Abstract
Background
Although cytoreductive surgery followed by adjuvant chemotherapy is effective as a standard treatment for early-stage ovarian cancer, the majority of ovarian cancer cases are diagnosed at the advanced stages with dissemination to the peritoneal cavity, leading to a poor prognosis. Therefore, it is crucial to understand the cellular and molecular mechanisms underlying metastasis and identify novel therapeutic targets.
Objective
In this study, we aimed to elucidate the mechanisms underlying gene expression alterations during the acquisition of metastatic potential and characterize the metastatic subpopulations within ovarian cancer cells.
Methods
We conducted single-cell RNA sequencing of two human ovarian cancer cell lines: SKOV-3 and SKOV-3-13, a highly metastatic subclone of SKOV-3. Suppression of NFE2L1 expression was performed through siRNA-mediated knockdown and CRISPR-Cas9-mediated knockout.
Results
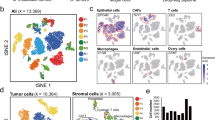
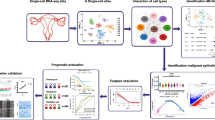
Clustering and pseudotime trajectory analysis revealed pro-metastatic subpopulation within these cells. Furthermore, gene set enrichment analysis and prognosis analysis indicated that NFE2L1 could be a key transcription factor in the acquisition of metastasis potential. Inhibition of NFE2L1 significantly reduced migration and viability of both cells. In addition, NFE2L1 knockout cells exhibited significantly reduced tumor growth in a mouse xenograft model, recapitulating in silico and in vitro results.
Conclusion
The results presented in this study deepen our understanding of the molecular pathogenesis of ovarian cancer metastasis with the ultimate goal of developing treatments targeting pro-metastatic subclones prior to metastasis.
Access this article
We’re sorry, something doesn't seem to be working properly.
Please try refreshing the page. If that doesn't work, please contact support so we can address the problem.




Similar content being viewed by others
References
Bacac M, Stamenkovic I (2008) Metastatic cancer cell. Annu Rev Pathol 3:221–247
Bast RC, Han CY, Lu Z, Lu KH (2021) Next steps in the early detection of ovarian cancer. Commun Med (Lond) 1
Forcina GC, Pope L, Murray M, Dong W, Abu-Remaileh M, Bertozzi CR, Dixon SJ (2022) Ferroptosis regulation by the NGLY1/NFE2L1 pathway. Proc Natl Acad Sci U S A 119:e2118646119
Galland F, Lacroix L, Saulnier P, Dessen P, Meduri G, Bernier M, Gaillard S, Guibourdenche J, Fournier T, Evain-Brion D et al (2010) Differential gene expression profiles of invasive and non-invasive non-functioning pituitary adenomas based on microarray analysis. Endocr Relat Cancer 17:361–371
Giornelli GH (2016) Management of relapsed ovarian cancer: a review. Springerplus 5:1197
Guo J, Han X, Li J, Li Z, Yi J, Gao Y, Zhao X, Yue W (2023) Single-cell transcriptomics in ovarian cancer identify a metastasis-associated cell cluster overexpressed RAB13. J Transl Med 21:254
Hanzelmann S, Castelo R, Guinney J (2013) GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics 14:7
Hao Y, Hao S, Andersen-Nissen E, Mauck WM 3rd, Zheng S, Butler A, Lee MJ, Wilk AJ, Darby C, Zager M et al (2021) Integrated analysis of multimodal single-cell data. Cell 184:3573–3587 e3529
Hashimoto K, Man S, Xu P, Cruz-Munoz W, Tang T, Kumar R, Kerbel RS (2010) Potent preclinical impact of metronomic low-dose oral topotecan combined with the antiangiogenic drug pazopanib for the treatment of ovarian cancer. Mol Cancer Ther 9:996–1006
Kan T, Zhang S, Zhou S, Zhang Y, Zhao Y, Gao Y, Zhang T, Gao F, Wang X, Zhao L et al (2022) Single-cell RNA-seq recognized the initiator of epithelial ovarian cancer recurrence. Oncogene 41:895–906
Kim HM, Han JW, Chan JY (2016) Nuclear factor Erythroid-2 like 1 (NFE2L1): structure, function and regulation. Gene 584:17–25
Kotschi S, Jung A, Willemsen N, Ofoghi A, Proneth B, Conrad M, Bartelt A (2022) NFE2L1-mediated proteasome function protects from ferroptosis. Mol Metab 57:101436
La Manno G, Soldatov R, Zeisel A, Braun E, Hochgerner H, Petukhov V, Lidschreiber K, Kastriti ME, Lonnerberg P, Furlan A et al (2018) RNA velocity of single cells. Nature 560:494–498
Lee J, Park J, Choi C (2014) Identification of phenotype deterministic genes using systemic analysis of transcriptional response. Sci Rep 4:4413
Lee YK, Kwon SM, Lee EB, Kim GH, Min S, Hong SM, Wang HJ, Lee DM, Choi KS, Park TJ et al (2020) Mitochondrial respiratory defect enhances Hepatoma Cell Invasiveness via STAT3/NFE2L1/STX12 Axis. Cancers (Basel) 12
Lessi F, Scatena C, Aretini P, Menicagli M, Franceschi S, Naccarato AG, Mazzanti CM (2019) Molecular profiling of microinvasive breast cancer microenvironment progression. J Transl Med 17:187
Li YF, Altman RB (2018) Systematic target function annotation of human transcription factors. BMC Biol 16:4
Li Y, Sun R, Fang X, Ruan Y, Hu Y, Wang K, Liu J, Wang H, Pi J, Chen Y et al (2022) Long-isoform NFE2L1 silencing inhibits acquisition of malignant phenotypes induced by arsenite in human bronchial epithelial cells. Ecotoxicol Environ Saf 232:113268
Malysheva V, Mendoza-Parra MA, Saleem MA, Gronemeyer H (2016) Reconstruction of gene regulatory networks reveals chromatin remodelers and key transcription factors in tumorigenesis. Genome Med 8:57
Matulonis UA, Sood AK, Fallowfield L, Howitt BE, Sehouli J, Karlan BY (2016) Ovarian cancer. Nat Rev Dis Primers 2:16061
Park J, Lee J, Choi C (2015) Evaluation of drug-targetable genes by defining modes of abnormality in gene expression. Sci Rep 5:13576
Pokhriyal R, Hariprasad R, Kumar L, Hariprasad G (2019) Chemotherapy Resistance in Advanced Ovarian Cancer Patients. Biomark Cancer 11:1179299x19860815
Schram AM, Chang MT, Jonsson P, Drilon A (2017) Fusions in solid tumours: diagnostic strategies, targeted therapy, and acquired resistance. Nat Rev Clin Oncol 14:735–748
Shih AJ, Menzin A, Whyte J, Lovecchio J, Liew A, Khalili H, Bhuiya T, Gregersen PK, Lee AT (2018) Identification of grade and origin specific cell populations in serous epithelial ovarian cancer by single cell RNA-seq. PLoS ONE 13:e0206785
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, Bray F (2021) Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and Mortality Worldwide for 36 cancers in 185 countries. CA Cancer J Clin 71:209–249
Tirosh I, Izar B, Prakadan SM, Wadsworth MH 2nd, Treacy D, Trombetta JJ, Rotem A, Rodman C, Lian C, Murphy G et al (2016) Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352:189–196
Trapnell C, Cacchiarelli D, Grimsby J, Pokharel P, Li S, Morse M, Lennon NJ, Livak KJ, Mikkelsen TS, Rinn JL (2014) The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat Biotechnol 32:381–386
Waku T, Katayama H, Hiraoka M, Hatanaka A, Nakamura N, Tanaka Y, Tamura N, Watanabe A, Kobayashi A (2020) NFE2L1 and NFE2L3 Complementarily maintain basal proteasome activity in Cancer cells through CPEB3-Mediated translational repression. Mol Cell Biol 40
Wang Y, Shan X, Dong H, Li M, Yue Y (2022) Prediction for 2-year mortality of metastatic ovarian cancer patients based on surveillance, epidemiology, and end results database. Front Surg 9:974536
Yang L, Xie HJ, Li YY, Wang X, Liu XX, Mai J (2022) Molecular mechanisms of platinum–based chemotherapy resistance in ovarian cancer (review). Oncol Rep 47
Yu G, Wang LG, Han Y, He QY (2012) clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16:284–287
Zheng P, Lin Z, Ding Y, Duan S (2023) Targeting the dynamics of cancer metabolism in the era of precision oncology. Metabolism 145:155615
Funding
This study was supported by the National Research Foundation of Korea (RS-2022-00165497, NRF-2019R1A5A2027588, NRF-2020R1H1A1102285).
Author information
Authors and Affiliations
Contributions
Conceptualization, JP, SHL, and Y-JC; Methodology, JP, Y-SK, SZ, DK, and SS; Software, JP and Y-SK; Validation, JP, Y-SK, and DK; Formal Analysis, JP, Y-SK, SZ, and DK; Investigation, JP, SZ, and DK; Resources, SS, SHL, and Y-JC; Data Curation, JP, Y-SK, and SZ; Writing – Original Draft, JP; Writing – Review & Editing, JP and Y-JC; Visualization, JP and SZ; Project Administration, SHL and Y-JC; Funding Acquisition, JP and Y-JC.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Park, J., Kim, YS., Zhang, S. et al. Single-cell RNA sequencing reveals a pro-metastatic subpopulation and the driver transcription factor NFE2L1 in ovarian cancer cells. Genes Genom 45, 1107–1115 (2023). https://doi.org/10.1007/s13258-023-01418-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-023-01418-1