Abstract
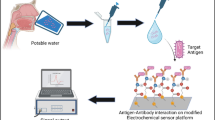
The contamination of food and drinking water by bacteria that cause food poisoning is a critical public health issue. In general, the procedures for testing the presence of pathogenic bacteria in public is not very effective, because it is time consuming and does not facilitate real-time monitoring in a field setting. Therefore, this study introduced a fully automated platform that would allow non-experts to easily examine the major food poisoning bacteria in a field setting, from pretreatment samples to final detection. Two enterobacteria species, of four different bacteria species (i.e., Escherichia coli and Salmonella typhimurium) were successfully immobilized on the surface of magnetic beads by exploiting the specific binding force with mannose. Subsequently, magneto-electrochemical impedance measurement technology allowed bacterial detection with a high sensitivity. E. coli, typical enterobacteria, was detected in 25 min with a detection limit of 100 CFU/mL. Moreover, we demonstrated that the automatic performance and the experimental consistency was greatly improved in comparison with those of manually conducted experiments.






Similar content being viewed by others
References
Mobed, A., Baradaran, B., Guardia, M., Agazadeh, M., Hasanzadeh, M., Rezaee, M.A., Mosafer, J., Mokhtarzadeh, A., Hamblin, M.R.: Advances in detection of fastidious bacteria: from microscopic observation to molecular biosensors. Trends Anal. Chem. 113, 157–171 (2019)
Gracias, K.S., McKillip, J.L.: A review of conventional detection and enumeration methods for pathogenic bacteria in food. Can. J. Microbiol. 50, 883–890 (2004)
Ma, F., Rehman, A., Liu, H., Zhang, J., Zhu, S., Zeng, X.: Glycosylation of quinone-fused polythiophene for reagentless and label-free detection of E. coli. Anal. Chem. 87, 1560–1568 (2015)
Liu, D., Zhu, Y., Li, N., Lu, Y., Cheng, J., Xu, Y.: A portable microfluidic analyzer for integrated bacterial detection using visible loop-mediated amplification. Sens. Actuator B Chem. 310, 127834 (2020)
Nguyen, V.D., Nguyen, H.V., Bui, K.H., Seo, T.S.: Smart phone-powered capillary electrophoresis on a chip for foodborne bacteria detection. Sens. Actuator B Chem. 301, 127108 (2019)
Yoon, T., Moon, H.S., Song, J.W., Hyun, K.A., Jung, H.I.: Automatically controlled microfluidic system for continuous separation of rare bacteria from blood. CYTOM PART A 95, 1135–1144 (2019)
Liu, Q., Zhang, X., Li, X., Liu, S., Sui, G.: A semi-quantitative method for point-of-care assessments of specific pathogenic bioaerosols using a portable microfluidics-based device. J. Aerosol Sci. 115, 173–180 (2018)
Lee, W.I., Park, Y., Park, J., Shrivastava, S., Son, Y.M., Choi, H.J., Lee, J., Jeon, B., Lee, H., Lee, N.E.: A smartphone fluorescence imaging-based mobile biosensing system integrated with a passive fluidic control cartridge for minimal user intervention and high accuracy. Lab Chip. 19, 1502–1511 (2019)
Schulz, M., Calabrese, S., Hausladen, F., Wurn, H., Drossart, D., Stock, K., Sobieraj, A.M., Eichenseher, F., Loessner, M.J., Schmelcher, M., Gerhardts, A., Goetz, U., Handel, M., Serr, A., Haecker, G., Li, J., Specht, M., Koch, P., Meyer, M., Tepper, P., Rother, R., Jehle, M., Wadle, S., Zengerle, R., Stetten, F., Paust, N., Borst, N.: Point-of-care testing system for digital single cell detection of MRSA directly from nasal swabs. Lab Chip. 20, 2549–2561 (2020)
Geng, Z., Gu, Y., Li, S., Lin, B., Liu, P.: A Fully integrated in vitro diagnostic microsystem for pathogen detection developed using a “3D Extensible” microfluidic design paradigm. Micromachines. 10, 873 (2019)
Stumpf, F., Schwemmer, F., Hutzenlaub, T., Baumann, D., Strohmeier, O., Dingemanns, G., Simons, G., Sager, C., Plobner, L., Stetten, F., Zengerle, R., Mark, D.: LabDisk with complete reagent prestorage for sample-to-answer nucleic acid-based detection of respiratory pathogens verified with influenza A H3N2 virus. Lab Chip. 16, 199–207 (2016)
Pignato, S., Marino, A.M., Emanuele, M.C., Lannotta, V., Caracappa, S., Giammanco, G.: Evaluation of new culture media for rapid detection and isolation of salmonellae in foods. Appl. Environ. Microbiol. 61, 1996–1999 (1995)
Fu, C.J., Carter, J.N., Li, Y., Porter, J.H., Kerley, M.S.: Comparison of agar plate and real-time PCR on enumeration of Lactobacillus, Clostridium perfringens and total anaerobic bacteria in dog faeces. Lett. Appl. Microbiol. 42, 490–494 (2006)
Clifford, R.J., Milillo, M., Prestwood, J., Quintero, R., Zurawski, D.V., Kwak, Y.I., Waterman, P.E., Lesho, E.P., Mc Gann, P.: Detection of bacterial 16S rRNA and identification of four clinically important bacteria by real-time PCR. PLoS ONE 7, e48558 (2012)
Yang, J., Zhang, N., Lv, J., Zhu, P., Pan, X., Hu, J., Wu, W., Li, S., Li, H.: Comparing the performance of conventional PCR, RTQ-PCR, and droplet digital PCR assays in detection of Shigella. Mol. Cell. Probes. 51, 101531 (2020)
Kwon, K., Park, J.W., Hyun, K.A., Kwak, B.S., Yong, D.E., Hwang, J., Jung, H.I.: Continuous adsorption and photothermal lysis of airborne bacteria using a gold-nanoparticle-embedded-geometrically activated surface interaction (gold-GASI) chip. Sens. Actuator B Chem. 248, 580–588 (2017)
Kwon, K., Gwak, H., Hyun, K.A., Kwak, B.S., Jung, H.I.: High-throughput microfluidic chip for magnetic enrichment and photothermal DNA extraction of foodborne bacteria. Sens. Actuator B Chem. 294, 62–68 (2019)
Kisiela, D., Laskowska, A., Sapeta, A., Kuczkowski, M., Wieliczko, A., Ugorski, M.: Functional characterization of the FimH adhesin from Salmonella enterica serovar Enteritidis. Microbiology 152, 1337–1346 (2006)
Kline, K.A., Fälker, S., Dahlberg, S., Normark, S., Henriques-Normark, B.: Bacterial adhesins in host-microbe interactions. Cell Host Microbe. 5, 580–592 (2009)
Wang, Q., Kaminska, I., Niedziolka-Jonsson, J., Opallo, M., Li, M., Boukherroub, R., Szunerits, S.: Sensitive sugar detection using 4-aminophenylboronic acid modified graphene. Biosens. Bioelectron. 50, 331–337 (2013)
Wang, B., Anzai, J.: Recent progress in lectin-based biosensors. Materials. 8, 8590–8607 (2015)
Raetz, C.R.H., Whitfield, C.: Lipopolysaccharide endotoxins. Annu. Rev. Biochem. 71, 635–700 (2002)
Abellan-Flos, M., Timmer, B.J.J., Altun, S., Aastrup, T., Vincent, S.P., Ramstrom, O.: QCM sensing of multivalent interactions between lectins and well-defined glycosylated nanoplatforms. Biosens. Bioelectron. 139, 111328 (2019)
Sousa, M.D., Martinez, D.S.T., Alves, O.L.: Alternative mannosylation method for nanomaterials: application to oxidized debris-free multiwalled carbon nanotubes. J. Nanopart. Res. 18, 143 (2016)
Yu, L., Hou, Y., Cheng, C., Schlaich, C., Noeske, P.M., Wei, Q., Haag, R.: High-antifouling polymer brush coatings on nonpolar surfaces via adsorption-cross-linking strategy. ACS Appl. Mater. Interfaces. 9, 44281–44292 (2017)
Telford, J., Barocchi, M., Margarit, I., et al.: Pili in Gram-positive pathogens. Nat Rev Microbiol 4, 509–519 (2006)
Spaulding, C. N., Schreiber IV, H. L., Zheng, W., Dodson, K. W., Hazen, J. E., Conover, M. S., Wang, F., Svenmarker, P., Luna-Rico, A., Francetic, O., Andersson, M., Hultgren, S., Egelman, E. H. Functional role of the type 1 pilus rod structure in mediating host–pathogen interactions. eLife. 7, 1–25 (2018)
Santos, M.B., Agusil, J.P., Prieto-Simon, B., Sporer, C., Teixeira, V., Samitier, J.: Highly sensitive detection of pathogen Escherichia coli O157:H7 by electrochemical impedance spectroscopy. Biosens. Bioelectron. 45, 174–180 (2013)
Wang, L., Huo, X., Qi, W., Xia, Z., Li, Y., Lin, J.: Rapid and sensitive detection of Salmonella Typhimurium using nickel nanowire bridge for electrochemical impedance amplification. Talanta 211, 120715 (2020)
Xu, M., Wang, R., Li, Y.: Rapid detection of Escherichia coli O157:H7 and Salmonella Typhimurium in foods using an electrochemical immunosensor based on screen-printed interdigitated microelectrode and immunomagnetic separation. Talanta 148, 200–208 (2016)
Acknowledgements
This research was supported by the National Research Foundation of Korea(NRF) Grant funded by the Korea Government (MSIT) (No. 2020R1A5A1018052), Korea Environment Industry & Technology Institute (KEITI) through Aquatic Ecosystem Conversion Research Program, funded by Korea Ministry of Environment (MOE) (2020003030007) and Korea Institute of Planning and Evaluation for Technology in Food, Agriculture and Forestry (IPET) through Crop Viruses and Pests Response Industry Technology Development Program, funded by Ministry of Agriculture, Food and Rural Affairs (MAFRA) (320035031HD030).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethical Consideration
Ethical approval was not required for this study, as it was not performed on animals or humans.
Conflict of Interests
There are no conflicts to declare.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Kwon, K., Yoon, T., Gwak, H. et al. Fully Automated System for Rapid Enrichment and Precise Detection of Enterobacteria Using Magneto-Electrochemical Impedance Measurements. BioChip J 15, 233–242 (2021). https://doi.org/10.1007/s13206-021-00024-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13206-021-00024-1