Abstract
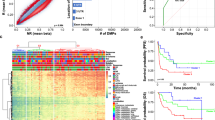
CpG island methylator phenotype (CIMP) involves methylation targeted toward the promoters of multiple genes. We determined a methylation profile of tumor-related genes in serum of sporadic breast cancer (SBC). The multigene methylation was examined by methylation-specific polymerase chain reaction assay in serum of 50 SBCs and 50 paired nontumors, and CIMP+ was defined as having three genes that are concordantly methylated. The methylation frequency of ten genes in serum of 50 SBCs varied from 10% in FHIT to 74% in RASSF1A. The methylation status of RASSF1A, BRCA1, p16, CDH1, ER, RARβ2, APC, and DAPK was significantly correlated with SBC and nontumor serum (P < 0.05). Methylation of at least one gene was found in 92% SBC; CIMP was more frequent in SBC than nontumor serum (P < 0.001). There was a significant association between CIMP and methylation of RASSF1A, BRCA1, p16, CDH1, ER, RARβ2, APC, and DAPK (P < 0.05); the methylation link profile of CDH1, RASSF1A, BRCA1, and RARβ2 as breast cancer marker may contribute high sensitivity (90%) and specificity (88%). ER and RARβ2 methylation was associated with elevated serum CA153 levels in 39 SBC samples with CIMP+ (P < 0.05). Multivariate analysis showed that living area of patients was found to provide independent prognostic information associated with a relative risk of tumor recurrence of 5.3. Multigene-specific methylation profile in serum was association with the recurrence risk of rural SBC, and positive correlation of CIMP can serve as a promising molecular marker of SBC.



Similar content being viewed by others
References
Goldhirsch A, Glick JH, Gelber RD, et al. Meeting highlights: international consensus panel on the treatment of primary breast cancer. Seventh international conference on adjuvant therapy of primary breast cancer. J Clin Oncol. 2001;19:3817–27.
Early Breast Cancer Trialists' Collaborative Group. Polychemotherapy for early breast cancer: an overview of the randomised trials. Early breast cancer trialists' collaborative group. Lancet. 1998;352:930–42.
Early Breast Cancer Trialists' Collaborative Group. Tamoxifen for early breast cancer: an overview of the randomised trials. Early breast cancer trialists' collaborative group. Lancet. 1998;351:1451–67.
Hayes DF, Isaacs C, Stearns V. Prognostic factors in breast cancer: current and new predictors of metastasis. J Mammary Gland Biol Neoplasia. 2001;6:375–92.
Couch FJ, Weber BL. Breast cancer. In: Scriver CR, Beaudet AL, Sly WS, et al., editors. The metabolic and molecular basis of human disease. New York: McGraw-Hill; 2001. p. 999–1031.
Leitch AM. Breast cancer screening: success amid conflict. Surg Oncol Clin N Am. 1999;8:657–72.
Dulaimi E, Hillinck J, Ibanez CI, et al. Tumor suppressor gene promoter hypermethylation in serum of breast cancer patients. Clin Cancer Res. 2004;15(10):6189–93.
Müller HM, Fiegl H, Widschwendter A, et al. Prognostic DNA methylation marker in serum of cancer patients. Ann N Y Acad Sci. 2004;1022:44–9.
Hu XC, Wong IH, Chow LW. Tumor-derived aberrant methylation in plasma of invasive ductal breast cancer patients: clinical implications. Oncol Rep. 2003;10(6):1811–5.
Umbricht CB, Evron E, Gabrielson E, et al. Hypermethylation of 14-3-3 sigma (stratifin) is an early event in breast cancer. Oncogene. 2001;20:3348–53.
Evron E, Dooley WC, Umbricht CB, et al. Detection of breast cancer cells in ductal lavage fluid by methylation-specific PCR. Lancet. 2001;357:1335–6.
Evron E, Umbricht CB, Korz D, et al. Loss of cyclin D2 expression in the majority of breast cancers is associated with promoter hypermethylation. Cancer Res. 2001;61:2782–7.
Ulivi P, Zoli W, Calistri D, et al. p16INK4A and CDH13 hypermethylation in tumor and serum of non-small cell lung cancer patients. J Cell Physiol. 2006;206(3):611–5.
Belinsky SA, Nikula KJ, Palmisano WA, et al. Aberrant methylation of p16(INK4a) is an early event in lung cancer and a potential biomarker for early diagnosis. PNAS. 1998;95:11891–6.
Tsou JA, Hagen JA, Carpenter CL. DNA methylation analysis: a powerful new tool for lung cancer diagnosis. Oncogene. 2002;21(35):5450–61.
Usadel H, Brabender J, Danenberg KD. Quantitative adenomatous polyposis coli promoter methylation analysis in tumor tissue, serum, and plasma DNA of patients with lung cancer. Cancer Res. 2002;62(2):371–5.
Barault L, Charon-Barra C, Jooste V. Hypermethylator phenotype in sporadic colon cancer: study on a population-based series of 582 cases. Cancer Res. 2008;68(20):8541–6.
Esteller M, Sparks A, Toyota M, et al. Analysis of adenomatous polyposis coli promoter hypermethylation in human cancer. Cancer Res. 2000;60:4366–71.
Ngan RK, Lau WH, Yip TT, et al. Remarkable application of serum EBV EBER-1 in monitoring response of nasopharyngeal cancer patients to salvage chemotherapy. Ann N Y Acad Sci. 2001;945:73–9.
Lo YM. Prognostic implication of pretreatment plasma/serum concentration of Epstein-Barr virus DNA in nasopharyngeal carcinoma. Biomed Pharmacother. 2001;55:362–5.
Sidransky D, Von Eschenbach A, Tsai YC, et al. Identification of p53 gene mutations in bladder cancers and urine samples. Science. 1991;252:706–9.
Muller HM, Widschwendter A, Fiegl H, et al. DNA methylation in serum of breast cancer patients: an independent prognostic marker. Cancer Res. 2003;63:7641–5.
Jeronimo C, Costa I, Martins MC, et al. Detection of gene promoter hypermethylation in fine needle washings from breast lesions. Clin Cancer Res. 2003;9:3413–7.
Hoque MO, Begum S, Topaloglu O, Jeronimo C, et al. Quantitative detection of promoter hypermethylation of multiple genes in the tumor, urine, and serum DNA of patients with renal cancer. Cancer Res. 2004;64:5511–17.
Kawakami K, Brabender J, Lord RV, et al. Hypermethylated APC DNA in plasma and prognosis of patients with esophageal adenocarcinoma. J Natl Cancer Inst. 2000;92:1805–11.
Topaloglu O, Hoque MO, Tokumaru Y, et al. Detection of promoter hypermethylation of multiple genes in the tumor and bronchoalveolar lavage of patients with lung cancer. Clin Cancer Res. 2004;10:2284–8.
Yang HJ, Liu VW, Wang Y, et al. Detection of hypermethylated genes in tumor and plasma of cervical cancer patients. Gynecol Oncol. 2004;93:435–40.
Usadel H, Brabender J, Danenberg KD, et al. Quantitative adenomatous polyposis coli promoter methylation analysis in tumor tissue, serum, and plasma DNA of patients with lung cancer. Cancer Res. 2002;62:371–5.
Toyota M, Ahuja N, Ohe-Toyota M, et al. CpG island methylator phenotype in colorectal cancer. Proc Natl Acad Sci USA. 1999;96(15):8681–6.
Ogino S, Cantor M, Kawasaki T, et al. CpG island methylator phenotype (CIMP) of colorectal cancer is best characterised by quantitative DNA methylation analysis and prospective cohort studies. Gut. 2006;55(7):1000–6.
Zhang C, Li Z, Cheng Y, et al. CpG island methylator phenotype association with elevated serum alpha-fetoprotein level in hepatocellular carcinoma. Clin Cancer Res. 2007;13(3):944–52.
Toyota M, Ahuja N, Ohe-Toyota M, Herman JG, Baylin SB, Issa JP. CpG island methylator phenotype in colorectal cancer. Proc Natl Acad Sci USA. 1999;96:8681–6.
Toyota M, Ahuja N, Suzuki H, et al. Aberrant methylation in gastric cancer associated with the CpG island methylator phenotype. Cancer Res. 1999;59:5438–42.
Weisenberger DJ, Siegmund KD, Campan M, et al. CpG island methylator phenotype underlies sporadic microsatellite instability and is tightly associated with BRAF mutation in colorectal cancer. Nat Genet. 2006;38:787–93.
Sambrook J, Russell DW. Molecular cloning. A laboratory manual. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press; 2001.
Herman JG, Graff JR, Myohanen S, Nelkin BD, Baylin SB. Methylation-specific PCR: a novel PCR assay for methylation status of CpG islands. Proc Natl Acad Sci USA. 1996;93:9821–6.
Grunau C, Clark SJ, Rosenthal A. Bisulifite genomic sequencing: systematic investigation of critical experimental parameters. Nucleic Acids Res. 2001;29:e65.
Esteller M, Silva JM, Dominguez G, Bonilla F, Xavier MG, Lerma E, et al. Promoter hypermethylation and BRCA1 inactivation in sporadic breast and ovarian tumors. J Natl Cancer Inst (Bethesda). 2000;92:564–9.
Sabine ZM, Fong KM, Maitra A, et al. 50 CpG island methylation of the FHIT gene is correlated with loss of gene expression in lung and breast cancer. Cancer Res. 2001;61:3581–5.
Katzenellenbogen RA, Baylin SB, Herman JG. Hypermethylation of the DAP-kinase CpG island is a common alteration in B-cell malignancies. Blood. 1999;93:4347–53.
Burbee DG, Forgacs E, Zochbauer-Muller S, et al. Epigenetic inactivation of RASSF1A in lung and breast cancers and malignant phenotype suppression. J Natl Cancer Inst. 2001;93:691–9.
Widschwendter M, Berger J, Hermann M, et al. Methylation and silencing of the retinoic acid receptor-beta2 gene in breast cancer. J Natl Cancer Inst. 2001;92:826–32.
Corn PG, Smith BD, Ruckdeschel ES, Douglas D, Baylin SB, Herman JG. E-cadherin expression is silenced by 5 CpG island methylation in acute leukemia. Clin Cancer Res. 2000;6:4243–8.
Mahmood A, Yulan C, Magno RM. Promoter methylation regulates helicobacter pylori-stimulated cyclooxygenase-2 expression in gastric epithelial cells. Cancer Res. 2001;61:2399–403.
Mori T, Martinez SR, O'Day SJ. Estrogen receptor-α methylation predicts melanoma progression. Cancer Res. 2006;66:6692–8.
Jones PA, Takai D. The role of DNA methylation in mammalian epigenetics. Science (Wash DC). 2001;293:1068–70.
Maike W, Andreas H, Andrea B, et al. Methylation of serum DNA is an independent prognostic marker in colorectal cancer. Clin Cancer Res. 2006;12:7347–52.
Misawa K, Ueda Y, Kanazawa T, et al. Epigenetic inactivation of galanin receptor 1 in head and neck cancer. Clin Cancer Res. 2008;14:7604–13.
Euhus DM, Bu D, Milchgrub S, et al. DNA methylation in benign breast epithelium in relation to age and breast cancer risk. Cancer Epidemiol Biomark Prev. 2008;17:1051–9.
Anders R, Jörg T, Solvang HK, et al. GSTP1 promoter haplotypes affect DNA methylation levels and promoter activity in breast carcinomas. Cancer Res. 2008;68:5562–71.
Oue N, Mitani Y, Motoshita J, et al. Accumulation of DNA methylation is associated with tumor stage in gastric cancer. Cancer. 2006;106:1250–9.
Eads CA, Lord RV, Wickramasinghe K, et al. Epigenetic patterns in the progression of esophageal adenocarcinoma. Cancer Res. 2001;61:3410–8.
Yamashita K, Dai T, Dai Y, Yamamoto F, Perucho M. Genetics supersedes epigenetics in colon cancer phenotype. Cancer Cell. 2003;4:121–31.
Van der Auwera I, Elst HJ, Van Laere SJ, et al. The presence of circulating total DNA and methylated genes is associated with circulating tumour cells in blood from breast cancer patients. Br J Cancer. 2009;100(8):1277–86.
Hawkins N, Norrie M, Cheong K, et al. CpG island methylation in sporadic colorectal cancers and its relationship to microsatellite instability. Gastroenterology. 2002;122:1376–87.
Samowitz WS, Albertsen H, Herrick J, et al. Evaluation of a large, population-based sample supports a CpG island methylator phenotype in colon cancer. Gastroenterology. 2005;129:837–45.
Grants provided by Science Funds for Distinguished Young Scholar of Hubei Province (2008154), Natural Science Foundation of Hubei Province (2007ABA371), Science Funds from Hubei Provincial Department of Education (Q20082407), and Innovation Foundation for Young Scholars of Yunyang Medical College (CXX200805).
Author information
Authors and Affiliations
Corresponding author
Additional information
Feng Jing and Wang Yuping contributed equally to this work.
Rights and permissions
About this article
Cite this article
Jing, F., Yuping, W., Yong, C. et al. CpG island methylator phenotype of multigene in serum of sporadic breast carcinoma. Tumor Biol. 31, 321–331 (2010). https://doi.org/10.1007/s13277-010-0040-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-010-0040-x