Abstract
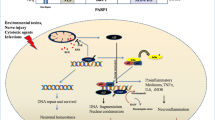
Poly (ADP-ribose) polymerase 1 (PARP1) protein is encoded by the PARP1 gene located on chromosome 1 (1q42.12) in human cells. It plays a crucial role in post-translational modification by adding poly (ADP-ribose) (PAR) groups to various proteins and PARP1 itself by utilizing nicotinamide adenine dinucleotide (NAD +) as a substrate. Since the discovery of PARP1, its role in DNA repair and cell death has been its identity. This is evident from an overwhelmingly high number of scientific reports in this regard. However, PARP1 also plays critical roles in inflammation, metabolism, tumor development and progression, chromatin modification and transcription, mRNA stability, and alternative splicing. In the present study, we attempted to compile all the scattered scientific information about this molecule, including the structure and multifunctional role of PARP1 in cancer and non-cancer diseases, along with PARP1 inhibitors (PARPis). Furthermore, for the first time, we have classified PARP1-mediated cell death for ease of understanding its role in cell death pathways.




Similar content being viewed by others
References
Ahmad SF, Zoheir KMA, Bakheet SA et al (2014) Poly(ADP-ribose) polymerase-1 inhibitor modulates T regulatory and IL-17 cells in the prevention of adjuvant induced arthritis in mice model. Cytokine 68:76–85. https://doi.org/10.1016/j.cyto.2014.04.006
Alemasova EE, Lavrik OI (2019) Poly(ADP-ribosyl)ation by PARP1: reaction mechanism and regulatory proteins. Nucleic Acids Res 47:3811–3827. https://doi.org/10.1093/nar/gkz120
Amé J-C, Spenlehauer C, de Murcia G (2004) The PARP superfamily. BioEssays 26:882–893. https://doi.org/10.1002/bies.20085
Andrabi SA, Kim NS, Yu S-W et al (2006) Poly(ADP-ribose) (PAR) polymer is a death signal. Proc Natl Acad Sci U S A 103:18308–18313. https://doi.org/10.1073/pnas.0606526103
Andrabi SA, Umanah GKE, Chang C et al (2014) Poly(ADP-ribose) polymerase-dependent energy depletion occurs through inhibition of glycolysis. Proc Natl Acad Sci U S A 111:10209–10214. https://doi.org/10.1073/pnas.1405158111
Arnold J, Grune T (2002) PARP-mediated proteasome activation: a co-ordination of DNA repair and protein degradation? BioEssays 24:1060–1065. https://doi.org/10.1002/bies.10179
Barboro P, Ferrari N, Capaia M et al (2015) Expression of nuclear matrix proteins binding matrix attachment regions in prostate cancer. PARP-1: new player in tumor progression. Int J Cancer 137:1574–1586. https://doi.org/10.1002/ijc.29531
Baumgartner M, Schneider R, Auer B et al (1992) Fluorescence in situ mapping of the human nuclear NAD+ ADP-ribosyltransferase gene (ADPRT) and two secondary sites to human chromosomal bands 1q42, 13q34, and 14q24. CGR 61:172–174. https://doi.org/10.1159/000133400
Bianchi AR, Ferreri C, Ruggiero S et al (2016) Automodification of PARP and fatty acid-based membrane lipidome as a promising integrated biomarker panel in molecular medicine. Biomark Med 10:229–242. https://doi.org/10.2217/bmm.16.3
Bieche I, Pennaneach V, Driouch K et al (2013) Variations in the mRNA expression of poly(ADP-ribose) polymerases, poly(ADP-ribose) glycohydrolase and ADP-ribosylhydrolase 3 in breast tumors and impact on clinical outcome. Int J Cancer 133:2791–2800. https://doi.org/10.1002/ijc.28304
Bouchard VJ, Rouleau M, Poirier GG (2003) PARP-1, a determinant of cell survival in response to DNA damage. Exp Hematol 31:446–454. https://doi.org/10.1016/s0301-472x(03)00083-3
Brighina L, Riva C, Bertola F et al (2011) Association analysis of PARP1 polymorphisms with Parkinson’s disease. Parkinsonism Relat Disord 17:701–704. https://doi.org/10.1016/j.parkreldis.2011.06.022
Bryant HE, Schultz N, Thomas HD et al (2005) Specific killing of BRCA2-deficient tumours with inhibitors of poly(ADP-ribose) polymerase. Nature 434:913–917. https://doi.org/10.1038/nature03443
Chaitanya GV, Alexander JS, Babu PP (2010) PARP-1 cleavage fragments: signatures of cell-death proteases in neurodegeneration. Cell Commun Signal 8:31. https://doi.org/10.1186/1478-811X-8-31
Choi J-R, Shin KS, Choi CY, Kang SJ (2016) PARP1 regulates the protein stability and proapoptotic function of HIPK2. Cell Death Dis 7:e2438–e2438. https://doi.org/10.1038/cddis.2016.345
Choi Y, Abdelmegeed MA, Song B-J (2017) Diet high in fructose promotes liver steatosis and hepatocyte apoptosis in C57BL/6J female mice: role of disturbed lipid homeostasis and increased oxidative stress. Food Chem Toxicol 103:111–121. https://doi.org/10.1016/j.fct.2017.02.039
Ciccarone F, Zampieri M, Caiafa P (2017) PARP1 orchestrates epigenetic events setting up chromatin domains. Semin Cell Dev Biol 63:123–134. https://doi.org/10.1016/j.semcdb.2016.11.010
D’Amours D, Desnoyers S, D’Silva I, Poirier GG (1999) Poly(ADP-ribosyl)ation reactions in the regulation of nuclear functions. Biochem J 342(Pt 2):249–268
Daar AS, Singer PA, Persad DL et al (2007) Grand challenges in chronic non-communicable diseases. Nature 450:494–496. https://doi.org/10.1038/450494a
de Murcia JM, Niedergang C, Trucco C et al (1997) Requirement of poly(ADP-ribose) polymerase in recovery from DNA damage in mice and in cells. Proc Natl Acad Sci U S A 94:7303–7307. https://doi.org/10.1073/pnas.94.14.7303
Dutta A, Eckelmann B, Adhikari S et al (2017) Microhomology-mediated end joining is activated in irradiated human cells due to phosphorylation-dependent formation of the XRCC1 repair complex. Nucleic Acids Res 45:2585–2599. https://doi.org/10.1093/nar/gkw1262
Eustermann S, Videler H, Yang J-C et al (2011) The DNA-binding domain of human PARP-1 interacts with DNA single-strand breaks as a monomer through its second zinc finger. J Mol Biol 407:149–170. https://doi.org/10.1016/j.jmb.2011.01.034
Farmer H, McCabe N, Lord CJ et al (2005) Targeting the DNA repair defect in BRCA mutant cells as a therapeutic strategy. Nature 434:917–921. https://doi.org/10.1038/nature03445
Fatokun AA, Dawson VL, Dawson TM (2014) Parthanatos: mitochondrial-linked mechanisms and therapeutic opportunities. Br J Pharmacol 171:2000–2016. https://doi.org/10.1111/bph.12416
Gagné J-P, Hendzel MJ, Droit A, Poirier GG (2006) The expanding role of poly(ADP-ribose) metabolism: current challenges and new perspectives. Curr Opin Cell Biol 18:145–151. https://doi.org/10.1016/j.ceb.2006.02.013
García S, Conde C (2015) The role of poly(ADP-ribose) polymerase-1 in rheumatoid arthritis. Mediat Inflamm 2015:e837250. https://doi.org/10.1155/2015/837250
Gariani K, Ryu D, Menzies KJ et al (2017) Inhibiting poly ADP-ribosylation increases fatty acid oxidation and protects against fatty liver disease. J Hepatol 66:132–141. https://doi.org/10.1016/j.jhep.2016.08.024
Gibson BA, Kraus WL (2012) New insights into the molecular and cellular functions of poly(ADP-ribose) and PARPs. Nat Rev Mol Cell Biol 13:411–424. https://doi.org/10.1038/nrm3376
Gobeil S, Boucher CC, Nadeau D, Poirier GG (2001) Characterization of the necrotic cleavage of poly(ADP-ribose) polymerase (PARP-1): implication of lysosomal proteases. Cell Death Differ 8:588–594. https://doi.org/10.1038/sj.cdd.4400851
Gross S, Kotova EYu, Maluchenko NV et al (2016) Evaluating Parp1 domains as gossypol targets. Moscow Univ BiolSci Bull 71:235–239. https://doi.org/10.3103/S0096392516040106
Haddad M, Rhinn H, Bloquel C et al (2006) Anti-inflammatory effects of PJ34, a poly(ADP-ribose) polymerase inhibitor, in transient focal cerebral ischemia in mice. Br J Pharmacol 149:23–30. https://doi.org/10.1038/sj.bjp.0706837
Hakmé A, Wong H-K, Dantzer F, Schreiber V (2008) The expanding field of poly(ADP-ribosyl)ation reactions. ‘protein modifications: beyond the usual suspects’ review series. EMBO Rep 9:1094–1100. https://doi.org/10.1038/embor.2008.191
Hans CP, Zerfaoui M, Naura AS et al (2009) Thieno[2,3-c]isoquinolin-5-one, a potent poly(ADP-ribose) polymerase inhibitor, promotes atherosclerotic plaque regression in high-fat diet-fed apolipoprotein E-deficient mice: effects on inflammatory markers and lipid content. J Pharmacol Exp Ther 329:150–158. https://doi.org/10.1124/jpet.108.145938
Hanzlikova H, Kalasova I, Demin AA et al (2018) The importance of poly(ADP-ribose) polymerase as a sensor of unligated Okazaki fragments during DNA replication. Mol Cell 71:319-331.e3. https://doi.org/10.1016/j.molcel.2018.06.004
Hassa PO, Hottiger MO (1999) A role of poly (ADP-ribose) polymerase in NF-kappaB transcriptional activation. Biol Chem 380:953–959. https://doi.org/10.1515/BC.1999.118
Hassa PO, Haenni SS, Buerki C et al (2005) Acetylation of poly(ADP-ribose) polymerase-1 by p300/CREB-binding protein regulates coactivation of NF-kappaB-dependent transcription. J Biol Chem 280:40450–40464. https://doi.org/10.1074/jbc.M507553200
Hegedűs C, Boros G, Fidrus E et al (2019) PARP1 inhibition augments UVB-mediated mitochondrial changes—implications for UV-induced DNA repair and photocarcinogenesis. Cancers (basel) 12:5. https://doi.org/10.3390/cancers12010005
Hochegger H, Dejsuphong D, Fukushima T et al (2006) Parp-1 protects homologous recombination from interference by Ku and Ligase IV in vertebrate cells. EMBO J 25:1305–1314. https://doi.org/10.1038/sj.emboj.7601015
Hoesel B, Schmid JA (2013) The complexity of NF-κB signaling in inflammation and cancer. Mol Cancer 12:86. https://doi.org/10.1186/1476-4598-12-86
Huang K, Du M, Tan X et al (2017) PARP1-mediated PPARα poly(ADP-ribosyl)ation suppresses fatty acid oxidation in non-alcoholic fatty liver disease. J Hepatol 66:962–977. https://doi.org/10.1016/j.jhep.2016.11.020
Huang A, Garraway LA, Ashworth A, Weber B (2020) Synthetic lethality as an engine for cancer drug target discovery. Nat Rev Drug Discov 19:23–38. https://doi.org/10.1038/s41573-019-0046-z
Jiang B-H, Tseng W-L, Li H-Y et al (2015) Poly(ADP-ribose) polymerase 1: cellular pluripotency, reprogramming, and tumorogenesis. Int J Mol Sci 16:15531–15545. https://doi.org/10.3390/ijms160715531
Kaelin WG (2005) The concept of synthetic lethality in the context of anticancer therapy. Nat Rev Cancer 5:689–698. https://doi.org/10.1038/nrc1691
Kam T-I, Mao X, Park H et al (2018) Poly(ADP-ribose) drives pathologic α-synuclein neurodegeneration in Parkinson’s disease. Science 362:eaat8407. https://doi.org/10.1126/science.aat8407
Ke Y, Han Y, Guo X et al (2017) PARP1 promotes gene expression at the post-transcriptiona level by modulating the RNA-binding protein HuR. Nat Commun 8:14632. https://doi.org/10.1038/ncomms14632
Ke Y, Wang C, Zhang J et al (2019) The role of PARPs in inflammation—and metabolic—related diseases: molecular mechanisms and beyond. Cells 8:1047. https://doi.org/10.3390/cells8091047
Kiss B, Szántó M, Szklenár M et al (2015) Poly(ADP) ribose polymerase-1 ablation alters eicosanoid and docosanoid signaling and metabolism in a murine model of contact hypersensitivity. Mol Med Rep 11:2861–2867. https://doi.org/10.3892/mmr.2014.3044
Ko HL, Ren EC (2012) Functional aspects of PARP1 in DNA repair and transcription. Biomolecules 2:524–548. https://doi.org/10.3390/biom2040524
Kossatz S, Brand C, Gutiontov S et al (2016a) Detection and delineation of oral cancer with a PARP1 targeted optical imaging agent. Sci Rep 6:21371. https://doi.org/10.1038/srep21371
Kossatz S, Weber WA, Reiner T (2016b) Optical imaging of PARP1 in response to radiation in oral squamous cell carcinoma. PLoS ONE 11:e0147752. https://doi.org/10.1371/journal.pone.0147752
Kossatz S, Pirovano G, França PDDS et al (2019) PARP1 as a biomarker for early detection and intraoperative tumor delineation in epithelial cancers – first-in-human results. BioRxiv. https://doi.org/10.1101/663385
Krishnakumar R, Kraus WL (2010) The PARP side of the nucleus: molecular actions, physiological outcomes, and clinical targets. Mol Cell 39:8–24. https://doi.org/10.1016/j.molcel.2010.06.017
Krishnakumar R, Gamble MJ, Frizzell KM et al (2008) Reciprocal binding of PARP-1 and histone H1 at promoters specifies transcriptional outcomes. Science 319:819–821. https://doi.org/10.1126/science.1149250
Krüger A, Bürkle A, Hauser K, Mangerich A (2020) Real-time monitoring of PARP1-dependent PARylation by ATR-FTIR spectroscopy. Nat Commun 11:2174. https://doi.org/10.1038/s41467-020-15858-w
Langelier M-F, Planck JL, Roy S, Pascal JM (2012) Structural basis for DNA damage-dependent poly(ADP-ribosyl)ation by human PARP-1. Science 336:728–732. https://doi.org/10.1126/science.1216338
Lindahl T, Satoh MS, Poirier GG, Klungland A (1995) Post-translational modification of poly(ADP-ribose) polymerase induced by DNA strand breaks. Trends Biochem Sci 20:405–411. https://doi.org/10.1016/s0968-0004(00)89089-1
Liu Y, Kadyrov FA, Modrich P (2011) PARP-1 enhances the mismatch-dependence of 5’-directed excision in human mismatch repair in vitro. DNA Repair (amst) 10:1145–1153. https://doi.org/10.1016/j.dnarep.2011.08.012
Liu Y, Zhang Y, Zhao Y et al (2016) High PARP-1 expression is associated with tumor invasion and poor prognosis in gastric cancer. Oncol Lett 12:3825–3835. https://doi.org/10.3892/ol.2016.5169
Luijsterburg MS, de Krijger I, Wiegant WW et al (2016) PARP1 links CHD2-mediated chromatin expansion and h3.3 deposition to DNA repair by non-homologous end-joining. Mol Cell 61:547–562. https://doi.org/10.1016/j.molcel.2016.01.019
Lüscher B, Bütepage M, Eckei L et al (2018) ADP-ribosylation, a multifaceted posttranslational modification involved in the control of cell physiology in health and disease. Chem Rev 118:1092–1136
Mangerich A, Bürkle A (2012) Pleiotropic cellular functions of PARP1 in longevity and aging: genome maintenance meets inflammation. Oxid Med Cell Longev 2012:321653. https://doi.org/10.1155/2012/321653
Mansour WY, Rhein T, Dahm-Daphi J (2010) The alternative end-joining pathway for repair of DNA double-strand breaks requires PARP1 but is not dependent upon microhomologies. Nucleic Acids Res 38:6065–6077. https://doi.org/10.1093/nar/gkq387
Mao K, Zhang G (2021) The role of PARP1 in neurodegenerative diseases and aging. FEBS J. https://doi.org/10.1111/febs.15716
Martinez-Zamudio R, Ha HC (2012) Histone ADP-ribosylation facilitates gene transcription by directly remodeling nucleosomes. Mol Cell Biol 32:2490–2502. https://doi.org/10.1128/MCB.06667-11
Martire S, Mosca L, d’Erme M (2015) PARP-1 involvement in neurodegeneration: a focus on Alzheimer’s and Parkinson’s diseases. Mech Ageing Dev 146–148:53–64. https://doi.org/10.1016/j.mad.2015.04.001
Mashimo M, Moss J (2016) Functional role of ADP-ribosyl-acceptor hydrolase 3 in poly(ADP-ribose) polymerase-1 response to oxidative stress. Curr Protein Pept Sci 17:633–640. https://doi.org/10.2174/1389203717666160419144603
Mashimo M, Kato J, Moss J (2013) ADP-ribosyl-acceptor hydrolase 3 regulates poly (ADP-ribose) degradation and cell death during oxidative stress. PNAS 110:18964–18969. https://doi.org/10.1073/pnas.1312783110
Mashimo M, Kato J, Moss J (2014) Structure and function of the ARH family of ADP-ribosyl-acceptor hydrolases. DNA Repair (amst) 23:88–94. https://doi.org/10.1016/j.dnarep.2014.03.005
Mashimo M, Onishi M, Uno A et al (2021) The 89-kDa PARP1 cleavage fragment serves as a cytoplasmic PAR carrier to induce AIF-mediated apoptosis. J Biol Chem 296:100046. https://doi.org/10.1074/jbc.RA120.014479
Matveeva E, Maiorano J, Zhang Q et al (2016) Involvement of PARP1 in the regulation of alternative splicing. Cell Discov 2:15046. https://doi.org/10.1038/celldisc.2015.46
Matveeva EA, Al-Tinawi QMH, Rouchka EC, Fondufe-Mittendorf YN (2019a) Coupling of PARP1-mediated chromatin structural changes to transcriptional RNA polymerase II elongation and cotranscriptional splicing. Epigenetics Chromatin 12:15. https://doi.org/10.1186/s13072-019-0261-1
Matveeva EA, Mathbout LF, Fondufe-Mittendorf YN (2019b) PARP1 is a versatile factor in the regulation of mRNA stability and decay. Sci Rep 9:3722. https://doi.org/10.1038/s41598-019-39969-7
Mazzon E, Dugo L, Li J-H et al (2002) GPI 6150, a PARP inhibitor, reduces the colon injury caused by dinitrobenzene sulfonic acid in the rat. Biochem Pharmacol 64:327–337. https://doi.org/10.1016/s0006-2952(02)01075-4
Meder VS, Boeglin M, de Murcia G, Schreiber V (2005) PARP-1 and PARP-2 interact with nucleophosmin/B23 and accumulate in transcriptionally active nucleoli. J Cell Sci 118:211–222. https://doi.org/10.1242/jcs.01606
Melikishvili M, Chariker JH, Rouchka EC, Fondufe-Mittendorf YN (2017) Transcriptome-wide identification of the RNA-binding landscape of the chromatin-associated protein PARP1 reveals functions in RNA biogenesis. Cell Discov 3:17043. https://doi.org/10.1038/celldisc.2017.43
Mohammad G, Siddiquei MM, Abu El-Asrar AM (2013) Poly (ADP-ribose) polymerase mediates diabetes-induced retinal neuropathy. Mediat Inflamm 2013:e510451. https://doi.org/10.1155/2013/510451
Morales JC, Li L, Fattah FJ et al (2014) Review of poly (ADP-ribose) polymerase (PARP) mechanisms of action and rationale for targeting in cancer and other diseases. Crit Rev Eukaryot Gene Expr 24:15–28
Mukhopadhyay P, Rajesh M, Cao Z et al (2014) Poly (ADP-ribose) polymerase-1 is a key mediator of liver inflammation and fibrosis. Hepatology 59:1998–2009. https://doi.org/10.1002/hep.26763
Mukhopadhyay P, Horváth B, Rajesh M et al (2017) PARP inhibition protects against alcoholic and non-alcoholic steatohepatitis. J Hepatol 66:589–600. https://doi.org/10.1016/j.jhep.2016.10.023
Murai J, Zhang Y, Morris J et al (2014) Rationale for poly(ADP-ribose) polymerase (PARP) inhibitors in combination therapy with camptothecins or temozolomide based on PARP trapping versus catalytic inhibition. J Pharmacol Exp Ther 349:408–416. https://doi.org/10.1124/jpet.113.210146
Nijman SMB (2011) Synthetic lethality: general principles, utility and detection using genetic screens in human cells. FEBS Lett 585:1–6. https://doi.org/10.1016/j.febslet.2010.11.024
Nomura F, Yaguchi M, Togawa A et al (2000) Enhancement of poly-adenosine diphosphate-ribosylation in human hepatocellular carcinoma. J Gastroenterol Hepatol 15:529–535. https://doi.org/10.1046/j.1440-1746.2000.02193.x
O’Donnell A, Yang S-H, Sharrocks AD (2013) PARP1 orchestrates variant histone exchange in signal-mediated transcriptional activation. EMBO Rep 14:1084–1091. https://doi.org/10.1038/embor.2013.164
Obrosova IG, Li F, Abatan OI et al (2004) Role of poly(ADP-ribose) polymerase activation in diabetic neuropathy. Diabetes 53:711–720. https://doi.org/10.2337/diabetes.53.3.711
Oliver F, Murcia J, Nacci C et al (1999) Resistance to endotoxic shock as a consequence of defective NF-kappaB activation in poly (ADP-ribose) polymerase-1 deficient mice. EMBO J 18: 4446–4454. EMBO J 18:4446–4454. https://doi.org/10.1093/emboj/18.16.4446
Ossovskaya V, Koo IC, Kaldjian EP et al (2010) Upregulation of poly (ADP-ribose) polymerase-1 (PARP1) in triple-negative breast cancer and other primary human tumor types. Genes Cancer 1:812–821. https://doi.org/10.1177/1947601910383418
Pascual M, López-Nevot MA, Cáliz R et al (2003) A poly(ADP-ribose) polymerase haplotype spanning the promoter region confers susceptibility to rheumatoid arthritis. Arthritis Rheum 48:638–641. https://doi.org/10.1002/art.10864
Pazzaglia S, Pioli C (2019) Multifaceted role of PARP-1 in DNA repair and inflammation: pathological and therapeutic implications in cancer and non-cancer diseases. Cells 9:41. https://doi.org/10.3390/cells9010041
Pletcher JP, Bhattacharjee S, Doan JP et al (2021) The emerging role of poly (ADP-ribose) polymerase inhibitors as effective therapeutic agents in renal cell carcinoma. Front Oncol 11:681441. https://doi.org/10.3389/fonc.2021.681441
Puthanveetil P, Zhang D, Wang Y et al (2012) Diabetes triggers a PARP1 mediated death pathway in the heart through participation of FoxO1. J Mol Cell Cardiol 53:677–686. https://doi.org/10.1016/j.yjmcc.2012.08.013
Quénet D, El Ramy R, Schreiber V, Dantzer F (2009) The role of poly(ADP-ribosyl)ation in epigenetic events. Int J Biochem Cell Biol 41:60–65. https://doi.org/10.1016/j.biocel.2008.07.023
Racz B, Hanto K, Tapodi A et al (2010) Regulation of MKP-1 expression and MAPK activation by PARP-1 in oxidative stress: a new mechanism for the cytoplasmic effect of PARP-1 activation. Free Radic Biol Med 49:1978–1988. https://doi.org/10.1016/j.freeradbiomed.2010.09.026
Ramazi S, Zahiri J (2021) Post-translational modifications in proteins: resources, tools and prediction methods. Database 2021. https://doi.org/10.1093/database/baab012
Rancourt A, Satoh MS (2009) Delocalization of nucleolar poly(ADP-ribose) polymerase-1 to the nucleoplasm and its novel link to cellular sensitivity to DNA damage. DNA Repair (amst) 8:286–297. https://doi.org/10.1016/j.dnarep.2008.11.018
Ray Chaudhuri A, Nussenzweig A (2017) The multifaceted roles of PARP1 in DNA repair and chromatin remodelling. Nat Rev Mol Cell Biol 18:610–621. https://doi.org/10.1038/nrm.2017.53
Robu M, Shah RG, Petitclerc N et al (2013) Role of poly(ADP-ribose) polymerase-1 in the removal of UV-induced DNA lesions by nucleotide excision repair. Proc Natl Acad Sci U S A 110:1658–1663. https://doi.org/10.1073/pnas.1209507110
Ronson GE, Piberger AL, Higgs MR et al (2018) PARP1 and PARP2 stabilise replication forks at base excision repair intermediates through Fbh1-dependent Rad51 regulation. Nat Commun 9:746. https://doi.org/10.1038/s41467-018-03159-2
Rouleau M, Patel A, Hendzel MJ et al (2010) PARP inhibition: PARP1 and beyond. Nat Rev Cancer 10:293–301. https://doi.org/10.1038/nrc2812
Salemi M, Mazzetti S, De Leonardis M et al (2021) Poly (ADP-ribose) polymerase 1 and Parkinson’s disease: a study in post-mortem human brain. Neurochem Int 144:104978. https://doi.org/10.1016/j.neuint.2021.104978
Sánchez-Fidalgo S, Villegas I, Martín A et al (2007) PARP inhibition reduces acute colonic inflammation in rats. Eur J Pharmacol 563:216–223. https://doi.org/10.1016/j.ejphar.2007.01.070
Santini D, Perrone G, Roato I et al (2011) Expression pattern of receptor activator of NFκB (RANK) in a series of primary solid tumors and related bone metastases. J Cell Physiol 226:780–784. https://doi.org/10.1002/jcp.22402
Satoh MS, Poirier GG, Lindahl T (1994) Dual function for poly(ADP-ribose) synthesis in response to DNA strand breakage. Biochemistry 33:7099–7106. https://doi.org/10.1021/bi00189a012
Schreiber V, Dantzer F, Ame J-C, de Murcia G (2006) Poly(ADP-ribose): novel functions for an old molecule. Nat Rev Mol Cell Biol 7:517–528. https://doi.org/10.1038/nrm1963
Scully R, Chen J, Plug A et al (1997) Association of BRCA1 with Rad51 in mitotic and meiotic cells. Cell 88:265–275. https://doi.org/10.1016/s0092-8674(00)81847-4
Sethi GS, Dharwal V, Naura AS (2017) Poly(ADP-ribose)polymerase-1 in lung inflammatory disorders: a review. Front Immunol. https://doi.org/10.3389/fimmu.2017.01172
Sharan SK, Morimatsu M, Albrecht U et al (1997) Embryonic lethality and radiation hypersensitivity mediated by Rad51 in mice lacking Brca2. Nature 386:804–810. https://doi.org/10.1038/386804a0
Shin H-J, Kwon H-K, Lee J-H et al (2015) Doxorubicin-induced necrosis is mediated by poly-(ADP-ribose) polymerase 1 (PARP1) but is independent of p53. Sci Rep 5:15798. https://doi.org/10.1038/srep15798
Singh N (1991) Enhanced poly ADP-ribosylation in human leukemia lymphocytes and ovarian cancers. Cancer Lett 58:131–135. https://doi.org/10.1016/0304-3835(91)90035-g
Smulson ME, Kang VH, Ntambi JM et al (1995) Requirement for the expression of poly(ADP-ribose) polymerase during the early stages of differentiation of 3T3-L1 preadipocytes, as studied by antisense RNA induction. J Biol Chem 270:119–127. https://doi.org/10.1074/jbc.270.1.119
Soldani C, Scovassi AI (2002) Poly(ADP-ribose) polymerase-1 cleavage during apoptosis: an update. Apoptosis 7:321–328. https://doi.org/10.1023/A:1016119328968
Soldani C, Lazzè MC, Bottone MG et al (2001) Poly(ADP-ribose) polymerase cleavage during apoptosis: when and where? Exp Cell Res 269:193–201. https://doi.org/10.1006/excr.2001.5293
Swindall AF, Stanley JA, Yang ES (2013) PARP-1: friend or foe of DNA damage and repair in tumorigenesis? Cancers (basel) 5:943–958. https://doi.org/10.3390/cancers5030943
Szabó É, Kovács I, Grune T et al (2011) PARP-1: a new player in the asthma field? Allergy 66:811–814. https://doi.org/10.1111/j.1398-9995.2011.02551.x
Szántó M, Gupte R, Kraus WL et al (2021) PARPs in lipid metabolism and related diseases. Prog Lipid Res. https://doi.org/10.1016/j.plipres.2021.101117
Thapa K, Khan H, Sharma U et al (2021) Poly (ADP-ribose) polymerase-1 as a promising drug target for neurodegenerative diseases. Life Sci 267:118975. https://doi.org/10.1016/j.lfs.2020.118975
Thomas C, Ji Y, Wu C et al (2019) Hit and run versus long-term activation of PARP-1 by its different domains fine-tunes nuclear processes. PNAS 116:9941–9946
Tomoda T, Kurashige T, Moriki T et al (1991) Enhanced expression of poly(ADP-ribose) synthetase gene in malignant lymphoma. Am J Hematol 37:223–227. https://doi.org/10.1002/ajh.2830370402
Ummarino S, Hausman C, Di Ruscio A (2021) The PARP way to epigenetic changes. Genes (basel) 12:446. https://doi.org/10.3390/genes12030446
Valabrega G, Scotto G, Tuninetti V et al (2021) Differences in PARP inhibitors for the treatment of ovarian cancer: mechanisms of action, pharmacology, safety, and efficacy. Int J Mol Sci 22:4203. https://doi.org/10.3390/ijms22084203
Wang Y, An R, Umanah GK et al (2016) A nuclease that mediates cell death induced by DNA damage and poly(ADP-ribose) polymerase-1. Science 354:aad6872. https://doi.org/10.1126/science.aad6872
Wang C, Xu W, Zhang Y et al (2018) PARP1 promote autophagy in cardiomyocytes via modulating FoxO3a transcription. Cell Death Dis 9:1–15. https://doi.org/10.1038/s41419-018-1108-6
Wang H, Kuusela S, Rinnankoski-Tuikka R et al (2020) Tankyrase inhibition ameliorates lipid disorder via suppression of PGC-1α PARylation in db/db mice. Int J Obes (lond) 44:1691–1702. https://doi.org/10.1038/s41366-020-0573-z
Weaver AN, Yang ES (2013) Beyond DNA repair: additional functions of PARP-1 in cancer. Front Oncol 3:290. https://doi.org/10.3389/fonc.2013.00290
Wei H, Yu X (2016) Functions of PARylation in DNA damage repair pathways. Genomics Proteomics Bioinform 14:131–139. https://doi.org/10.1016/j.gpb.2016.05.001
Westera L, Jennings AM, Maamary J et al (2019) Poly-ADP ribosyl polymerase 1 (PARP1) regulates influenza A virus polymerase. Adv Virol 2019:e8512363. https://doi.org/10.1155/2019/8512363
Wu X, Dong Z, Wang CJ et al (2016) FASN regulates cellular response to genotoxic treatments by increasing PARP-1 expression and DNA repair activity via NF-κB and SP1. Proc Natl Acad Sci U S A 113:E6965–E6973. https://doi.org/10.1073/pnas.1609934113
Xu W, Hu X, Anwaier A et al (2020a) Fatty acid synthase correlates with prognosis-related abdominal adipose distribution and metabolic disorders of clear cell renal cell carcinoma. Front Mol Biosci 7:610229. https://doi.org/10.3389/fmolb.2020.610229
Xu W-H, Xu Y, Tian X et al (2020b) Large-scale transcriptome profiles reveal robust 20-signatures metabolic prediction models and novel role of G6PC in clear cell renal cell carcinoma. J Cell Mol Med 24:9012–9027. https://doi.org/10.1111/jcmm.15536
Yalcintepe L, Turker-Sener L, Sener A et al (2005) Changes in NAD/ADP-ribose metabolism in rectal cancer. Braz J Med Biol Res 38:361–365. https://doi.org/10.1590/s0100-879x2005000300006
Yang L, Huang K, Li X et al (2013) Identification of poly(ADP-ribose) polymerase-1 as a cell cycle regulator through modulating sp1 mediated transcription in human hepatoma cells. PLoS ONE 8:e82872. https://doi.org/10.1371/journal.pone.0082872
Yu S-W, Andrabi SA, Wang H et al (2006) Apoptosis-inducing factor mediates poly(ADP-ribose) (PAR) polymer-induced cell death. PNAS 103:18314–18319. https://doi.org/10.1073/pnas.0606528103
Zaffini R, Gotte G, Menegazzi M (2018) Asthma and poly(ADP-ribose) polymerase inhibition: a new therapeutic approach. Drug Des Devel Ther 12:281–293. https://doi.org/10.2147/DDDT.S150846
Zhou P, Wang J, Mishail D, Wang C-Y (2020) Recent advancements in PARP inhibitors-based targeted cancer therapy. Precis Clin Med 3:187–201. https://doi.org/10.1093/pcmedi/pbaa030
Zhou Y, Liu L, Tao S et al (2021) Parthanatos and its associated components: promising therapeutic targets for cancer. Pharmacol Res 163:105299. https://doi.org/10.1016/j.phrs.2020.105299
Zingarelli B, Hake PW, Burroughs TJ et al (2004) Activator protein-1 signalling pathway and apoptosis are modulated by poly(ADP-ribose) polymerase-1 in experimental colitis. Immunology 113:509–517. https://doi.org/10.1111/j.1365-2567.2004.01991.x
Acknowledgements
Vikas Kumar is a recipient of a senior research fellowship from the Indian Council of Medical Research, New Delhi, India.
Funding
Indian Council of Medical Research (ICMR), Government of India Grant no. BMS/7/2020-21, New Delhi, India financially supported this study.
Author information
Authors and Affiliations
Contributions
VK and AK conceptualized and drafted the manuscript. KIM assisted in reviewing the literature. VY helped prepare the manuscript. SSC obtained funding and critically revised the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest to declare.
Rights and permissions
About this article
Cite this article
Kumar, V., Kumar, A., Mir, K.U.I. et al. Pleiotropic role of PARP1: an overview. 3 Biotech 12, 3 (2022). https://doi.org/10.1007/s13205-021-03038-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-021-03038-6