Abstract
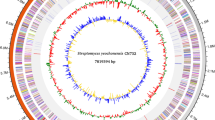
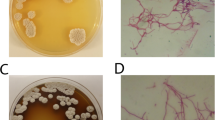
The present study scrutinizes the presence of Streptomyces strains in the soil sample collected from industrial area of Bahadurgarh (Haryana) India. The morphological approach manifested the isolated strain belong to Streptomyces species and named as Streptomyces sp. KD18. Sequencing of Streptomyces sp. KD18 genome was performed by Illumina Nextseq500 platform. 65 contigs were generated via SPAdes v3.11.1 and harboured genome size of 7.2 Mb. AntiSMASH server revealed the presence of 25 biosynthetic gene clusters in KD18 genome where BGC of lipstatin was of more interest from industrial and pharmaceutical purpose. The draft genome sequence represented via ANI values claimed that the KD18 strain belongs to Streptomyces toxytricini and finally named as S. toxytricini KD18. The LC–MS analysis of the extracted metabolite confirmed the production of lipstatin. The genome sequence data have been deposited to NCBI under the accession number of GCA_014748315.1.




Similar content being viewed by others
Data availability
This whole-genome shotgun project has been deposited at DDBJ/ENA/GenBank under the accession number GCA_014748315.1. (BioProject number PRJNA662161 and BioSample number SAMN16077200). The version described in this paper is JACVWW010000000 and consists of sequences JACVWW010000001 to JACVWW010000065. The raw reads are available in NCBI under BioProject accession number PRJNA662161.
References
Aziz RK, Bartels D, Best AA et al (2008) The RAST Server: rapid annotations using subsystems technology. BMC Genomics 9:1–15
Baltz RH (2019) Natural product drug discovery in the genomic era: realities, conjectures, misconceptions, and opportunities. J Ind Microbiol Biotechnol 46:281–299
Bankevich A, Nurk S, Antipov D et al (2012) SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol 19:455–477
Beld J, Sonnenschein EC, Vickery CR et al (2014) The phosphopantetheinyl transferases: catalysis of a post-translational modification crucial for life. Nat Prod Rep 31:61–108
Benaud N, Edwards RJ, Amos TG et al (2021) Antarctic desert soil bacteria exhibit high novel natural product potential, evaluated through long-read genome sequencing and comparative genomics. Environ Microbiol 23:3646–3664
Bible KC, Ryder M (2016) Evolving molecularly targeted therapies for advanced-stage thyroid cancers. Nat Rev Clin Oncol 13:403–416
Boetzer M, Henkel C, Jansen HJ et al (2011) Scaffolding pre-assembled contigs using SSPACE. Bioinformatics 27:578–579
Boldogköi Z, Murvai J, Fodor I (1995) G and C accumulation at silent positions of codons produces additional ORFs. Trends Genet 11:125–126
Chaudhari NM, Gupta VK, Dutta C (2016) BPGA-an ultra-fast pan-genome analysis pipeline. Sci Rep 6:1–10
Chen AY, Jemal A, Ward EM (2009) Increasing incidence of differentiated thyroid cancer in the United States, 1988–2005. Cancer 115:3801–3807
Choi YJ, Lee J-E, Ji HD et al (2021) Tunicamycin as a novel redifferentiation agent in radioiodine therapy for anaplastic thyroid cancer. Int J Mol Sci 22:1077
Claessen D, Rozen DE, Kuipers OP et al (2014) Bacterial solutions to multicellularity: a tale of biofilms, filaments and fruiting bodies. Nat Rev Microbiol 12:115–124
Galanie S, Entwistle D, Lalonde J (2020) Engineering biosynthetic enzymes for industrial natural product synthesis. Nat Prod Rep 37:1122–1143
Goodfellow M (2012) Phylum XXVI. Actinobacteria phyl. nov. In: Bergey’s manual® of systematic bacteriology. Springer, pp 33–2028
Goris J, Konstantinidis KT, Klappenbach JA et al (2007) DNA–DNA hybridization values and their relationship to whole-genome sequence similarities. Int J Syst Evol Microbiol 57:81–91
Goto Y, Li B, Claesen J et al (2010) Discovery of unique lanthionine synthetases reveals new mechanistic and evolutionary insights. PLoS Biol 8:e1000339
Hahn J, Seeber F, Kolodziej H et al (2013) High sensitivity of Giardia duodenalis to tetrahydrolipstatin (orlistat) in vitro. PLoS One 8:e71597
Ikeda H, Ishikawa J, Hanamoto A et al (2003) Complete genome sequence and comparative analysis of the industrial microorganism Streptomyces avermitilis. Nat Biotechnol 21:526–531
Kim J-N, Kim Y, Jeong Y et al (2015) Comparative genomics reveals the core and accessory genomes of Streptomyces species. J Microbiol Biotechnol 25:1599–1605
Komaki H, Sakurai K, Hosoyama A et al (2018) Diversity of nonribosomal peptide synthetase and polyketide synthase gene clusters among taxonomically close Streptomyces strains. Sci Rep 8:1–11
Konstantinidis KT, Tiedje JM (2005) Towards a genome-based taxonomy for prokaryotes. J Bacteriol 187:6258–6264
Kumar MS, Verma V, Soni S, Rao ASP (2012) Identification and process development for enhanced production of Lipstatin from Streptomyces toxytricini (Nrrl 15443) and further downstream processing. Adv Bio Tech 11:11
Labeda DP, Goodfellow M, Brown R et al (2012) Phylogenetic study of the species within the family Streptomycetaceae. Antonie Van Leeuwenhoek 101:73–104
Letunic I, Bork P (2021) Interactive Tree Of Life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res 49:W293–W296
Luo Y, Cobb RE, Zhao H (2014) Recent advances in natural product discovery. Curr Opin Biotechnol 30:230–237
Majer HM, Ehrlich RL, Ahmed A et al (2021) Whole genome sequencing of Streptomyces actuosus ISP-5337, Streptomyces sioyaensis B-5408, and Actinospica acidiphila B-2296 reveals secondary metabolomes with antibiotic potential. Biotechnol Reports 29:e00596
Medema MH, Blin K, Cimermancic P et al (2011) antiSMASH: rapid identification, annotation and analysis of secondary metabolite biosynthesis gene clusters in bacterial and fungal genome sequences. Nucleic Acids Res 39:W339–W346
Mulzer M, Tiegs BJ, Wang Y et al (2014) Total synthesis of tetrahydrolipstatin and stereoisomers via a highly regio-and diastereoselective carbonylation of epoxyhomoallylic alcohols. J Am Chem Soc 136:10814–10820
Nikodinovic J, Barrow KD, Chuck JA (2003) High yield preparation of genomic DNA from Streptomyces. Biotechniques 35:932–936. https://doi.org/10.2144/03355bm05
Seemann T (2014) Prokka: rapid prokaryotic genome annotation. Bioinformatics 30:2068–2069
Seipke RF (2015) Strain-level diversity of secondary metabolism in Streptomyces albus. PLoS One 10:e0116457
Shiraishi T, Yoshida T, Nakata S et al (2005) Tunicamycin enhances tumor necrosis factor–related apoptosis-inducing ligand–induced apoptosis in human prostate cancer cells. Cancer Res 65:6364–6370
Sladič G, Urukalo M, Kirn M et al (2014) Identification of lipstatin-producing ability in Streptomyces virginiae CBS 314.55 using dereplication approach. Food Technol Biotechnol 52:276–284
Stalder H, Oesterhelt G, Borgström B (1992) Tetrahydrolipstatin: degradation products produced by human carboxyl-ester lipase. Helv Chim Acta 75:1593–1603
Telfer TJ, Richardson-Sanchez T, Gotsbacher MP et al (2019) Analogues of desferrioxamine B (DFOB) with new properties and new functions generated using precursor-directed biosynthesis. Biometals 32:395–408
Vicente CM, Thibessard A, Lorenzi J-N et al (2018) Comparative genomics among closely related Streptomyces strains revealed specialized metabolite biosynthetic gene cluster diversity. Antibiotics 7:86
Wang Y, Coleman-Derr D, Chen G, Gu YQ (2015) OrthoVenn: a web server for genome wide comparison and annotation of orthologous clusters across multiple species. Nucleic Acids Res 43:W78–W84
Williams GJ (2013) Engineering polyketide synthases and nonribosomal peptide synthetases. Curr Opin Struct Biol 23:603–612
Wu M, Scott AJ (2012) Phylogenomic analysis of bacterial and archaeal sequences with AMPHORA2. Bioinformatics 28:1033–1034
Wyszynski FJ, Lee SS, Yabe T et al (2012) Biosynthesis of the tunicamycin antibiotics proceeds via unique exo-glycal intermediates. Nat Chem 4:539–546
Yang H, Zhang Z, Yan L et al (2019) Characterization of lipstatin and the minor components from Streptomyces toxytricini fermentation broth by HPLC–ESI–Q-TOF–MS. Chromatographia 82:1791–1800
Zerouki C, Bensalah F, Kuittinen S et al (2021) Whole-genome sequencing of two Streptomyces strains isolated from the sand dunes of Sahara. BMC Genomics 22:1–21. https://doi.org/10.1186/s12864-021-07866-x
Acknowledgements
The authors sincerely acknowledge the Central University of Haryana, Mahendergarh for necessary facilities and PhiXgen Pvt. Ltd. for the support in the analysis. We also wish to acknowledge University Grant Commission (UGC), New Delhi, for doctoral fellowship.
Author information
Authors and Affiliations
Contributions
Conceptualization: KKD; methodology: NS; investigation: K and ND; resources: KKD and VG; writing—original draft preparation: K; writing—review and editing: all authors.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest in the publication.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Consent for publication
Not applicable.
Accession number
The genome sequence data have been deposited to NCBI under the accession number of GCA_014748315.1.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Khushboo, Singhvi, N., Gupta, V. et al. Draft genome sequence of Streptomyces sp. KD18, isolated from industrial soil. 3 Biotech 13, 34 (2023). https://doi.org/10.1007/s13205-022-03453-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-022-03453-3