Abstract
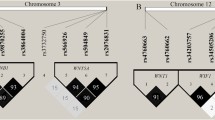
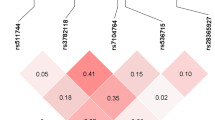
Multiple host genes determine susceptibility or resistance to tuberculosis. In an exome-wide association study conducted among tuberculosis patients and their exposed but clinically asymptomatic household contacts, we found that the SNP rs61104666 located in the fifth exon of SIGLEC15 gene is associated with the disease. No other variant in SIGLEC15 has been reported previously to be associated with tuberculosis. The associated polymorphism results in a synonymous change (E292E) and therefore is unlikely to be involved in disease pathogenesis. Bioinformatic analysis of epigenetic marks in the genomic region reveals an enhancer mark present in lung and blood, downstream to the SIGLEC15 gene may harbor candidate causal SNPs which are in strong LD with the index SNP. This region overlaps with the 3′UTR region of the neighboring gene EPG5. EPG5 has role in autophagy, a phenomenon relevant to clearing of the infection. The region also harbors DNAse I sensitive sites with SNPs of low RegulomeDB score indicative of potential transcription binding sites. All these evidences suggest further exploration of the enhancer region to understand its role in disease manifestation.



Similar content being viewed by others
Change history
19 July 2019
Due to an oversight an error with respect to SNP number and associated polymorphism was crept in the abstract of the original version. The corrected version of the same should read as follows.
References
Angata T, Ishii T, Motegi T, Oka R, Taylor RE, Soto PC, Chang YC, Secundino I, Gao CX, Ohtsubo K, et al. Loss of Siglec-14 reduces the risk of chronic obstructive pulmonary disease exacerbation. Cell Mol Life Sci. 2013;70:3199–210.
Bhattacharyya C, Majumder PP, Pandit B. An exome wide association study of pulmonary tuberculosis patients and their asymptomatic household contacts. Infect Genet Evol. 2019;71:76–81.
Bornhofft KF, Goldammer T, Rebl A, Galuska SP. Siglecs: a journey through the evolution of sialic acid-binding immunoglobulin-type lectins. Dev Comp Immunol. 2018;86:219–31.
Bukvic BK, Blekic M, Simpson A, Marinho S, Curtin JA, Hankinson J, Aberle N, Custovic A. Asthma severity, polymorphisms in 20p13 and their interaction with tobacco smoke exposure. Pediatr Allergy Immunol. 2013;24:10–8.
Chang YC, Olson J, Beasley FC, Tung C, Zhang J, Crocker PR, Varki A, Nizet V. Group B Streptococcus engages an inhibitory Siglec through sialic acid mimicry to blunt innate immune and inflammatory responses in vivo. PLoS Path. 2014;10:e1003846.
Chen GY, Chen X, King S, Cavassani KA, Cheng J, Zheng X, Cao H, Yu H, Qu J, Fang D, et al. Amelioration of sepsis by inhibiting sialidase-mediated disruption of the CD24-SiglecG interaction. Nat Biotechnol. 2011;29:428–35.
Delgado JC, Baena A, Thim S, Goldfeld AE. Ethnic-specific genetic associations with pulmonary tuberculosis. J Infect Dis. 2002;186:1463–8.
Ernst JD. The immunological life cycle of tuberculosis. Nat Rev Immunol. 2012;12:581–91.
Fol M, Druszczynska M, Wlodarczyk M, Ograczyk E, Rudnicka W. Immune response gene polymorphisms in tuberculosis. Acta Biochim Pol. 2015;62:633–40.
Gao PS, Shimizu K, Grant AV, Rafaels N, Zhou LF, Hudson SA, Konno S, Zimmermann N, Araujo MI, Ponte EV, et al. Polymorphisms in the sialic acid-binding immunoglobulin-like lectin-8 (Siglec-8) gene are associated with susceptibility to asthma. Eur J Hum Genet. 2010;18:713–9.
Global Tuberculosis Report 2017. World Health Organization 2017.
Graustein AD, Horne DJ, Fong JJ, Schwarz F, Mefford HC, Peterson GJ, Wells RD, Musvosvi M, Shey M, Hanekom WA, et al. The SIGLEC14 null allele is associated with Mycobacterium tuberculosis- and BCG-induced clinical and immunologic outcomes. Tuberculosis. 2017;104:38–45.
Hammonds JE, Beeman N, Ding L, Takushi S, Francis AC, Wang JJ, Melikyan GB, Spearman P. Siglec-1 initiates formation of the virus-containing compartment and enhances macrophage-to-T cell transmission of HIV-1. PLoS Pathog. 2017;13:e1006181.
Hiruma Y, Tsuda E, Maeda N, Okada A, Kabasawa N, Miyamoto M, Hattori H, Fukuda C. Impaired osteoclast differentiation and function and mild osteopetrosis development in Siglec-15-deficient mice. Bone. 2013;53:87–93.
Kameda Y, Takahata M, Komatsu M, Mikuni S, Hatakeyama S, Shimizu T, Angata T, Kinjo M, Minami A, Iwasaki N. Siglec-15 regulates osteoclast differentiation by modulating RANKL-induced phosphatidylinositol 3-kinase/Akt and Erk pathways in association with signaling Adaptor DAP12. J Bone Miner Res. 2013;28:2463–75.
Kawasaki N, Rillahan CD, Cheng TY, Van Rhijn I, Macauley MS, Moody DB, Paulson JC. Targeted delivery of mycobacterial antigens to human dendritic cells via Siglec-7 induces robust T cell activation. J Immunol. 2014;193:1560–6.
Lu Q, Yokoyama CC, Williams JW, Baldridge MT, Jin X, DesRochers B, Bricker T, Wilen CB, Bagaitkar J, Loginicheva E, et al. Homeostatic control of innate lung inflammation by Vici syndrome gene Epg5 and additional autophagy genes promotes influenza pathogenesis. Cell Host Microbe. 2016;19:102–13.
Meilang Q, Zhang Y, Zhang J, Zhao Y, Tian C, Huang J, Fan H. Polymorphisms in the SLC11A1 gene and tuberculosis risk: a meta-analysis update. Int J Tuberc Lung Dis. 2012;16:437–46.
Meyer CG, Thye T. Host genetic studies in adult pulmonary tuberculosis. Semin Immunol. 2014;26:445–53.
Mroz RM, Holownia A, Wielgat P, Sitko A, Skopinski T, Chyczewska E, Braszko JJ. Siglec-8 in induced sputum of COPD patients. Adv Exp Med Biol. 2013;788:19–23.
Mukhopadhyay S, Thatoi PK, Pandey AD, Das BK, Ravindran B, Bhattacharjee S, Mohapatra SK. Transcriptomic meta-analysis reveals up-regulation of gene expression functional in osteoclast differentiation in human septic shock. PLoS ONE. 2017;12:e0171689.
Naranbhai V. The role of host genetics (and genomics) in tuberculosis. Microbiol Spectr. 2016;4:1–36.
Singh V, Jamwal S, Jain R, Verma P, Gokhale R, Rao KV. Mycobacterium tuberculosis-driven targeted recalibration of macrophage lipid homeostasis promotes the foamy phenotype. Cell Host Microbe. 2012;12:669–81.
Souza de Lima D, Nunes VCL, Ogusku MM, Sadahiro A, Pontillo A, Alencar BC. Polymorphisms in SIGLEC1 contribute to susceptibility to pulmonary active tuberculosis possibly through the modulation of IL-1ss. Infect Genet Evol. 2017;55:313–7.
Stein CM, Sausville L, Wejse C, Sobota RS, Zetola NM, Hill PC, Boom WH, Scott WK, Sirugo G, Williams SM. Genomics of human pulmonary tuberculosis: from genes to pathways. Curr Genet Med Rep. 2017;5:149–66.
Thye T, Vannberg FO, Wong SH, Owusu-Dabo E, Osei I, Gyapong J, Sirugo G, Sisay-Joof F, Enimil A, Chinbuah MA, et al. Genome-wide association analyses identifies a susceptibility locus for tuberculosis on chromosome 18q11.2. Nat Genet. 2010;42:739–41.
van Tong H, Velavan TP, Thye T, Meyer CG. Human genetic factors in tuberculosis: an update. Trop Med Int Health. 2017;22:1063–71.
Acknowledgements
We are grateful to all the study participants. Mr Bijan Bhusan Bairagya is thanked for excellent technical assistance. We are thankful to Dr. Arindam Maitra and CoTeRi for conducting the genotyping and sequencing experiments.
Funding
This work was supported by the grant BT/01/CEIB/11/VI/05 dated 23/11/2011, from Department of Biotechnology (DBT), Government of India, India. Partha P. Majumder was supported by the J.C.Bose Fellowship of the Government of India, Department of Science and Technology. Chandrika Bhattacharyya was supported by Junior and Senior Research Fellowship from University Grant Commission, India.
Author information
Authors and Affiliations
Contributions
BP and PPM planned the study and designed experiments. CB performed the experiments. Data was analysed by CB, PPM. Bioinformatic data mining and analysis done by BP. PPM provided critical insights. BP wrote the manuscript. All the authors read the manuscript, provided suggestions and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is based on the presentation made during 18th All India Congress of Cytology and Genetics and International Symposium on “Translating Genes and Genomes” held at CSIR-Indian Institute of Chemical Biology, Kolkata in collaboration with Archana Sharma Foundation of Calcutta during January 29–31, 2018.
Rights and permissions
About this article
Cite this article
Pandit, B., Bhattacharyya, C. & Majumder, P.P. SIGLECs and their contribution to tuberculosis. Nucleus 62, 119–125 (2019). https://doi.org/10.1007/s13237-019-00279-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13237-019-00279-y