Abstract
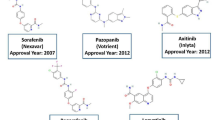
Epidermal growth factor receptor (EGFR) promotes tumorigenic characteristics and activates cancer-associated signaling pathways such as Wnt/-catenin, transforming growth factor (TGF-β), and phosphoinositide-3-kinase (PI3K). Several inhibitors have been reported to suppress the activity of EGFR and are being used in cancer treatment. However, patients in the malignant stage of cancer show resistance to those inhibitors, opening a wide space for research to discover novel inhibitors. Therefore, we carried out machine learning and virtual screening to discover novel inhibitors with high affinity against EGFR-TK. Initially, a library of 2640 chalcones were screened out using a machine-learning model developed based on the random forest algorithm, exhibiting high sensitivity and a Receiver Operating Characteristic curve (ROC area) of 0.99. Furthermore, out of the initial 2640 screened compounds, 412 compounds exhibiting potential activity are subjected to evaluation for drug-likeness properties through different filters: Blood–brain barrier penetration, Lipinski’s rule, CMC-50 like rule, Veber rule, and Ghose filter, alongside Cell Line Cytotoxicity Prediction. A total of 30 compounds that successfully pass through all these filters are selected for molecular docking. Of these, 6 compounds display substantial binding affinity and closer interaction with the conserved catalytic residues of the target EGFR-TK compared to the reference molecule (erlotinib). Furthermore, molecular dynamics simulation studies were conducted on four compounds (CID-375861, CID-375862, CID-23636403, and CID-259166) to confirm the stability of the docked complexes over a 100 ns simulation trajectory. Additionally, the binding free energy calculations by MMPBSA reveal that these four chalcone compounds exhibit strong affinity towards the EGFR-TK enzyme, with binding free energies of − 65.421 kJ/mol, − 94.266 kJ/mol, − 80.044 kJ/mol, and − 79.734 kJ/mol, respectively. The findings from this investigation highlight a set of promising chalcone compounds that have the potential to be developed into effective drugs for the treatment of various cancers. Further research and development on these compounds could pave the way for novel therapeutic interventions.







Similar content being viewed by others
Data availability
All the data cited in this manuscript is generated by the authors and available upon request from the corresponding author.
Code availability
Not applicable.
References
Ani R, Manohar R, Anil G, Deepa OS, (2018) Virtual Screening of Drug Likeness using Tree Based Ensemble Classifier. Biomed Pharmacol J. https://doi.org/10.13005/bpj/1518
Araujo DV, Watson GA, Oliva M et al (2021) Bugs as drugs: the role of microbiome in cancer focusing on immunotherapeutics. Cancer Treat Rev 92:102125. https://doi.org/10.1016/j.ctrv.2020.102125
Arteaga CL, Johnson DH (2001) Tyrosine kinase inhibitors-ZD1839 (Iressa). Curr Opin Oncol 13:491–498. https://doi.org/10.1097/00001622-200111000-00012
Asha Kiranmai S, Jaya Laxmi A (2018) Data mining for classification of power quality problems using WEKA and the effect of attributes on classification accuracy. Prot Control Mod Power Syst 3:29. https://doi.org/10.1186/s41601-018-0103-3
Chauhan SS, Singh AK, Meena S et al (2014) Synthesis of novel β-carboline based chalcones with high cytotoxic activity against breast cancer cells. Bioorg Med Chem Lett 24:2820–2824. https://doi.org/10.1016/j.bmcl.2014.04.109
Connelly PR, Thomson JA (1992) Heat capacity changes and hydrophobic interactions in the binding of FK506 and rapamycin to the FK506 binding protein. Proc Natl Acad Sci 89:4781–4785. https://doi.org/10.1073/pnas.89.11.4781
Cyrański MK, Jezierska A, Klimentowska P et al (2008) Impact of intermolecular hydrogen bond on structural properties of phenylboronic acid: quantum chemical and X-ray study. J Phys Org Chem 21:472–482. https://doi.org/10.1002/poc.1389
Deng W, Huang Z, Zhang J, Xu J (2021) A data mining based system for transaction fraud detection. IEEE Int Conf Consum Electron Comput Eng ICCECE 2021:542–545. https://doi.org/10.1109/ICCECE51280.2021.9342376
Di Lorenzo G, Tortora G, D’Armiento FP et al (2002) Expression of epidermal growth factor receptor correlates with disease relapse and progression to androgen-independence in human prostate cancer. Clin Cancer Res 8:3438–3444
Gacche RN, Dhole NA, Kamble SG, Bandgar BP (2008) In-vitro evaluation of selected chalcones for antioxidant activity. J Enzyme Inhib Med Chem 23:28–31. https://doi.org/10.1080/14756360701306370
Ghose AK, Viswanadhan VN, Wendoloski JJ (1999) A knowledge-based approach in designing combinatorial or medicinal chemistry libraries for drug discovery. 1. A qualitative and quantitative characterization of known drug databases. J Comb Chem 1:55–68. https://doi.org/10.1021/cc9800071
Guleria V, Pal T, Sharma B et al (2021) Pharmacokinetic and molecular docking studies to design antimalarial compounds targeting Actin I. Int J Health Sci (qassim) 15:4–15
Hennequin LF, Stokes ESE, Thomas AP et al (2002) Novel 4-anilinoquinazolines with C-7 basic side chains: design and structure activity relationship of a series of potent, orally active, VEGF receptor tyrosine kinase inhibitors. J Med Chem 45:1300–1312. https://doi.org/10.1021/jm011022e
Hirsch FR, Scagliotti GV, Langer CJ et al (2003) Epidermal growth factor family of receptors in preneoplasia and lung cancer: perspectives for targeted therapies. Lung Cancer 41:29–42. https://doi.org/10.1016/S0169-5002(03)00137-5
Holbro T, Hynes NE (2004) ErbB Receptors: directing key signaling networks throughout life. Annu Rev Pharmacol Toxicol 44:195–217. https://doi.org/10.1146/annurev.pharmtox.44.101802.121440
Housman G, Byler S, Heerboth S et al (2014) Drug resistance in cancer: an overview. Cancers (basel) 6:1769–1792. https://doi.org/10.3390/cancers6031769
Huang M, Shen A, Ding J, Geng M (2014) Molecularly targeted cancer therapy: some lessons from the past decade. Trends Pharmacol Sci 35:41–50. https://doi.org/10.1016/j.tips.2013.11.004
Jain U, Bhatia R, Rao A et al (2014) Design and development of halogenated chalcone derivatives as potential anticancer agents. Trop J Pharm Res 13:73. https://doi.org/10.4314/tjpr.v13i1.11
Kalmegh S (2015) Analysis of WEKA data mining algorithm REPTree, simple cart and RandomTree for classification of Indian news. Int J Innov Sci Eng Technol 2:438–446
Ke X, Shen L (2017) Molecular targeted therapy of cancer: the progress and future prospect. Front Lab Med 1:69–75. https://doi.org/10.1016/j.flm.2017.06.001
Kersemaekers AM, Fleuren GJ, Kenter GG et al (1999) Oncogene alterations in carcinomas of the uterine cervix: overexpression of the epidermal growth factor receptor is associated with poor prognosis. Clin Cancer Res 5:577–586
Kolundžija B, Marković V, Stanojković T et al (2014) Novel anthraquinone based chalcone analogues containing an imine fragment: synthesis, cytotoxicity and anti-angiogenic activity. Bioorg Med Chem Lett 24:65–71. https://doi.org/10.1016/j.bmcl.2013.11.075
Kostal J, (2016) Computational Chemistry in Predictive Toxicology. pp 139–186
Kumari R, Kumar R, Lynn A (2014) g_mmpbsa —A GROMACS tool for high-throughput MM-PBSA calculations. J Chem Inf Model 54:1951–1962. https://doi.org/10.1021/ci500020m
Lagunin AA, Dubovskaja VI, Rudik AV et al (2018) CLC-Pred: A freely available web-service for in silico prediction of human cell line cytotoxicity for drug-like compounds. PLoS ONE 13:e0191838. https://doi.org/10.1371/journal.pone.0191838
Lee SH, Seo GS, Kim JY et al (2006) Heme oxygenase 1 mediates anti-inflammatory effects of 2′,4′,6′-tris(methoxymethoxy) chalcone. Eur J Pharmacol 532:178–186. https://doi.org/10.1016/j.ejphar.2006.01.005
Lipinski CA (2000) Drug-like properties and the causes of poor solubility and poor permeability. J Pharmacol Toxicol Methods 44:235–249. https://doi.org/10.1016/S1056-8719(00)00107-6
Madhavi Sastry G, Adzhigirey M, Day T et al (2013) Protein and ligand preparation: parameters, protocols, and influence on virtual screening enrichments. J Comput Aided Mol Des 27:221–234. https://doi.org/10.1007/s10822-013-9644-8
Matsuda N, Lim B, Wang X, Ueno NT (2017) Early clinical development of epidermal growth factor receptor targeted therapy in breast cancer. Expert Opin Investig Drugs 26:463–479. https://doi.org/10.1080/13543784.2017.1299707
Moyer JD, Barbacci EG, Iwata KK et al (1997) Induction of apoptosis and cell cycle arrest by CP-358,774, an inhibitor of epidermal growth factor receptor tyrosine kinase. Cancer Res 57:4838–4848
Nowakowska Z (2007) A review of anti-infective and anti-inflammatory chalcones. Eur J Med Chem 42:125–137. https://doi.org/10.1016/j.ejmech.2006.09.019
Nowakowska Z, Kędzia B, Schroeder G (2008) Synthesis, physicochemical properties and antimicrobial evaluation of new (E)-chalcones. Eur J Med Chem 43:707–713. https://doi.org/10.1016/j.ejmech.2007.05.006
O’Boyle NM, Banck M, James CA et al (2011) Open Babel: An open chemical toolbox. J Cheminform 3:33. https://doi.org/10.1186/1758-2946-3-33
Ogiso H, Ishitani R, Nureki O et al (2002) Crystal structure of the complex of human epidermal growth factor and receptor extracellular domains. Cell 110:775–787. https://doi.org/10.1016/S0092-8674(02)00963-7
Pronk S, Páll S, Schulz R et al (2013) GROMACS 4.5: a high-throughput and highly parallel open source molecular simulation toolkit. Bioinformatics 29:845–854. https://doi.org/10.1093/bioinformatics/btt055
Pryczynicz A, Guzińska-Ustymowicz K, Kemona A, Czyzewska J Expression of EGF and EGFR strongly correlates with metastasis of pancreatic ductal carcinoma. Anticancer Res 28:1399–404
Rabindran SK, Discafani CM, Rosfjord EC et al (2004) Antitumor activity of HKI-272, an orally active, irreversible inhibitor of the HER-2 tyrosine kinase. Cancer Res 64:3958–3965. https://doi.org/10.1158/0008-5472.CAN-03-2868
Rao C, Yejella RP, Rehman R, Basha SH (2015) Molecular docking based screening of novel designed chalcone series of compounds for their anti-cancer activity targeting EGFR kinase domain. Bioinformation 11:322–329. https://doi.org/10.6026/97320630011322
Rusnak DW, Lackey K, Affleck K et al (2001) The effects of the novel, reversible epidermal growth factor receptor/ErbB-2 tyrosine kinase inhibitor, GW2016, on the growth of human normal and tumor-derived cell lines in vitro and in vivo. Mol Cancer Ther 1:85–94
Sharma SV, Settleman J (2007) Oncogene addiction: setting the stage for molecularly targeted cancer therapy. Genes Dev 21:3214–3231. https://doi.org/10.1101/gad.1609907
Siddiqui ZN, Praveen S, Musthafa TNM et al (2012) Thermal solvent-free synthesis of chromonyl chalcones, pyrazolines and their in vitro antibacterial, antifungal activities. J Enzyme Inhib Med Chem 27:84–91. https://doi.org/10.3109/14756366.2011.577035
Singh P, Anand A, Kumar V (2014) Recent developments in biological activities of chalcones: a mini review. Eur J Med Chem 85:758–777. https://doi.org/10.1016/j.ejmech.2014.08.033
Smaill JB, Rewcastle GW, Loo JA et al (2000) Tyrosine kinase inhibitors. 17. irreversible inhibitors of the epidermal growth factor receptor: 4-(Phenylamino)quinazoline- and 4-(Phenylamino)pyrido[3,2- d ]pyrimidine-6-acrylamides bearing additional solubilizing functions. J Med Chem 43:1380–1397. https://doi.org/10.1021/jm990482t
Stamos J, Sliwkowski MX, Eigenbrot C, (2002) Structure of the Epidermal Growth Factor Receptor Kinase Domain Alone and in Complex with a 4-Anilinoquinazoline Inhibitor *. 277:46265–46272. https://doi.org/10.1074/jbc.M207135200
Syam S, Abdelwahab SI, Al-Mamary MA, Mohan S (2012) Synthesis of chalcones with anticancer activities. Molecules 17:6179–6195. https://doi.org/10.3390/molecules17066179
Thakur A, Sharma B, Parashar A et al (2023) (2023) 2D-QSAR, molecular docking and MD simulation based virtual screening of the herbal molecules against Alzheimer’s disorder: an approach to predict CNS activity. J Biomol Struct Dyn. https://doi.org/10.1080/07391102.2192805
Vanommeslaeghe K, Hatcher E, Acharya C et al (2009) CHARMM general force field: a force field for drug-like molecules compatible with the CHARMM all-atom additive biological force fields. J Comput Chem NA-NA. https://doi.org/10.1002/jcc.21367
Vanommeslaeghe K, Hatcher E, Acharya C et al (2010) CHARMM general force field: A force field for drug-like molecules compatible with the CHARMM all-atom additive biological force fields. J Comput Chem 31:671–690
Veber DF, Johnson SR, Cheng H-Y et al (2002) Molecular properties that influence the oral bioavailability of drug candidates. J Med Chem 45:2615–2623. https://doi.org/10.1021/jm020017n
Wan M, Xu L, Hua L et al (2014) Synthesis and evaluation of novel isoxazolyl chalcones as potential anticancer agents. Bioorg Chem 54:38–43. https://doi.org/10.1016/j.bioorg.2014.03.004
Wee P, Wang Z (2017) Epidermal growth factor receptor cell proliferation signaling pathways. Cancers (basel) 9:52. https://doi.org/10.3390/cancers9050052
Xu G, Abad MC, Connolly PJ et al (2008a) 4-Amino-6-arylamino-pyrimidine-5-carbaldehyde hydrazones as potent ErbB-2/EGFR dual kinase inhibitors. Bioorg Med Chem Lett 18:4615–4619. https://doi.org/10.1016/j.bmcl.2008.07.020
Xu G, Searle LL, Hughes TV et al (2008b) Discovery of novel 4-amino-6-arylaminopyrimidine-5-carbaldehyde oximes as dual inhibitors of EGFR and ErbB-2 protein tyrosine kinases. Bioorg Med Chem Lett 18:3495–3499. https://doi.org/10.1016/j.bmcl.2008.05.024
Yang C-H, Chou H-C, Fu Y-N et al (2015) EGFR over-expression in non-small cell lung cancers harboring EGFR mutations is associated with marked down-regulation of CD82. Biochim Biophys Acta - Mol Basis Dis 1852:1540–1549. https://doi.org/10.1016/j.bbadis.2015.04.020
Yap CW (2011) PaDEL-descriptor: An open source software to calculate molecular descriptors and fingerprints. J Comput Chem 32:1466–1474. https://doi.org/10.1002/jcc.21707
Yun C-H, Boggon TJ, Li Y et al (2007) Structures of lung cancer-derived EGFR mutants and inhibitor complexes: mechanism of activation and insights into differential inhibitor sensitivity. Cancer Cell 11:217–227. https://doi.org/10.1016/j.ccr.2006.12.017
Zhang X, Gureasko J, Shen K et al (2006) An allosteric mechanism for activation of the kinase domain of epidermal growth factor receptor. Cell 125:1137–1149. https://doi.org/10.1016/j.cell.2006.05.013
Acknowledgements
The authors would like to thank the Head of the Botany Department Soban Singh Jeena University, S.S.J. Campus, Almora, India, for providing the requisite facilities to conduct this research work. Department of biotechnology, Bhimtal campus, (Kumaun University), is also acknowledged by the authors for providing high-speed Internet facilities. We also extend our appreciation to Rashtriya Uchchattar Shiksha Abhiyan(RUSA), Ministry of Human Resource Development, Government of India, for the deployment of computer infrastructure to establish Bioinformatics Centre in Kumaun University, S.S.J. Campus, Almora.
Funding
This research is not supported by any funding source from the public, commercial, or not-for-profit sectors.
Author information
Authors and Affiliations
Contributions
SM wrote the manuscript. TJ and PS has done all experiments parts. PM and MN analyzed and interpreted data. SC conceptualized and designed the project. VP and SC supervised the study. The whole manuscript was approved by all authors.
Corresponding author
Ethics declarations
Conflict of interest
The authors report there are no competing interests to declare.
Ethical approval
Not applicable.
Informed consent
Not applicable. No human trials involved.
Research involving human participants and/or animals
Not applicable. No human or animal trials were conducted.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Mathpal, S., Joshi, T., Sharma, P. et al. In silico screening of chalcone derivatives as promising EGFR-TK inhibitors for the clinical treatment of cancer. 3 Biotech 14, 18 (2024). https://doi.org/10.1007/s13205-023-03858-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-023-03858-8