Abstract
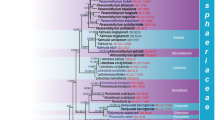
The family Pleosporaceae includes numerous saprobic, opportunistic human, and plant pathogenic taxa. The classification of genera and species Pleosporaceae has been a major challenge due to the lack of a clear understanding of the importance of the morphological characters used to distinguish taxa as well as the lack of reference strains. Recent treatments concluded that Pleospora and some other genera in Pleosporaceae are likely polyphyletic. In order to establish the evolutionary relationships and to resolve the polyphyletic nature of Pleospora and allied genera, we sequenced the 18S nrDNA, 28S nrDNA, ITS, GAPDH, RPB2 and TEF1-alpha gene regions of Pleosporaceae species and phylogenetically analysed this data. Multigene phylogenies strongly support the monophyletic nature of Pleosporaceae among the other families in Pleosporales, and the acceptance of the genera Alternaria, Bipolaris, Clathrospora, Comoclathris, Curvularia, Dactuliophora, Decorospora, Diademosa, Exserohilum, Extrawettsteinina, Gibbago, Neocamarosporium, Paradendryphiella, Platysporoides, Pleospora, Porocercospora, Pseudoyuconia and Pyrenophora. Austropleospora, Dendryphion, Edenia and Macrospora are excluded from the family based on morphology coupled with molecular data. Two novel species, Alternaria murispora in this paper and Comoclathris sedi are introduced. The sexual morph of Alternaria alternata is re-described and illustrated using modern concepts from fresh collections. The paraphyletic nature of Pleospora is resolved based on the available morpho-molecular data, but further sampling with fresh collections, reference or ex-type strains and molecular data are needed to obtain a natural classification of genera and the family.




























Similar content being viewed by others
References
Abler R (2003) Global change and local places: estimating, understanding, and reducing greenhouse gases
Agrios GN (2005) Plant pathology, 5th edn. California Academic Press, Elsevier San Diego
Alves A, Correia A, Phillips AJL (2006) Multi-gene genealogies and morphological data support Diplodia cupressi sp. nov., previously recognized as D. pinea f. sp. cupressi, as a distinct species. Fungal Divers 23:1–15
Amaradasa BS, Madrid H, Groenewald JZ, Crous PW, Amundsen K (2014) Porocercospora seminalis gen. et comb. nov., the causal organism of buffalograss false smut. Mycologia 106(1):77–85
Ariyawansa HA, Jones EBG, Suetrong S, Alias SA, Kang JC, Hyde KD (2013a) Halojulellaceae a new family of the order Pleosporales. Phytotaxa 130:14–24
Ariyawansa HA, Kang JC, Alias SA, Chukeatirote E, Hyde KD (2013b) Towards a natural classification of Dothideomycetes 1: the genera Dermatodothella, Dothideopsella, Grandigallia, Hysteropeltella and Gloeodiscus (Dothideomycetes incertae sedis). Phytotaxa 147(2):35–47
Ariyawansa HA, Maharachchikumbura SSN, Karunarathne SC, Chukeatirote E, Bahkali AH, Kang JC, Bhat JD, Hyde KD (2013c) Deniquelata barringtoniae from Barringtonia asiatica, associated with leaf spots of Barringtonia asiatica. Phytotaxa 105:11–20
Ariyawansa H, Phookamsak R, Tibpromma S, Kang JC, Hyde KD (2014a) A molecular and morphological reassessment of Diademaceae. Sci World J 2014:1–11
Ariyawansa HA, Camporesi E, Thambugala KM, Mapook A, Kang JC, Alias SA, Chukeatirote E, Thines M, Mckenzie EHC, Hyde KD (2014b) Confusion surrounding Didymosphaeria — phylogenetic and morphological evidence suggest Didymosphaeriaceae is not a distinct family. Phytotaxa 176(1):102–119
Ariyawansa HA, Hawksworth DL, Hyde KD, Jones EBG, Maharachchikumbura SSN, Manamgoda DS, Thambugala KM, Udayanga D, Camporesi E, Daranagama A, Jayawardena R, Liu JK, McKenzie EHC, Phookamsak R, Senanayake IC, Shivas RG, Tian Q, Xu JC (2014c) Epitypification and neotypification: guidelines with appropriate and inappropriate examples. Fungal Divers 69(1):57–91
Ariyawansa HA, Kang JC, Alias SA, Chukeatirote E, Hyde KD (2014d) Pyrenophora. Mycosphere 5(2):351–362
Ariyawansa HA, Tanaka K, Thambugala KM, Phookamsak R, Tian Q, Camporesi E, Hongsanan S, Monkai J, Wanasinghe DN, Chukeatirote E, Kang JC, Xu JC, McKenzie EHC, Jones EBG, Hyde KD (2014e) A molecular phylogenetic reappraisal of the Didymosphaeriaceae (= Montagnulaceae). Fungal Divers 68:69–104
Ariyawansa HA, Thambugala KM, Kang JC, Alias SA, Chukeatirote E, Hyde KD (2014f) Towards a natural classification of Dothideomycetes 2: The genera Cucurbidothis, Heterosphaeriopsis, Hyalosphaera, Navicella and Pleiostomellina (Dothideomycetes incertae sedis). Phytotaxa 176(1):7–17
Barr ME (1972) Preliminary studies on the Dothideales in temperate North America. Contrib Univ Mich Herb 9:523–638
Barr ME (1975) A note on Extrawettsteinina. Mycotaxon 2:104–106
Barr ME (1981) The genus Curreya: an example of taxonomic confusion in the Ascomycetes. Mycologia 73:599–609
Barr ME (1987a) New taxa and combinations in the Loculoascomycetes. Mycotaxon 29:501–505
Barr ME (1987b) Prodromus to class loculoascomycetes. Amherst. University of Massachusetts, Massachusetts
Barr ME (1990) Some dictyosporous genera and species of Pleosporales in North America. Mem N Y Bot Gard 62:1–92
Batista AC, Costa CA, Peres GEP, Leal FB (1959) Novos e antigos fungos Microthyriaceae. Anais Soc Biol Pernambuco 16:129–140
Berbee ML (1996) Loculoascomycete origins and evolution of filamentous ascomycete morphology based on 18S rRNA gene sequence data. Mol Biol Evol 13:462–470
Berbee ML, Pirseyedi M, Hubbard S (1999) Cochliobolus phylogenetics and the origin of known, highly virulent pathogens, inferred from ITS and glyceraldehyde-3-phosphate dehydrogenase gene sequences. Mycologia 91:964–977
Boedijn KB (1933) Uebr Einige Phragmosporen Dematiazeen. Bull Jard Bot Buitenz 13(3 Sér):123
Boonmee S, Rossman AY, Lui JK, LiWJ DDQ, Bhat JD, Jones EBG, McKenzie EHC, Xu JC, Hyde KD (2014) Tubeufiales, ord. nov. integrating sexual and asexual generic names. Fungal Divers 68(1):239–298
Cai L, Wu WP, Hyde KD (2009) Phylogenetic relationships of Chalara and allied species inferred from ribosomal DNA sequences. Mycol Prog 8(2):133–143
Carbone I, Kohn LM (1999) A method for designing primer sets for speciation studies in filamentous ascomycetes. Mycologia 91:553–556
Carter E, Boudreaux C (2004) Fatal cerebral phaeohyphomycosis due to Curvularia lunata in an immunocompetent patient. J Clin Microbiol 42(11):5419–5423
Chomnunti P, Hongsanan S, Aguirre-Hudson B, Tian Q, Peršoh D, Dhami MK, Alis AS, Xu J, Liu X, Stadler M, Hyde KD (2014) The sooty moulds. Fungal Divers 66:1–36
Clements FE (1909) The genera of fungi. The HW Wilson Company
Crous PW, Slippers B, Wingfield MJ, Rheeder J, Marasas WFO et al (2006) Phylogenetic lineages in the Botryosphaeriaceae. Stud Mycol 55:235–253
Crous PW, Shivas RG, Quaedvlieg W, van der Bank M, Zhang Y, Summerell BA, Tan YP (2014) Fungal Planet description sheets: 214–280. Persoonia 32:184
de Gruyter J, Woudenberg JHC, Aveskamp MM, Verkley GJM, Groenewald JZ, Crous PW (2013) Redisposition of phoma-like anamorphs in pleosporales. Stud Mycol 75:1–36
Desmazières JBHJ (1851) Dix-neuvième notice sur les plantes cryptogames récemment découvertes en France. Ann Sci Nat Botanique 16:296–330
Dettman JR, Jacobson DJ, Turner E, Pringle A, Taylor JW (2003) Reproductive isolation and phylogenetic divergence in Neurospora: comparing methods of species recognition in a model eukaryote. Evolution 57:2721–2741
Dong JW, Chen WD, Crane JL (1998) Phylogenetic studies of the Leptosphaeriaceae, Pleosporaceae and some other Loculoascomycetes based on nuclear ribosomal DNA sequences. Mycol Res 102:151–156
Drechsler C (1934) Phytopathological and taxonomical aspects of Ophilobolus, Pyrenophora, Helminthosporium and a new genus Cochliobolus. Phytopathology 24(9):953–981
Ellis MB (1971) Dematiaceous hyphomycetes. CMI, Kew
Eriksson OE (1981) The families of bitunicate ascomycetes. Opera Bot 60:1–220
Eriksson OE (2006) Outline of ascomycota – 2006. Myconet 12:1–82
Eriksson OE, Hawksworth DL (1986) An alphabetical list of the generic names of ascomycetes – 1986. Syst Ascomyc 5:3–111
Eriksson OE, Hawksworth DL (1991) Outline of the ascomycetes – 1990. Syst Ascomyc Reprint of Volumes 1–4 (1982–1985) 9: 39–271
Eriksson OE, Hawksworth DL (1998) Outline of the ascomycetes −1998. Syst Ascomyc 16:83–296
Farr DF, Bills GF, Chamuris GP, Rossman AY (1989) Fungi on plants and plant products in the United States. APS Press, St. Paul
Goh KT, Hyde KD, Lee KLD (1998) Generic distinction in Helminthosporium complex based on restriction analysis of the nuclear ribosomal RNA gene. Fungal Divers 1:85–107
Hibbett DS, Binder M, Bischoff JF, Blackwell M, Cannon PF, Eriksson OE, Huhndorf S, James T, Kirk PM, Lucking R, Lumbsch HT, Lutzoni F, Matheny PB, Mclaughlin DJ, Powell MJ, Redhead S, Schoch CL, Spatafora JW, Stalpers JA, Vilgalys R, Aime MC, Aptroot A, Bauer R, Begerow D, Benny GL, Castlebury LA, Crous PW, Dai YC, Gams W, Geiser DM, Griffith GW, Gueidan C, Hawksworth DL, Hestmark G, Hosaka K, Humber RA, Hyde KD, Ironside JE, Koljalg U, Kurtzman CP, Larsson KH, Lichtwardt R, Longcore J, Miadlikowska J, Miller A, Moncalvo JM, Mozley- Standridge S, Oberwinkler F, Parmasto E, Reeb V, Rogers JD, Roux C, Ryvarden L, Sampaio JP, Schussler A, Sugiyama J, Thorn RG, Tibell L, Untereiner WA, Walker C, Wang Z, Weir A, Weiss M, White MM, Winka K, Yao YJ, Zhang N (2007) A higher-level phylogenetic classification of the fungi. Mycol Res 111:509–547
Huelsenbeck JP, Ronquist F (2001) MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 17:754–755
Hyde KD, Jones EBG, Liu JK, Ariyawansa H, Boehm E, Boonmee S, Braun U, Chomnunti P, Crous PW, Dai DQ, Diederich P, Dissanayake A, Doilom M, Doveri F, Hongsanan S, Jayawardena R, Lawrey JD, Li YM, Liu YX, Lücking R, Monkai J, Muggia L, Nelsen MP, Pang KL, Phookamsak R, Senanayake IC, Shearer CA, Suetrong S, Tanaka K, Thambugala KM, Wijayawardene NN, Wikee S, Wu HX, Zhang Y, Aguirre-Hudson B, Alias SA, Aptroot A, Bahkali AH, Bezerra JL, Bhat DJ, Camporesi E, Chukeatirote E, Gueidan C, Hawksworth DL, Hirayama K, Hoog SD, Kang JC, Knudsen K, Li WJ, Li XH, Liu ZY, Mapook A, McKenzie EHC, Miller AN, Mortimer PE, Phillips AJL, Raja HA, Scheuer C, Schumm F, Taylor JE, Tian Q, Tibpromma S, Wanasinghe DN, Wang Y, Xu JC, Yacharoen S, Yan JY, Zhang M (2013) Families of dothideomycetes. Fungal Divers 63:1–313
Hyde KD, Nilsson RH, Alias SA, Ariyawansa HA, Blair JE, Cai L, de Cock AWAM, Dissanayake AJ, Glockling SL, Goonasekara ID, Gorczak M, Hahn M, Jayawardena RS, van Kan JAL, Laurence MH, Lévesque CA, Li X, Liu JK, Maharachchikumbura SSN, Manamgoda DS, Martin FN, McKenzie EHC, McTaggart AR, Mortimer PE, Nair PVR, Pawłowska J, Rintoul TL, Shivas RG, Spies CFJ, Summerell BA, Taylor PWJ, Terhem RB, Udayanga D, Vaghefi N, Walther G, Wilk M, Wrzosek M, Xu JC, Yan JY, Zhou N (2014) One stop shop: backbones trees for important phytopathogenic 5 genera: I (2014). Fungal Divers 67(1):21–125
Inderbitzin P, Kohlmeyer J, Volkmann-Kohlmeyer B, Berbee ML (2002) Decorospora, a new genus for the marine ascomycete Pleospora gaudefroyi. Mycologia 94(4):651–659
Inderbitzin P, Shoemaker RA, O’Neill NR, Turgeon BG, Berbee ML (2006) Systematics and mating systems of two fungal pathogens of opium poppy: the heterothallic Crivellia papaveracea and a homothallic species with a Brachycladium papaveris asexual state. Can J Bot 84:1304–1326
Index Fungorum (2015) http://www.indexfungorum.org
Jones EB, Stanley SJ, Pinruan U (2008) Marine endophyte sources of new chemical natural products: a review. Bot Mar 51(3):163–170
Khan JA, Hussain ST, Hasan S, McEvoy P, Sarwari A (2000) Disseminated bipolaris infection in an immunocompetent host: an atypical presentation. J Pak Med Assoc 50:68–71
Kirk PM, Cannon PF, David JC, Stalpers JA (2001) Dictionary of the fungi 9th edn. CABI, Wallingford
Kodsueb R, Dhanasekaran V, Aptroot A, Lumyong S, McKenzie EHC, Hyde KD, Jeewon R (2006) The family Pleosporaceae: intergeneric relationships and phylogenetic perspectives based on sequence analyses of partial 28S rDNA. Mycologia 98:571–583
Lawrence DP, Park MS, Pryor BM (2012) Nimbya and Embellisia revisited, with nov. comb for Alternaria celosiae and A. perpunctulata. Mycol Prog 11:799–815
Lawrence DP, Gannibal PB, Peever TL, Pryor BM (2013) The sections of Alternaria: formalizing species-groups concepts. Mycologia 105:530–546
Leakey CLA (1964) Dactuliophora, a new genus of mycelia sterilia from tropical Africa. Trans Br Mycol Soc 47(3):341–IN10
Leonard KJ, Suggs EG (1974) Setosphaeria prolata, the ascigerous state of Exserohilum prolatum. Mycologia 281–297
Leuchtmann A (1984) Über Phaeosphaeria Miyake und andere bitunicate Ascomyceten mit mehrfach querseptierten Ascosporen. Sydowia 37:75–194
Liu YJ, Whelen S, Hall BD (1999) Phylogenetic relationships among ascomycetes: evidence from an RNA polymerse II subunit. Mol Biol Evol 16:1799–1808
Liu JK, Phookamsak R, Mingkhuan M, Wikee S, Li YM et al (2012) Towards a natural classification of Botryosphaeriales. Fungal Divers 57:149–210
Lumbsch HT, Huhndorf SM (eds.) (2007) Outline of ascomycota −2007. Myconet 13:1–58
Lumbsch HT, Huhndorf SM (2010) Outline of ascomycota – 2009. Fieldiana Life Earth Sci 1:1–60
Luttrell ES (1951) Taxonomy of the pyrenomycetes. Univ Mo Stud 24:1–120
Luttrell ES (1955) The ascostromatci ascomycetes. Mycologia 47:511–532
Luttrell ES (1973) Loculoascomycetes. In: Ainsworth GC, Sparrow FK, Sussman AS (eds) The fungi, an advanced treatise, a taxonomic review with keys: ascomycetes and fungi imperfecti. Academic, New York
Malathrakis NE (1979) A study of an olive tree disease caused by the fungus Phoma incompta Sacc. et Mart. Dissertation, Agricultural College of Athens
Manamgoda DS, Cai L, Bahkali AH, Chukeatirote E, Hyde KD (2011) Cochliobolus: an overview and current status of species. Fungal Divers 51:3–42
Manamgoda DS, Cai L, McKenzie EHC, Crous PW, Madrid H, Chukeatirote E et al (2012) A phylogenetic and taxonomic re-evaluation of the Bipolaris-Cochliobolus-Curvularia complex. Fungal Divers 56(1):131–144
Manamgoda DS, Rossman AY, Castlebury LA, Crous PW, Madrid H, Chukeatirote E, Hyde KD (2014) The genus bipolaris. Stud in mycol. doi:10.1016/j.simyco.2014.10.002
Morin L, Shivas RG, Piper MC, Tan YP (2010) Austropleospora osteospermi gen. et sp. nov. and its host specificity and distribution on Chrysanthemoides monilifera ssp. rotundata in Australia. Fungal Divers 40(1):65–74
Nilsson RH, Hyde KD, Pawłowska J, Ryberg M et al (2014) Improving ITS sequence data for identification of plant pathogenic fungi. Fungal Divers 67(1):11–19
Nitschke TRJ (1869) Grundlage eines Systems der Pyrenomyceten Verh Naturhist Vereines Preuss Rheinl 26:70–77
Nylander JAA (2004) MrModeltest 2.0. Program distributed by the author. Evolutionary Biology Centre, Uppsala University
Page RD (1996) TreeView: an application to display phylogenetic trees on personal computers. Comput Appl Biosci 12:357–358
Petrak F (1952) Ergebnisse einer Revision der Grundtypen verschiedener Gattungen der Askomyzeten und Fungi Imperfecti. Sydowia 6:336–343
Phillips AJL, Alves A, Pennycook SR, Johnston PR, Ramaley A et al (2008) Resolving the phylogenetic and taxonomic status of dark-spored teleomorph genera in the Botryosphaeriaceae. Persoonia 21:29–55
Phookamsak R, Liu JK, McKenzie EH, Manamgoda DS, Ariyawansa H, Thambugala KM et al (2014) Revision of Phaeosphaeriaceae. Fungal Divers 68(1):159–238.
Pryor BM, Bigelow DM (2003) Molecular characterization of Embellisia and Nimbya species and their relationship to Alternaria, Ulocladium and Stemphylium. Mycologia 95:1141–1154
Pryor BM, Creamer R, Shoemaker RA, McLain-Romero J, Hambleton S (2009) Undifilum, a new genus for endophytic Embellisia oxytropis and parasitic Helminthosporium bornmuelleri on legumes. Botany 87:178–194
Ramaley AW, Barr ME (1995) New dictyosporous species from leaves of Agavaceae. Mycotaxon 54:75–90
Rambaut A, Drummond A (2008) FigTree: Tree figure drawing tool, version 1.2. 2. Institute of Evolutionary Biology, University of Edinburgh
Rannala B, Yang Z (1996) Probability distribution of molecular evolutionary trees: a new method of phylogenetic inference. J Mol Evol 43:304–311
Runa F, Park M, Pryor B (2009) Ulocladium systematics revisited: phylogeny and taxonomic status. Mycol Prog 8:35–47
Schoch CL, Crous PW, Groenewald JZ, Boehm EWA, Burgess TI, de Gruyter J, de Hoog GS, Dixon LJ, Grube M, Gueidan C, Harada Y, Hatakeyama S, Hirayama K, Hosoya T, Huhndorf SM, Hyde KD, Jones EBG, Kohlmeyer J, Kruys Å, Li YM, Lücking R, Lumbsch HT, Marvanová L, Mbatchou JS, McVay AH, Miller AN, Mugambi GK, Muggia L, Nelsen MP, Nelson P, Owensby CA, Phillips AJL, Phongpaichit S, Pointing SB, Pujade-Renaud V, Raja HA, Rivas Plata E, Robbertse B, Ruibal C, Sakayaroj J, Sano T, Selbmann L, Shearer CA, Shirouzu T, Slippers B, Suetrong S, Tanaka K, Volkmann-Kohlmeyer B, Wingfield MJ, Wood AR, Woudenberg JHC, Yonezawa H, Zhang Y, Spatafora JW (2009) A class–wide phylogenetic assessment of dothideomycetes. Stud Mycol 64:1–15
Schoch CL, Robbertse B, Robert V, Vu D, Cardinali G, Irinyi L, Meyer W, Nilsson RH, Hughes K, Miller AN, Kirk PM, Abarenkov K, Aime MC, Ariyawansa HA, Bidartondo M, Boekhout T, Buyck B, Cai Q, Chen J, Crespo A, Crous PW, Damm U, De Beer ZW, Dentinger BTM, Divakar PK, Dueñas M, Feau N, Fliegerova K, García MA, Ge Z-W, Griffith GW, Groenewald JZ, Groenewald M, Grube M, Gryzenhout M, Gueidan C, Guo L, Hambleton S, Hamelin R, Hansen K, Hofstetter V, Hong S-B, Houbraken J, Hyde KD, Inderbitzin P, Johnston PR, Karunarathna SC, Kõljalg U, Kovács GM, Kraichak E, Krizsan K, Kurtzman CP, Larsson K-H, Leavitt S, Letcher PM, Liimatainen K, Liu J-K, Lodge DJ, Luangsa-ard JJ, Lumbsch HT, Maharachchikumbura SSN, Manamgoda D, Martín MP, Minnis AM, Moncalvo J-M, Mulè G, Nakasone KK, Niskanen T, Olariaga I, Papp T, Petkovits T, Pino-Bodas R, Powell MJ, Raja HA, Redecker D, Sarmiento-Ramirez JM, Seifert KA, Shrestha B, Stenroos S, Stielow B, Suh S-O, Tanaka K, Tedersoo L, Telleria MT, Udayanga D, Untereiner WA, Uribeondo JD, Subbarao KV, Vágvölgyi C, Visagie C, Voigt K, Walker DM, Weir BS, Weiß M, Wijayawardene NN, Wingfield MJ, Xu JP, Yang ZL, Zhang N, Zhuang W-Y, Federhen S (2014) Finding needles in haystacks: linking scientific names, reference specimens and molecular data for Fungi. Database 2014: bau061
Seifert KA, Morgan-Jones G, Gams W, Kendrick WB (2011) The genera of hyphomycetes. CBS Biodiversity Series no. 9. Utrecht, CBS-KNAW Fungal Biodiversity Centre
Shimizu K, Tanaka C, Peng YL, Tsuda M (1998) Phylogeny of Bipolaris inferred from nucleotide sequences of Brn1, a reductase gene involved in melanin biosynthesis. J Gen Appl Microbiol 44:251–25
Shoemaker RA (1959) Nomenclature of Drechslera and Bipolaris, grass parasites segregated from Helminthosporium. Can J Bot 37:879–887
Shoemaker RA (1998) Marielliottia, a new genus of cereal and grass parasites segregated from Drechslera. Can J Bot 76(9):1558–1569
Shoemaker RA, Babcock CE (1992) Applanodictyosporous Pleosporales: Clathrospora, Comoclathris, Graphyllium, Macrospora, and Platysporoides. Can J Bot 70:1617–1658
Siboe GM, Kirk PM, Cannon PF (1999) New dematiaceous hyphomycetes from Kenyan rare plants. Mycotaxon 73:283–302
Simmons EG (1986) Gibbago, a new phaeodictyoconidial genus of hyphomycetes. Mycotaxon 27:107–111
Sivanesan A (1984) The bitunicate ascomycetes and their anamorphs. J. Cramer, Vaduz
Sivanesan A (1987) Graminicolous species of Bipolaris, Curvularia, Drechslera, Exserohilum and their teleomorphs. Mycol Pap 158:1–261
Stamatakis A (2006) RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22:2688–269
Stamatakis A, Hoover P, Rougemont J (2008) A rapid bootstrap algorithm for the RAxML web servers. Syst Biol 57:758–777
Stielow B, Bubner B, Hensel G, Munzenberger B, Hoffmann P et al (2010) The neglected hypogeous fungus Hydnotrya bailii Soehner (1959) is a widespread sister taxon of Hydnotrya tulasnei (Berk.) Berk. and Broome (1846). Mycol Prog 9:195–203
Suetrong S, Schoch CL, Spatafora JW, Kohlmeyer J, Volkmann- Kohlmeyer B, Sakayaroj J, Phongpaichit S, Tanaka K, Hirayama K, Jones EBG (2009) Molecular systematics of the marine dothideomycetes. Stud Mycol 64:155–173
Sung G-H, Sung J-M, Hywel-Jones NL, Spatafora JW (2007) A multi-gene phylogeny of Clavicipitaceae (Ascomycota, Fungi): identification of localized incongruence using a combinational bootstrap approach. Mol Phylogenet Evol 44:1204–1223
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28(10):2731–2739
Tan YP, Madrid H, Crous PW, Shivas RG (2014) Johnalcornia gen. et. comb. nov., and nine new combinations in Curvularia based on molecular phylogenetic analysis. Australas Plant Path 1–15
Taylor JW, Jacobson DJ, Kroken S, Kasuga T, Geiser DM et al (2000) Phylogenetic species recognition and species concepts in fungi. Fungal Genet Biol 31:21–32
Telle S, Thines M (2008) Amplification of cox2 (620 bp) from 2 mg of up to 129 years old herbarium specimens, comparing 19 extraction methods and 15 polymerases. PLoS One 3:e3584
Thambugala KM, Ariyawansa HA, Li YM, Boonmee S, Hongsanan S, Tian Q, Singtripop C, Bhat DJ, Camporesi E, Jayawardena R, Liu ZY, Chukeatirote E, Hyde KD (2014a) Dothideales. Fungal Divers 68(1):105–158
Thambugala KM, Singtripop C, Chunfang Y, Mckenzie EH, Liu ZY, Chukeatirote E, Hyde KD (2014b) Towards a natural classification of Dothideomycetes 7: the genera Allosoma, Austropleospora, Dangeardiella, Griggsia and Karschia (Dothideomycetes incertae sedis). Phytotaxa 181(1):34–46
Theissen F, Sydow H (1918) Vorentw?rfe zu den Pseudosphaeriales. Ann Mycol 16:1–34
Udayanga D, Castlebury LA, Rossman AY, Chukeatirote E, Hyde KD (2014) Insights into the genus Diaporthe: phylogenetic species delimitation in the D. eres species complex. Fungal Divers 67(1):203–229
Uilenberg G, Thiaucourt F, Jongejan F (2004) On molecular taxonomy: what is in a name? Exp Appl Acarol 32(4):301–312
Vassiljeva LN (1983) De Buergenerula thalictri (Wint.) E. Müller. Nov Sist Nizsh Rast 20:70–72
Verkley GJM, Dukik K, Renfurm R, Göker M, Stielow JB (2014) Novel genera and species of coniothyrium-like fungi in Montagnulaceae (Ascomycota). Persoonia 32:25–51
von Arx JA, Müller E (1975) A re-evaluation of the bitunicate ascomycetes with keys to families and genera. Stud Mycol 9:1–159
von Arx JA, van der Aa HA (1983) Notes on Curreya (Ascomycetes, Dothideales). Sydowia 36:1–5
von Höhnel F (1918) Dritte vorläufige Mitteilung mycologischer Ergebnisse (nr. 201–304). Ber Deut Bot Ges 36:309–317
Wang Y, Geng Y, Ma J, Wang Q, Zhang X-G (2011) Sinomyces: a new genus of anamorphic Pleosporaceae. Fungal Biol 115:188–195
Wehmeyer LE (1961) A world monograph of the genus pleospora and its segregates. University of Michigan Press, Michigan
Wehmeyer LE (1975) The pyrenomycetous fungi. Mycologia Memoir No. 6. The New York Botanical Garden. J. Cramer, Germany
White TJ, Bruns TD, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. PCR Protocols: Guide Methods Appl 18:315–322
Wijayawardene NN, Crous PW, Kirk PM, Hawksworth DL, Boonmee S, Braun U, Chomnunti P, Dai DQ, D’souza MJ, Diederich P, Dissanayake A, Doilom M, Hongsanan S, Jones EBG, Groenewald JZ, Jayawardena R, Lawrey JD, Liu JK, Lücking R, Madrid H, Manamgoda DS, Muggia L, Nelsen MP, Phookamsak R, Suetrong S, Tanaka K, Thambugala KM, Wikee S, Zhang Y, Aptroot A, Ariyawansa HA, Bahkali AH, Bhat JD, Gueidan C, De Hoog GS, Knudsen K, McKenzie EHC, Miller AN, Mortimer PE, Wanasinghe DN, Phillips AJL, Raja HA, Slippers B, Shivas RS, Taylor JE, Wang Y, Woudenberg JHC, Piątek M, Cai L, Jaklitsch WM, Hyde KD (2014) Naming and outline of dothideomycetes–2014 including proposals for the protection or suppression of generic names. Fungal Divers 69(1):1–55
Woudenberg JHC, Groenewald JZ, Binder M, Crous PW (2013) Alternaria redefined. Stud Mycol 75:171–212
Yusoff M, Moss ST, Jones EBG (1994) Ascospore ultrastructure of Pleospora gaudefroyi (Pleosporaceae, Loculoascomycetes, Ascomycotina). Can J Bot 72(1):1–6
Zhang Y, Schoch CL, Fournier J, Crous PW, De Gruyter J, Woudenberg JHC, Hirayama K, Tanaka K, Pointing SB, Hyde KD (2009) Multi-locus phylogeny of the Pleosporales: a taxonomic, ecological and evolutionary re-evaluation. Stud Mycol 64:85–102
Zhang Y, Crous PW, Schoch CL, Hyde KD (2012) Pleosporales. Fungal Divers 52:1–225
Zhaxybayeva O, Gogarten JP (2002) Bootstrap, Bayesian probability and maximum likelihood mapping: exploring new tools for comparative genome analyses. BMC Genomics 3(1):4
Acknowledgments
MFLU grant number 56101020032 is thanked for supporting studies on Dothideomycetes. We are grateful to the Mushroom Research Foundation, Chiang Rai, Thailand for supporting studies on Dothideomycetes. Kevin D. Hyde thanks the Chinese Academy of Sciences, project number 2013T2S0030, for the award of Visiting Professorship for Senior International Scientists at Kunming Institute of Botany. Jian-Chu Xu and Peter E Mortimer would like to thank Humidtropics, a CGIAR Research Program that aims to develop new opportunities for improved livelihoods in a sustainable environment, for partially funding this work. H.A Ariyawansa and J.C. Kang are grateful to the agricultural science and technology foundation of Guizhou province (Nos. NY[2013]3042 ), the international collaboration plan of Guizhou province (No. G [2012]7006) and the innovation team construction for science and technology of Guizhou province (No. [2012]4007) from the Science and Technology Department of Guizhou province, China. Hiran Ariyawansa is grateful to A.D Ariyawansa, D.M.K Ariyawansa and Dhanuska Udayanga for their valuable suggestions. E.B. Gareth Jones is supported by the Distinguished Scientist Fellowship Program (DSFP), King Saud University, Saudi Arabia.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Table 1
(DOC 253 kb)
Rights and permissions
About this article
Cite this article
Ariyawansa, H.A., Thambugala, K.M., Manamgoda, D.S. et al. Towards a natural classification and backbone tree for Pleosporaceae . Fungal Diversity 71, 85–139 (2015). https://doi.org/10.1007/s13225-015-0323-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13225-015-0323-z