Abstract
Background
A large proportion of eukaryote nuclear genomes is composed of repetitive DNA. Tracing the dynamics of repetitive elements in the genomes of related taxa can reveal important information about their phylogenic relationships as well as traits that have become distinct to a lineage.
Objective
Study the genomic abundance and chromosomal location of repetitive DNA in Capsicum annuum L. to understand the repeat dynamics.
Method
We quantified repeated DNA content in the C. annuum genome using the RepeatExplorer pipeline.
Results
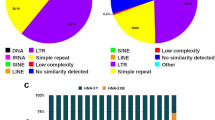
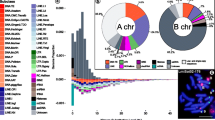
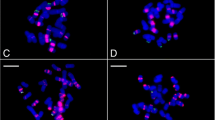
About 42% of the C. annuum genome dataset comprised repetitive elements. Of these, 0.011, 0.98, 3.09, and 0.024% represented high and low confidence satellite repeats, putative long-terminal repeats (LTRs), and rDNA sequences, respectively. One novel high confidence 167-bp satellite repeat with a genomic proportion of 0.011%, Ca167TR, was identified. Furthermore, FISH with Ca167TR on metaphase chromosomes of C. annuum revealed signals in the subtelomeric regions of the short and long arms of chromosome 3 and 4, respectively.
Conclusion
Further understanding of the origin and associated functions of Ca167TR and other repeats in Capsicum will give us insights into the genomic relationships and functions of the genome.


Similar content being viewed by others
References
Allen G, Flores-Vergara M, Krasynanski S, Kumar S, Thompson W (2006) A modified protocol for rapid DNA isolation from plant tissues using cetyltrimethylammonium bromide. Nat Protoc 1:2320–2325
Cai Z, Liu H, He Q, Pu M, Chen J, Lai J, Li X, Jin W (2014) Differential genome evolution and speciation of Coix lacryma-jobi L. and Coix aquatica Roxb. hybrid guangxi revealed by repetitive sequence analysis and fine karyotyping. BMC Genomics 15:1025–1040
Castro-Concha LA, Tuyub-Che J, Moo-Mukul A, Vazquez-Flota FA, Miranda-Ham ML (2014) Antioxidant capacity and total phenolic content in fruit tissues from accessions of Capsicum chinense Jacq. (Habanero Pepper) at different stages of ripening. Sci World J 2014:809073
Cheema S, Pant M (2013) Karyotype analysis of seven cultivated varieties of Capsicum annuum L. Caryologia 66:70–75
Consortium TG (2012) The tomato genome sequence provides insights into fleshy fruit evolution. Nature 485:635–641
Dixit P (1931) A cytological study of Capsicum annuum. Indian J Agric Sci 1:419–433
Galian JA, Rosato M, Rossello JA (2012) Early evolutionary colocalization of the nuclear ribosomal 5S and 45S gene families in seed plants: evidence from the living fossil gymnosperm Ginkgo biloba. Heredity 108:640–646
Galindo-González L, Mhiri C, Deyholos MK, Grandbastien M-A (2017) LTR-retrotransposons in plants: engines of evolution. Gene 626:14–25
Garrido-Ramos MA (2015) Satellite DNA in plants: more than just rubbish. Cytogenet Genome Res 146:153–170
Gupta PK, Varshney RK (2000) The development and use of microsatellite markers for genetic analysis and plant breeding with emphasis on bread wheat. Euphytica 113:163–185
Huskins C, La-Cour L (1930) Chromosome numbers in Capsicum. Am Nat 64:382–384
Hwang YJ, Lee SN, Song KA, Ryu KB, Ryu KH, Kim HH (2010) Karyotype analyses of genetically modified (GM) and Non-GM hot peppers by conventional staining and FISH method. Hortic Environ Biotechnol 51:525–530
Jia J, Zhao S, Kong X, Li Y, Zhao G, He W, Appels R, Pfeifer M, Tao Y, Zhang X et al (2013) Aegilops tauschii draft genome sequence reveals a gene repertoire for wheat adaptation. Nature 496:91–95
Kalendar R, Tanskanen J, Immonen S, Nevo E, Schulman AH (2000) Genome evolution of wild barley (Hordeum spontaneum) by BARE-1 retrotransposon dynamics in response to sharp microclimatic divergence. Proc Natl Acad Sci USA 97:6603–6607
Kim S, Park M, Yeom S-I, Kim Y-M, Lee JM, Lee H-A, Seo E, Choi J, Cheong K, Kim K-T (2014) Genome sequence of the hot pepper provides insights into the evolution of pungency in Capsicum species. Nat Genet 46:270–278
Kwon J-K, Kim B-D (2009) Localization of 5S and 25S rRNA genes on somatic and meiotic chromosomes in Capsicum species of chili pepper. Mol Cells 27:205–209
Li S-F, Su T, Cheng G-Q, Wang B-X, Li X, Deng C-L, Gao W-J (2017) Chromosome evolution in connection with repetitive sequences and epigenetics in plants. Genes 8:290–308
Lim K-B, De Jong H, Yang T-J, Park J-Y, Kwon S-J, Kim JS, Lim M-H, Kim JA, Jin M, Jin Y-M (2005) Characterization of rDNAs and tandem repeats in the heterochromatin of Brassica rapa. Mol Cells 19:436–444
Lippert L, Smith P, Bergh B (1966) Cytogenetics of the vegetable crops. Garden pepper, Capsicum sp. Bot Rev 32:24–55
Liu Z, Liu Y, Liu F, Zhang S, Wang X, Lu Q, Wang K, Zhang B, Peng R (2018) Genome-wide survey and comparative analysis of long terminal repeat (LTR) retrotransposon families in four Gossypium species. Sci Rep 8:9399
Mehrotra S, Goyal V (2014) Repetitive sequences in plant nuclear DNA: types, distribution, evolution and function. Genomics Proteomics Bioinform 12:164–171
Moscone EA, Scaldaferro MA, Grabiele M, Cecchini NM, Sánchez García Y, Jarret R, Daviña JR, Ducasse DA, Barboza GE, Ehrendorfer F (2007) The evolution of chili peppers (Capsicum-Solanaceae): a cytogenetic perspective. Acta Hortic 745:137–170
Muratović E, Robin O, Bogunić F, Šoljan D, Siljak-Yakovlev S (2010) Karyotype evolution and speciation of European lilies from Lilium sect. Liriotypus. Taxon 59:165–175
Novák P, Neumann P, Pech J, Steinhaisl J, Macas J (2013) RepeatExplorer: a Galaxy-based web server for genome-wide characterization of eukaryotic repetitive elements from next-generation sequence reads. Bioinformatics 29:792–793
Novák P, Ávila Robledillo L, Koblížková A, Vrbová I, Neumann P, Macas J (2017) TAREAN: a computational tool for identification and characterization of satellite DNA from unassembled short reads. Nucleic Acids Res 45:e111–e111
Paran I, Ben-Chaim A, Kang B-C, Jahn M (2007) Capsicums. In: Kole C (ed) Genome mapping and molecular breeding in plants, vol 5. Vegetables. Springer, Berlin, pp 209–226
Park M, Park J, Kim S, Kwon JK, Park HM, Bae IH, Yang TJ, Lee YH, Kang BC, Choi D (2012) Evolution of the large genome in Capsicum annuum occurred through accumulation of single-type long terminal repeat retrotransposons and their derivatives. Plant J 69:1018–1029
Pertea M (2012) The human transcriptome: an unfinished story. Genes 3:344–360
Pickersgill B (1991) Cytogenetics and evolution of Capsicum L. Chromosome Eng Plants Genet Breed Evol 33:139–160
Piegu B, Guyot R, Picault N, Roulin A, Saniyal A, Kim H, Collura K, Brar DS, Jackson S, Wing RA (2006) Doubling genome size without polyploidization: dynamics of retrotransposition-driven genomic expansions in Oryza australiensis, a wild relative of rice. Genome Res 16:1262–1269
Qin C, Yu C, Shen Y, Fang X, Chen L, Min J, Cheng J, Zhao S, Xu M, Luo Y (2014) Whole-genome sequencing of cultivated and wild peppers provides insights into Capsicum domestication and specialization. Proc Natl Acad Sci 111:5135–5140
Rao SR, Trivedi S, Emmanuel D, Merita K, Hynniewta M (2010) DNA repetitive sequences-types, distribution and function: a review. J Cell Mol Biol 7:1–11
Richard G-F, Kerrest A, Dujon B (2008) Comparative genomics and molecular dynamics of DNA repeats in eukaryotes. Microbiol Mol Biol Rev 72:686–727
Rohami M, Mohammadi A, Khosroshahli M, Ahmadi H, Darandeh N (2010) Karyotype analysis of several ecotypes of Capsicum annuum L. in Iran. Not Bot Horti Agrobot Cluj-Napoca 38:177–180
Romero-da Cruz MV, Urdampilleta JD, Forni Martins ER, Moscone EA (2017) Cytogenetic markers for the characterization of Capsicum annuum L. cultivars. Plant Biosyst 151:84–91
Schnable PS, Ware D, Fulton RS, Stein JC, Wei F, Pasternak S, Liang C, Zhang J, Fulton L, Graves TA (2009) The B73 maize genome: complexity, diversity, and dynamics. Science 326:1112–1115
Schrader L, Kim JW, Ence D, Zimin A, Klein A, Wyschetzki K, Weichselgartner T, Kemena C, Stökl J, Schultner E (2014) Transposable element islands facilitate adaptation to novel environments in an invasive species. Nat Commun 5:5495
Waminal NE, Kim NS, Kim HH (2011) Dual-color FISH karyotype analyses using rDNAs in three Cucurbitaceae species. Genes Genomics 33:521–528
Waminal NE, Pellerin RJ, Kim N-S, Jayakodi M, Park JY, Yang T-J, Kim HH (2018) Rapid and efficient FISH using pre-labeled oligomer probes. Sci Rep 8:8224
Wang D, Su Y, Wang X, Lei H, Yu J (2012) Transposon-derived and satellite-derived repetitive sequences play distinct functional roles in mammalian intron size expansion. Evol Bioinform Online 8:301–319
Witsenboer H, Michelmore R, Vogel J (1997) Identification, genetic localization, and allelic diversity of selectively amplified microsatellite polymorphic loci in lettuce and wild relatives (Lactuca spp.). Genome 40:923–936
Xu X, Pan S, Cheng S, Zhang B, Mu D, Ni P, Zhang G, Yang S, Li R, Wang J et al (2011) Genome sequence and analysis of the tuber crop potato. Nature 475:189–195
Acknowledgements
This paper was supported by the Sahmyook University Research Fund in 2017 (Grant no. RI12017006).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zhou, H.C., Waminal, N.E. & Kim, H.H. In silico mining and FISH mapping of a chromosome-specific satellite DNA in Capsicum annuum L.. Genes Genom 41, 1001–1006 (2019). https://doi.org/10.1007/s13258-019-00832-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-019-00832-8