Abstract
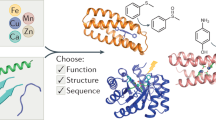
Functional proteins designed de novo have potential application in chemical engineering, agriculture and healthcare. Metal binding sites are commonly used to incorporate functions. Based on a de novo designed protein DS119 with a βαβ structure, we have computationally engineered zinc binding sites into it using a home-made searching program. Seven out of the eight designed sequences tested were shown to bind Zn2+ with micromolar affinity, and one of them bound Zn2+ with 1:1 stoichiometry. This is the first time that metalloproteins with an α, β mixed structure have been designed from scratch.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Ambroggio, X.I., and Kuhlman, B. (2006). Computational design of a single amino acid sequence that can switch between two distinct protein folds. J Am Chem Soc 128, 1154–1161.
Berg, J.M., and Shi, Y.G. (1996). The galvanization of biology: a growing appreciation for the roles of zinc. Science 271, 1081–1085.
Berman, H.M., Westbrook, J., Feng, Z., Gilliland, G., Bhat, T.N., Weissig, H., Shindyalov, I.N., and Bourne, P.E. (2000). The Protein Data Bank. Nucleic Acids Res 28, 235–242.
Cerasoli, E., Sharpe, B.K., and Woolfson, D.N. (2005). ZiCo: a peptide designed to switch folded state upon binding zinc. J Am Chem Soc 127, 15008–15009.
Choma, C.T., Lear, J.D., Nelson, M.J., Dutton, P.L., Robertson, D.E., and Degrado, W.F. (1994). DESIGN OF A HEME-BINDING 4-HELIX BUNDLE. J Am Chem Soc 116, 856–865.
Christianson, D.W., and Fierke, C.A. (1996). Carbonic anhydrase: Evolution of the zinc binding site by nature and by design. Acc Chem Res 29, 331–339.
Clarke, N.D., and Yuan, S.M. (1995). Metal search: a computer program that helps design tetrahedral metal-binding sites. Proteins 23, 256–263.
De Maeyer, M., Desmet, J., and Lasters, I. (1997). All in one: a highly detailed rotamer library improves both accuracy and speed in the modelling of sidechains by dead-end elimination. Fold Des 2, 53–66.
Dwyer, M.A., Looger, L.L., and Hellinga, H.W. (2003). Computational design of a Zn2 + receptor that controls bacterial gene expression. Proc Natl Acad Sci U S A 100, 11255–11260.
Dyer, R.B., Gai, F., and Woodruff, W.H. (1998). Infrared studies of fast events in protein folding. Acc Chem Res 31, 709–716.
Fry, H.C., Lehmann, A., Saven, J.G., DeGrado, W.F., and Therien, M. J. (2010). Computational design and elaboration of a de novo heterotetrameric alpha-helical protein that selectively binds an emissive abiological (porphinato)zinc chromophore. J Am Chem Soc 132, 3997–4005.
Handel, T.M., Williams, S.A., and DeGrado, W.F. (1993). Metal ion-dependent modulation of the dynamics of a designed protein. Science 261, 879–885.
Hellinga, H.W., and Richards, F.M. (1991). Construction of new ligand binding sites in proteins of known structure. I. Computer-aided modeling of sites with pre-defined geometry. J Mol Biol 222, 763–785.
Holm, R.H., Kennepohl, P., and Solomon, E.I. (1996). Structural and functional aspects of metal sites in biology. Chem Rev 96, 2239–2314.
Kaplan, J., and DeGrado, W.F. (2004). De novo design of catalytic proteins. Proc Natl Acad Sci U S A 101, 11566–11570.
Kiyokawa, T., Kanaori, K., Tajima, K., Koike, M., Mizuno, T., Oku, J.I., and Tanaka, T. (2004). Binding of Cu(II) or Zn(II) in a de novo designed triple-stranded alpha-helical coiled-coil toward a prototype for a metalloenzyme. J Pept Res 63, 347–353.
Li, W.F., Zhang, J., and Wang, W. (2007). Understanding the folding and stability of a zinc finger-based full sequence design protein with replica exchange molecular dynamics simulations. Proteins 67, 338–349.
Li, W.F., Zhang, J., Wang, J., and Wang, W. (2008). Metal-coupled folding of Cys2His2 zinc-finger. J Am Chem Soc 130, 892–900.
Liang, H.H., Chen, H., Fan, K.Q., Wei, P., Guo, X.R., Jin, C.W., Zeng, C., Tang, C., and Lai, L.H. (2009). De novo design of a beta alpha beta motif. Angew Chem Int Ed Engl 48, 3301–3303.
Lombardi, A., Summa, C.M., Geremia, S., Randaccio, L., Pavone, V., and DeGrado, W.F. (2000). Retrostructural analysis of metalloproteins: application to the design of a minimal model for diiron proteins. Proc Natl Acad Sci U S A 97, 6298–6305.
Lu, Y., Berry, S.M., and Pfister, T.D. (2001). Engineering novel metalloproteins: design of metal-binding sites into native protein scaffolds. Chem Rev 101, 3047–3080.
Lu, Y., Yeung, N., Sieracki, N., and Marshall, N.M. (2009). Design of functional metalloproteins. Nature 460, 855–862.
Matzapetakis, M., Farrer, B.T., Weng, T.C., Hemmingsen, L., Penner-Hahn, J.E., and Pecoraro, V.L. (2002). Comparison of the binding of cadmium(II), mercury(II), and arsenic(III) to the de novo designed peptides TRI L12C and TRI L16C. J Am Chem Soc 124, 8042–8054.
Meinnel, T., Blanquet, S., and Dardel, F. (1996). A new subclass of the zinc metalloproteases superfamily revealed by the solution structure of peptide deformylase. J Mol Biol 262, 375–386.
Miller, J.C., Holmes, M.C., Wang, J.B., Guschin, D.Y., Lee, Y.L., Rupniewski, I., Beausejour, C.M., Waite, A.J., Wang, N.S., Kim, K. A., et al. (2007). An improved zinc-finger nuclease architecture for highly specific genome editing. Nat Biotechnol 25, 778–785.
Müller, H.N., and Skerra, A. (1994). GGrafting of a high-affinity Zn(II)-binding site on the beta-barrel of retinol-binding protein results in enhanced folding stability and enables simplified purification. Biochemistry 33, 14126–14135.
Nomura, A., and Sugiura, Y. (2004). Hydrolytic reaction by zinc finger mutant peptides: successful redesign of structural zinc sites into catalytic zinc sites. Inorg Chem 43, 1708–1713.
Pasquinelli, R.S., Shepherd, R.E., Koepsel, R.R., Zhao, A., and Ataai, M.M. (2000). Design of affinity tags for one-step protein purification from immobilized zinc columns. Biotechnol Prog 16, 86–91.
Petros, A.K., Reddi, A.R., Kennedy, M.L., Hyslop, A.G., and Gibney, B.R. (2006). Femtomolar Zn(II) affinity in a peptide-based ligand designed to model thiolate-rich metalloprotein active sites. Inorg Chem 45, 9941–9958.
Phillips, N.B., Wan, Z.L., Whittaker, L., Hu, S.Q., Huang, K., Hua, Q. X., Whittaker, J., Ismail-Beigi, F., and Weiss, M.A. (2010). Supramolecular protein engineering: design of zinc-stapled insulin hexamers as a long acting depot. J Biol Chem 285, 11755–11759.
Proudfoot, C., McPherson, A.L., Kolb, A.F., and Stark, W.M. (2011). Zinc finger recombinases with adaptable DNA sequence specificity. PLoS One 6, e19537.
Reddi, A.R., Guzman, T.R., Breece, R.M., Tiemey, D.L. and Gibney, B.R. (2007). Deducing the Energetic Cost of Protein Folding in Zinc Finger Proteins Using Designed Metallopeptides. J Am Chem Soc, 129, 12815–12827.
Regan, L., and Clarke, N.D. (1990). A tetrahedral zinc(II)-binding site introduced into a designed protein. Biochemistry 29, 10878–10883.
Shults, M.D., Pearce, D.A., and Imperiali, B. (2003). Modular and tunable chemosensor scaffold for divalent zinc. J Am Chem Soc 125, 10591–10597.
Smith, B.A., and Hecht, M.H. (2011). Novel proteins: from fold to function. Curr Opin Chem Biol 15, 421–426.
Stevens, F.J. (1986). Analysis of protein-protein interaction by simulation of small-zone size-exclusion chromatography: application to an antibody-antigen association. Biochemistry 25, 981–993.
Wade, W.S., Koh, J.S., Han, N., Hoekstra, D.M., and Lerner, R.A. (1993). Engineering Metal Coordination Sites into the Antibody Light-Chain. J Am Chem Soc 115, 4449–4456.
Winzor, D.J. (2003). Analytical exclusion chromatography. J Biochem Biophys Methods 56, 15–52.
Wu, J., Kandavelou, K., and Chandrasegaran, S. (2007). Custom-designed zinc finger nucleases: what is next? Cell Mol Life Sci 64, 2933–2944.
Author information
Authors and Affiliations
Corresponding author
Additional information
These authors contributed equally to the work.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Zhu, C., Zhang, C., Liang, H. et al. Engineering a zinc binding site into the de novo designed protein DS119 with a βαβ structure. Protein Cell 2, 1006–1013 (2011). https://doi.org/10.1007/s13238-011-1121-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13238-011-1121-3