Abstract
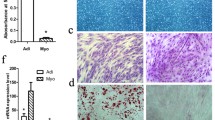
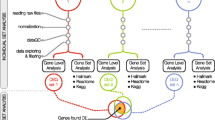
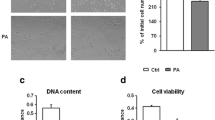
The molecular mechanisms underlying myogenic satellite cells (MSCs) differentiation into myotube-formed cells (MFCs) and transdifferentiation into adipocyte-like cells (ALCs) are unclear. As a step towards understanding the molecular mechanisms underlying MSC differentiation and transdifferentiation, we attempted to identify the genes differentially expressed during differentiation and transdifferentiation using gene microarray analysis (GMA). Thirty oligonucleotide arrays were used with two technical replicates and nine and six biological replicates for MFCs vs. MSCs and ALCs vs. MSCs, respectively, to contrast expression profile differences. GMA identified 1,224 differentially expressed genes by at least 2-fold during differentiation and transdifferentiation of MSCs. To select the highly expressed genes for future functional study, genes with a 4-fold expression difference were selected for validation by real time RT-PCR and approximately 96.9% of the genes were validated. The up-regulation of marker genes for myogenesis (MYL2, MYH3) and adipogenesis (PPARγ, and FABP4) was observed during the differentiation and transdifferentiation of MSCs into MFCs and ALCs, respectively. KOG analysis revealed that the most of the genes up-regulated during differentiation and transdifferentiation of MSCs were related to signal transduction. Again the exact location of 109 differentially expressed genes by 4-fold were analyzed by chromosome mapping. Among those, co-localization of 29 genes up-regulated during transdifferentiation with QTL for marbling score and intramuscular fat percentage supports the involvement of these genes in cellular transdifferentiation. Interestingly, some genes with unknown function were also identified during the process. Functional studies on these genes may unfold the molecular mechanisms controlling MSC differentiation and transdifferentiation.
Similar content being viewed by others
References
Asakura A, Komaki M, and Rudnicki M (2001) Muscle satellite cells are multipotential stem cells that exhibit myogenic, osteogenic, and adipogenic differentiation. Differentiation 68: 245–253.
Buckingham M, Bajard L, Chang T, Daubas P, Hadchouel J, Meilhac S, Montarras D, Hadchouel J, Meilhac S, and Montarras D (2003) The formation of skeletal muscle: from s&omite to limb. J. Anat. 202: 59–68.
Campbell SE, Tandon NN, Woldegiorgis G, Luiken JJ, Glatz JF, and Bonen A (2004) A novel function for fatty acid translocase (FAT)/CD36: involvement in long chain fatty acid transfer into the mitochondria. J. Biol. Chem. 279: 36235–36241.
Casas E, Shackelford SD, Keele JW, Koohmaraie M, Smith TPL, and Stone RT (2003) Detection of quantitative trait loci for growth and carcass composition in cattle. J. Anim. Sci. 81: 2976–2983.
Coburn CT, Knapp Jr. FF, Febbraio M, Beets AL, Silverstein RL, and Abumrad NA (2000) Defective uptake and utilization of long-chain fatty acids in muscle and adipose tissues of CD36 knockout mice. J. Biol. Chem. 275: 32523–32529.
Drover VA, Nguyen DV, Bastie CC, Darlington YF, Abumrad NA, Pessin JE, London E, Sahoo D, and Phillips MC. (2008) CD36 mediates both cellular uptake of very long chain fatty acids and their intestinal absorption in mice. J. Biol. Chem. 283: 13108–13115.
Fux C, Mitta B, Kramer BP, and Fussenegger M. (2004) Dual-regulated expression of C/EBP-alpha and BMP-2 enables differential differentiation of C2C12 cells into adipocytes and osteoblasts. Nucleic Acids Res. 2:32:e1–e9.
Garbe JR, Elsik CG, Antoniou E, Reecy JM, Clark KJ, Venkatraman V, Kim JW, Schnabel RD, Dickens MC, Wolfinger RD et al. (2010) Development and application of bovine and porcine oligonucleotide arrays with protein based annotation. J. Biomed. Biotechnol. Vol. 2010. Doi:10.1155/2010/453638.
Hawke TJ, and Garry DJ (2001) Myogenic satellite cells: physiology to molecular biology. J. Appl. Physiol. 91: 534–551.
Hu E, Tontonoz P, and Spiegelman BM (1995) Transdifferentiation of myoblasts by the adipogenic transcription factors PPAR gamma and C/EBPα. Proc. Natl. Acad. Sci. USA 92: 9856–9860.
Kook SH, Choi KC, Son YO, Lee KY, Hwang IH, Lee HJ, Chang JS, Choi IH, and Lee JC (2006) Satellite cells isolated from adult Hanwoo muscle can proliferate and differentiate into myoblasts and adipose-like cells. Mol. Cells 22: 239–245.
Lee DM, Bajracharya P, Lee EJ, Lee HJ, Kim JE, Lee HJ, Chun T, Kim J, Cho KH, Chang J et al. (2011) In Vitro Cell. Develop. Biol. — Animal 47: 438–444.
Mauro A (1961) Satellite cells of skeletal muscle fibres. J. Biophys. Biochem. Cytol. 9: 493–495.
McClure MC, Morsci NS, Schnabel RD, Kim JW, Yao P, Rolf MM, McKay SD, Gregg SJ, Chapple RH, Northcutt SL et al. (2010) A Genome scan for quantitative trait loci influencing carcass, post-natal growth, and reproductive traits in commercial Angus cattle. Anim. Genet. 41: 597–607.
Mody N, Graham TE, Tsuji Y, Yang Q, and Kahn BB (2008). Decreased clearance of serum retinol-binding protein and elevated levels of transthyretin in insulin-resistant ob/ob mice. Endocrinol. Metabol. 294: E785–E793.
Moran JL, Li Y, Hill AA, Mounts WM, and Miller CP (2002) Gene expression changes during mouse skeletal myoblast differentiation revealed by transcriptional profiling. Physiol. Genomics 10: 103–111.
Poulos SP, and Hausman GJA (2006) comparison of thiazolidined ione-induced adipogenesis and myogenesis in stromal-vascular cells from subcutaneous adipose tissue or semitendinosus muscle of postnatal pigs. J. Anim. Sci. 84: 1076–1082.
Rudnicki MA, Schnegelsberg PN, Stead RH, Braun T, Arnold HH, and Jaenisch R (1993) MyoD or Myf-5 is required for the formation of skeletal muscle. Cell 75: 1351–1359.
Shen X, Collier JM, Hlaing M, Zhang L, Delshad EH, Bristow J, and Bernstein HS (2003) Genome-wide examination of myoblast cell cycle withdrawal during differentiation. Dev. Dyn. 226: 128–138.
Singh NK, Chae HS, Hwang IH, Yoo YM, Ahn CN, Lee SH, Lee HJ, Park HJ, and Chung HY (2007) Transdifferentiation of porcine satellite cells to adipoblasts with ciglitizone. J. Anim. Sci. 85: 1126–1135.
Sterrenburg E, Turk R, ’t Hoen PAC, van Deutekom JCT, Boer JM, van Ommen GJ, and den Dunnen JT (2004) Large-scale gene expression analysis of human skeletal myoblast differentiation. Neuromuscul. Disord. 14: 507–518.
Tajbakhsh S, Rocancourt D, and Buckingham M (1996) Muscle progenitor cells failing to respond to positional cues adopts non-myogenic fates in myf-5 null mice. Nature 384: 266–270.
Tan SH, Reverter A, Wang YH, Byrne KA, McWilliam SM, and Lehnert SA (2006) Gene expression profiling of bovine in vitro adipogenesis using a cDNA microarray. Funct Integr Genomics. 6: 235–249.
Taylor JF, Coutinho LL, Herring KL, Gallagher DS, Brenneman Jr. RA, Burney N, Sanders JO, Turner JW, Smith SB, Miller RK, et al. (1998) Davis SK. Candidate gene analysis of GH1 for effects on growth and carcass composition of cattle. Anim. Genet. 29: 194–201.
Teboul L, Gaillard D, Staccini L, Inadera H, Amri EZ, and Grimaldi PA. (1995) Thiazolidinediones and fatty acids convert myogenic cells into adipose-like cells. J. Biol. Chem. 270: 28183–28187.
Tomczak KK, Marinescu VD, Tamoni MF, Sanoudou D, Montanaro F, Han M, Kunkel LM, Kohane IS, and Beggs AH (2003) Expression profiling and identification of novel genes involved in myogenic differentiation. The FASEB J. 18: 403–405.
Wheeler TL, Cundiff LV, and Koch RM (1994) Effect of marbling degree on beef palatability in Bos taurus and Bos indicus cattle. J. Anim. Sci. 72: 3145–3151.
Zhou J, Febbraio M, Zhai Y, Kuruba R, Wada T, Khadem S, Ren S, Li S, Silverstein RL, and Xie W (2008) LXR, PXR, and PPARγ cooperate in regulating fatty acid transporter CD36 and promoting hepatic lipogenesis. Gastroenterology 134: 556–567.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Lee, E.J., Bajracharya, P., Lee, DM. et al. Gene expression profiles during differentiation and transdifferentiation of bovine myogenic satellite cells. Genes Genom 34, 133–148 (2012). https://doi.org/10.1007/s13258-011-0096-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-011-0096-z