Abstract
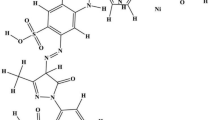
In the current investigation, the capacity of different yeast strains to decolorize reactive black 5 (RB-5) was assessed. A comparative study between the different strains demonstrated that Saccharomyces cerevisiae X19G2 exhibited the highest decolorization rate (69.20 ± 1.16%) after 48 h of incubation. This strain was selected to optimize the medium components’ concentrations for maximum RB-5 decolorization. Response-surface methodology (RSM) was tested for the most significant parameters (glucose, yeast extract and RB-5 dye concentrations) that were previously determined by Plackett–Burman design. A dye decolorization rate of 99.59 ± 0.24% was achieved within 48 h using a maximum RB-5 concentration (0.15 g/L) with glucose and yeast extract concentrations equalling to 10.5 g/L and 1 g/L, respectively. Experimental data results proved to fit well with the pseudo-second order kinetics model. The phytotoxicity assessment was carried out using Raphanus sativus seeds to determine the toxicity of RB-5 before and after treatment by S. cerevisiae. Results suggested that germination rate and the length of seeds radical irrigated with 0.15 g/L of RB-5 decreased by 30 and 53%, compared to those irrigated with treated solution. Therefore, metabolites derived from decolorization of RB-5 by S. cerevisiae X19G2 were significantly less toxic than the original dye.




Similar content being viewed by others
References
Al-Tohamy R, Sun J, Fareed MF, Kenawy E, Ali SS (2020) Ecofriendly biodegradation of reactive black 5 by newly isolated Sterigmatomyces halophilus SSA1575, valued for textile azo dye wastewater processing and detoxification. Sci Rep 10:12370. https://doi.org/10.1038/s41598-020-69304-4
Ameenudeen S, Unnikrishnan S, Ramalingam K (2021) Statistical optimization for the efficacious degradation of reactive azo dyes using Acinetobacter baumannii JC359. J Environ Manag 279:111512. https://doi.org/10.1016/j.jenvman.2020
Bankole PO, Adekunle AA, Govindwar SP (2018) Enhanced decolorization and biodegradation of acid red 88 dye by newly isolated fungus, Achaetomium strumarium. J Environ Chem Eng 6:1589–1600. https://doi.org/10.1016/j.jece.2018.01.069
Beluci NDCL, Mateus GAP, Miyashiro CS et al (2019) Hybrid treatment of coagulation/flocculation process followed by ultrafiltration in TiO2-modified membranes to improve the removal of reactive black 5 dye. Sci Total Environ 664:222–229. https://doi.org/10.1016/j.scitotenv.2019.01.199
Ben Atitallah I, Arous F, Louati I et al (2021) Efficient bioethanol production from date palm (Phoenix dactylifera L.) sap by a newly isolated Saccharomyces cerevisiae X19G2. Process Biochem 105:102–112. https://doi.org/10.1016/j.procbio.2021.03.019
Das A, Mishra S (2017) Removal of textile dye reactive green-19 using bacterial consortium: process optimization using response surface methodology and kinetics study. J Environ Chem Eng 5:612–627. https://doi.org/10.1016/j.jece.2016.10.005
Droguett T, Mora-Gómez J, García-Gabaldón M et al (2020) Electrochemical degradation of reactive black 5 using two-different reactor configuration. Sci Rep 10:1–11. https://doi.org/10.1038/s41598-020-61501-5
Emadi Z, Sadeghi R, Forouzandeh S et al (2021) Simultaneous anaerobic decolorization/degradation of reactive black-5 azo dye and chromium(VI) removal by Bacillus cereus strain MS038EH followed by UV-C/H2O2 post-treatment for detoxification of biotransformed products. Arch Microbiol 203:4993–5009. https://doi.org/10.1007/s00203-021-02462-9
Fadaei S, Noorisepehr M, Pourzamani H et al (2021) Heterogeneous activation of peroxymonosulfate with Fe3O4 magnetic nanoparticles for degradation of reactive black 5: batch and column study. J Environ Chem Eng 9(4):105414. https://doi.org/10.1016/j.jece.2021.105414
Felista MM, Wanyonyi WC, Ongera G (2020) Adsorption of anionic dye (reactive black 5) using macadamia seed husks: kinetics and equilibrium studies. Sci Afr 7:e00283. https://doi.org/10.1016/j.sciaf.2020.e00283
Ghaedi M, Hajati S, Barazesh B, Krimi F, Ghezelbash G (2013) Saccharomyces cerevisiae for the biosorption of basic dyes from binary component systems and the high order derivative spectrophotometric method for simultaneous analysis of Brilliant green and Methylene blue. J Ind Eng Chem 19:227–233. https://doi.org/10.1016/j.jiec.2012.08.006
Guo G, Tian F, Zhao Y et al (2019) Aerobic decolorization and detoxification of Acid Scarlet GR by a newly isolated salt-tolerant yeast strain Galactomyces geotrichum GG. Int Biodeterior Biodegrad 145:104818. https://doi.org/10.1016/j.ibiod.2019.104818
Hameed BB, Ismail ZZ (2018) Decolorization, biodegradation and detoxification of reactive red azo dye using non-adapted immobilized mixed cells. Biochem Eng J 137:71–77. https://doi.org/10.1016/j.bej.2018.05.018
Heo N, Jun Y, Park JH (2013) Dye molecules in electrolytes: new approach for suppression of dye-desorption in dye-sensitized solar cells. Sci Rep 3(1):1–6. https://doi.org/10.1038/srep01712
Herrero R, Lodeiro P, Rojo R, Ciorba A, Rodríguez P, de Vicente MES (2008) The efficiency of the red alga Mastocarpus stellatus for remediation of cadmium pollution. Bioresour Technol 99:4138–4146. https://doi.org/10.1016/j.biortech.2007.08.065
Jadhav JP, Govindwar SP (2006) Biotransformation of malachite green by Saccharomyces cerevisiae MTCC 463. Yeast 23:315–323. https://doi.org/10.1002/yea.1356
Jadhav JP, Parshetti GK, Kalme SD, Govindwar SP (2007) Decolourization of azo dye methyl red by Saccharomyces cerevisiae MTCC 463. Chemosphere 68:394–400. https://doi.org/10.1016/j.chemosphere.2006.12.087
Khan SA, Mehmood S, Iqbal A, Hamayun M (2020) Industrial polluted soil borne fungi decolorize the recalcitrant azo dyes synozol red HF–6BN and synozol black B. Ecotoxicol Environ Saf 206:111381. https://doi.org/10.1016/j.ecoenv.2020.111381
Kiayi Z, Lotfabad TB, Heidarinasab A, Shahcheraghi F (2019) Microbial degradation of azo dye carmoisine in aqueous medium using Saccharomyces cerevisiae ATCC 9763. J Hazard Mater 373:608–619. https://doi.org/10.1016/j.jhazmat.2019.03.111
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33(7):1870–1874
Kumar N, Sinha S, Mehrotra T, Singh R, Tandon S, Thakur IS (2019) Biodecolorization of azo dye acid black 24 by Bacillus pseudomycoides: process optimization using Box Behnken design model and toxicity assessment. Bioresour Technol Rep 8:100311. https://doi.org/10.1016/j.biteb.2019.100311
Lim CK, Bay HH, Aris A, Abdul Majid Z, Ibrahim Z (2013) Biosorption and biodegradation of acid orange 7 by Enterococcus faecalis strain ZL: optimization by response surface methodological approach. Environ Sci Pollut Res 20:5056–5066. https://doi.org/10.1007/s11356-013-1476-5
Louati I, Fersi M, Hadrich B, Ghariani B, Nasri M, Mechichi T (2018) Prickly pear cactus cladodes powder of Opuntia ficus indica as a cost effective biosorbent for dyes removal from aqueous solutions. 3 Biotech 8(11):1–10. https://doi.org/10.1007/s13205-018-1499-1
Louati I, Hadrich B, Nasri M, Belbahri L, Woodward S, Mechichi T (2019) Modelling of reactive black 5 decolourization in the presence of heavy metals by the newly isolated Pseudomonas aeruginosa strain Gb30. J Appl Microbiol 126:1761–1771. https://doi.org/10.1111/jam.14262
Louati I, Elloumi-Mseddi J, Cheikhrouhou W, Hadrich B, Nasri M, Aifa S, Woodward S, Mechichi T (2020) Simultaneous cleanup of reactive black 5 and cadmium by a desert soil bacterium. Ecotoxicol Environ Saf 190:110103. https://doi.org/10.1016/j.ecoenv.2019.110103
Maheshwari N, Kumar M, Thakur IS, Srivastava S (2018) Carbon dioxide biofixation by free air CO2 enriched (FACE) bacterium for biodiesel production. J CO2 Util 27:423–432. https://doi.org/10.1016/j.jcou.2018.08.010
Mahmoud ME, Khalifa MA, El-Mallah NM, Hassouba HM, Nabil GM (2022) Performance of MnO2 nanoparticles-coated cationic CTAB for detoxification and decolorization of sulfonated remazol red and reactive black 5 dyes from water. Int J Environ Sci Technol 19:141–158. https://doi.org/10.1007/s13762-021-03153-0
Malairuang K, Krajang M, Sukna J, Rattanapradit K, Chamsart S (2020) High cell density cultivation of Saccharomyces cerevisiae with intensive multiple sequential batches together with a novel technique of fed-batch at cell level (FBC). Processes 8:1321. https://doi.org/10.3390/pr8101321
Marcharchand S, Ting ASY (2017) Trichoderma asperellum cultured in reduced concentrations of synthetic medium retained dye decolourization efficacy. J Environ Manag 203:542–549. https://doi.org/10.1016/j.jenvman.2017.06.068
Martorell MM, Pajot HF, Ahmed PM, de Figueroa LIC (2017) Biodecoloration of reactive black 5 by the methylotrophic yeast Candida boidinii MM 4035. J Environ Sci (china) 53:78–87. https://doi.org/10.1016/j.jes.2016.01.033
Mathivanan M, Chinnaiah S, Rs SS (2018) Dye degradation using Saccharomyces cerevisiae. Int J Eng Technol 7(3–12):180–184
Mitter EK, Corso CR (2013) Acid dye biodegradation using saccharomyces cerevisiae immobilized with polyethyleneimine-treated sugarcane bagasse. Water Air Soil Pollut 224(1):1–8. https://doi.org/10.1007/s11270-012-1391-2
M-Ridha MJ, Hussein SI, Alismaeel ZT, Atiya MA, Aziz GM (2020) Biodegradation of reactive dyes by some bacteria using response surface methodology as an optimization technique. Alex Eng J 59(5):3551–3563. https://doi.org/10.1016/j.aej.2020.06.001
Navaei A, Yazdani M, Alidadi H, Dankooba M, Bonyadid Z, Dehghand A, Hosseinie A (2019) Biosorption of reactive red 120 dye from aqueous solution using Saccharomyces cerevisiae: RSM analysis, isotherms and kinetic studies. Desalin Water Treat 171:418–427. https://doi.org/10.5004/dwt.2019.24780
Ramalho PA, Cardoso MH, Cavaco-Paulo A, Ramalho MT (2004) Characterization of azo reduction activity in a novel ascomycete yeast strain. Appl Environ Microbiol 70:2279–2288. https://doi.org/10.1128/AEM.70.4.2279-2288.2004
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425. https://doi.org/10.1093/oxfordjournals.molbev.a040454
Samarbaf S, Tahmasebi Birgani Y, Yazdani M, Babaei AA (2019) A comparative removal of two dyes from aqueous solution using modified oak waste residues: process optimization using response surface methodology. J Ind Eng Chem 73:67–77. https://doi.org/10.1016/j.jiec.2018.12.011
Santos J, Leitão-Correia F, Sousa MJ, Leão C (2016) Nitrogen and carbon source balance determines longevity, independently of fermentative or respiratory metabolism in the yeast Saccharomyces cerevisiae. Oncotarget. https://doi.org/10.18632/oncotarget.8656
Saratale RG, Saratale GD, Chang JS, Govindwar SP (2009) Ecofriendly degradation of sulfonated diazo dye C.I. Reactive Green 19A using Micrococcus glutamicus NCIM-2168. Bioresour Technol 100:3897–3905. https://doi.org/10.1016/j.biortech.2009.03.051
Sharma B, Tiwari S, Bisht N, Tewari L (2021) Eco-friendly bioprocess using agar plug immobilized Penicillium crustosum PWWS-6 biomass for treatment of wastewater contaminated with toxic Congo red dye for use in agriculture. Ind Crops Prod 170:133755. https://doi.org/10.1016/j.indcrop.2021.113755
Singhal A, Kumar M, Bhattacharya M, Kumari N, Jha PK, Chauhan DK, Thakur IS (2018) Pretreatment of Leucaena leucocephala wood by acidified glycerol: optimization, severity index and correlation analysis. Bioresour Technol 265:214–223. https://doi.org/10.1016/j.biortech.2018.05.084
Singhopon T, Shinoda K, Rujakom S, Kazama F (2021) Factors affecting the simultaneous removal of nitrate and reactive black 5 dye via hydrogen-based denitrification. Water (switzerland) 13(7):922. https://doi.org/10.3390/w13070922
Song Z, Song L, Shao Y, Tan L (2018) Degradation and detoxification of azo dyes by a salt-tolerant yeast Cyberlindnera samutprakarnensis S4 under high-salt conditions. World J Microbiol Biotechnol 34(9):1–13. https://doi.org/10.1007/s11274-018-2515-7
Srinivasan S, Sadasivam SK, Gunalan S, Shanmugam G, Kothandan G (2019) Application of docking and active site analysis for enzyme linked biodegradation of textile dyes. Environ Pollut 248:599–608. https://doi.org/10.1016/j.envpol.2019.02.080
Srivastava A, Dangi LK, Kumar S, Rani R (2022) Microbial decolorization of reactive black 5 dye by Bacillus albus DD1 isolated from textile water effluent: kinetic, thermodynamics & decolorization mechanism. Heliyon 8:e08834. https://doi.org/10.1016/j.heliyon.2022.e08834
Tan L, He M, Song L, Fu X, Shi S (2016) Aerobic decolorization, degradation and detoxification of azo dyes by a newly isolated salt-tolerant yeast Scheffersomyces spartinae TLHS-SF1. Bioresour Technol 203:287–294. https://doi.org/10.1016/j.biortech.2015.12.058
Vatandoostarani S, Bagheri Lotfabad T, Heidarinasab A, Yaghmaei S (2017) Degradation of azo dye methyl red by Saccharomyces cerevisiae ATCC 9763. Int Biodeterior Biodegrad 125:62–72. https://doi.org/10.1016/j.ibiod.2017.08.009
Acknowledgements
Authors are grateful to Tunisian Ministry of Higher Education, Scientific Research and Technology for supporting this research work.
Funding
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Author information
Authors and Affiliations
Contributions
ID: Investigation, writing-original draft, conceptualization, methodology. IBA and IL: Investigation (identification of strain). BH: Conceptualization, methodology, formal analysis, visualization. TM: Supervision.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest in the publication.
Rights and permissions
About this article
Cite this article
Dammak, I., Ben Atitallah, I., Louati, I. et al. Optimization of reactive black 5 decolorization by the newly isolated Saccharomyces cerevisiae X19G2 using response-surface methodology. 3 Biotech 12, 142 (2022). https://doi.org/10.1007/s13205-022-03191-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-022-03191-6