visconte V, Scarpato N, Carriero MV, et al. The metastasis-associated 67-kDa laminin receptor is involved in G-CSF-induced hematopoietic stem cell mobilization. Blood. 2006;108(7):2476–84.
Berno V, Porrini D, Castiglioni F, Campiglio M, Casalini P, Pupa SM, et al. The 67 kDa laminin receptor increases tumor aggressiveness by remodeling laminin-1. Endocrinol Relat Cancer. 2005;12(2):393–406.
Kumazoe M, Sugihara K, Tsukamoto S, Huang Y, Tsurudome Y, Suzuki T, et al. Tachibana H.67-kDa laminin receptor increases cGMP to induce cancer-selective apoptosis. J Clin Invest. 2013;123(2):787–99.
Kim DG, Choi JW, Lee JY, Kim H, Oh YS, Lee JW, et al. Interaction of two translational components, lysyl-tRNA synthetase and p40/37LRP, in plasma membrane promotes laminin-dependent cell migration. FASEB J. 2012;26(10):4142–59.
Ardini E, Sporchia B, Pollegioni L, Modugno M, Ghirelli C, Castiglioni F, et al. Identification of a novel function for 67-kDa laminin receptor: increase in laminin degradation rate and release of motility fragments. Cancer Res. 2002;62(5):1321–5.
Travis WD, Brambilla E, Müller-Hermelink HK, Harris CC. Pathology and genetics: tumours of the lung, pleura, thymus and heart. Lyon: IARC; 2004.
Goldstraw P. Updated staging system for lung cancer. Surg Oncol Clin N Am. 2011;20:655–66.
Givant-Horwitz V, Davidson B, Reich R. Laminin-induced signaling in tumor cells: the role of the M(r) 67,000 laminin receptor. Cancer Res. 2004;64(10):3572–9.
Sou PW, Delic NC, Halliday GM, Lyons JG. Snail transcription factors in keratinocytes: enough to make your skin crawl. Int J Biochem Cell Biol. 2010;42(12):1940–4.
Bonavida B, Baritaki S. The novel role of yin yang 1 in the regulation of epithelial to mesenchymal transition in cancer via the dysregulated NF-κB/snail/YY1/RKIP/PTEN circuitry. Crit Rev Oncog. 2011;16(3–4):211–26.
Tania M, Khan MA, Fu J. Epithelial to mesenchymal transition inducing transcription factors and metastatic cancer. Tumour Biol. 2014;35(8):7335–42.
Zhao H, Zhao Y, Jiang G, Zhang X, Zhang Y, Dong Q, Luan L, Papavassiliou P, Wang E, Wang E. Dishevelled-3 activates p65 to upregulate p120-catenin transcription via a p38-dependent pathway in non-small cell lung cancer. Mol Carcinog. 2014.
Scheiman J, Tseng JC, Zheng Y, Meruelo D. Multiple functions of the 37/67-kd laminin receptor make it a suitable target for novel cancer gene therapy. Mol Ther. 2010;18(1):63–74.
Acknowledgments
We are grateful to Dr. Hiroshi Kijima (Department of Pathology and Bioscience, Hirosaki University Graduate School of Medicine, Japan) for providing the cell lines mentioned in this manuscript. This work was supported by grants from the National Natural Science Foundation of China (no. 81272606 to Enhua Wang, no. 81302312 to Yang Liu, no. 81472805 to Yuan Miao, no. 81402369 to Guiyang Jiang, and no. 81301837 to Juanhan Yu).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhang, X., Miao, Y., Yu, X. et al. C6orf106 enhances NSCLC cell invasion by upregulating vimentin, and downregulating E-cadherin and P120ctn. Tumor Biol. 36, 5979–5985 (2015). https://doi.org/10.1007/s13277-015-3274-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-015-3274-9