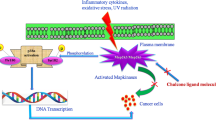
Abstract
In the present study, we have focused on to elucidate potential bioactive pyrrolopyridine (PYP23) analogs against human mitogen-activated protein kinase-activated protein kinase-2 (MK-2). Here, in silico methods and computational systems biology tools were used as rational strategies to predict novel PYP23 analogs against the MK-2. Initially, crystal structure (PDB-ID: 2P3G) consists steriochemical conflicts were rectified by structure-optimization approaches using the Modeller program, and a new optimized-high resolution model was generated. The stereochemical qualities of the predicted MK-2 model were judged; these showed that the model was reliable for docking assessments. SAR-based bioactivity analysis showed that among the 197 datasets only 15 candidates contained bioactivity data and were accepted as probable MK-2 inhibitors. Virtual screening and docking strategies of dataset compounds against the ligand-binding domain of MK-2 recognized 13 composites containing high binding affinity than known compounds. Furthermore, the comparative structure clustering, in silico toxicogenomics and QSAR-based anticancer properties prediction approaches were successful in the recognition of five best potential compounds such as 60118340, 60118338, 60117736, 60118473 and 60118322, which have great anticancer and drug-likeness with non-toxicity class indices. Leu70, Glu139, Leu141, Glu145, Glu190, Thr206 and Asp207 were found to be novel hotspot residues prominently involved in H-bonds framing with ligands. Interestingly, they have shown better molecular similarity with known bioactive PYP inhibitors. Thus, predicted five compounds can useful as possible chemotherapeutic agents for MK-2 and show similar molecular actions like known PYP inhibitors. Overall, these streamlined new methods may have great potential to reveal possible ligands toward other molecular targets and biomarkers.





Similar content being viewed by others
References
Anderson DR, Meyers MJ, Vernier WF et al (2007) Pyrrolopyridine Inhibitors of mitogen-activated protein kinase-activated protein kinase 2 (MK-2). J Med Chem 50:2647–2654
Anton R, Bauer SM, Peter RWEF, Keck et al (2014) A p38 substrate-specific MK2-EGFP translocation assay for identification and validation of new p38 inhibitors in living cells: a comprising alternative for acquisition of cellular p38 inhibition Data. PLoS One. https://doi.org/10.1371/journal.pone.0095641
Bommu UD, Konidala KK, Pabbaraju N, Yeguvapalli S (2017) Ligand-based virtual screening, molecular docking, QSAR and pharmacophore analysis of quercetin-associated potential novel analogs against epidermal growth factor receptor. J Recept Signal Transduct Res 37:600–610
Cargnello M, Roux PP (2011) Activation and function of the MAPKs and their substrates, the MAPK-activated protein kinases. Microbiol Mol Biol Rev 75:50–83
Chen K, Huang J, Gong W et al (2007) Toll-like receptors in inflammation, infection and cancer. Int Immunopharmacol 7:1271–1285
Cheng F, Li W, Zhou Y et al (2012) admetSAR: a comprehensive source and free tool for assessment of chemical ADMET properties. J Chem Inf Model 52:3099–3105
Chikanza IC (2004) Corticosteroid resistance in rheumatoid arthritis: molecular and cellular perspectives. Rheumatology 43:1337–1345
Colovos C, Yeates TO (1993) Verification of protein structures: patterns of nonbonded atomic interactions. Protein Sci 2:1511–1519
Daina A, Michielin O, Zoete V (2017) SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci Rep 7:42717
Dallakyan S, Olson AJ (2014) Small-molecule library screening by docking with PyRx. Methods Mol Biol 1263:243–250
Dobbelstein M, Moll U (2014) Targeting tumour-supportive cellular machineries in anticancer drug development. Nat Rev Drug Discov 13:179–196
Fiore M, Forli S, Manetti F (2015) Targeting mitogen-activated protein kinase-activated protein kinase 2 (MAPKAPK2, MK2): medicinal chemistry efforts to lead small molecule inhibitors to clinical trials. J Med Chem 59:3609–3634
Gurgis FMS, Ziaziaris W, Munoz L (2013) Mitogen-activated protein kinase-activated protein kinase 2 in neuroinflammation, heat shock protein 27 phosphorylation, and cell cycle: role and targeting. Mol Pharmacol 85(2):345–356
Höpker K, Hagmann H, Khurshid S et al (2012) AATF/Che-1 acts as a phosphorylation-dependent molecular modulator to repress p53-driven apoptosis. EMBO J 31:3961–3975
Kaplan W (2001) Swiss-PDB viewer (deep view). Brief Bioinform 2:195–197
Kim S, Han L, Yu B et al (2015a) PubChem structure–activity relationship (SAR) clusters. J Chem Inform. https://doi.org/10.1186/s13321-015-0070-x
Kim S, Thiessen PA, Bolton EE et al (2015b) PubChem substance and compound databases. Nucleic Acids Res. https://doi.org/10.1093/nar/gkv951
Kumar KK, Devi BU, Neeraja P (2017) Integration of in silico approaches to determination of endocrine-disrupting perfluorinated chemicals binding potency with steroidogenic acute regulatory protein. Biochem Biophys Res Commun 491:1007–1014
Laskowski RA, Macarthur MW, Moss DS, Thornton JM (1993) PROCHECK: a program to check the stereochemical quality of protein structures. J Appl Crystallogr 26:283–291
Leelananda SP, Lindert S (2016) Computational methods in drug discovery. Beilstein J Org Chem 12:2694–2718
Lüthy R, Bowie JU, Eisenberg D (1992) Assessment of protein models with three-dimensional profiles. Nature 356:83–85
Manke IA, Nguyen A, Lim D et al (2005) MAPKAP kinase-2 is a cell cycle checkpoint kinase that regulates the G2/M transition and S phase progression in response to UV irradiation. Mol Cell 17:37–48
Mourey RJ, Burnette BL, Brustkern SJ et al (2010) A benzothiophene inhibitor of mitogen-activated protein kinase-activated protein kinase 2 inhibits tumor necrosis factor production and has oral anti-inflammatory efficacy in acute and chronic models of inflammation. J Pharmacol Exp Ther 333:797–807
Phatak SS, Stephan CC, Cavasotto CN (2009) High-throughput and in silico screenings in drug discovery. Expert Opin Drug Discov 4:947–959
Putz M, Duda-Seiman C, Duda-Seiman D et al (2016) Chemical structure-biological activity models for pharmacophores’ 3D-interactions. Int J Mol Sci 17:1087
Šali A, Blundell TL (1993) Comparative protein modelling by satisfaction of spatial restraints. J Mol Biol 234:779–815
Sancar A, Lindsey-Boltz LA, Gaddameedhi S et al (2014) Circadian clock, cancer, and chemotherapy. Biochemistry 54:110–123
Singh H, Kumar R, Singh S et al (2016) Prediction of anticancer molecules using hybrid model developed on molecules screened against NCI-60 cancer cell lines. BMC Cancer. https://doi.org/10.1186/s12885-016-2082-y
Sliwoski G, Kothiwale S, Meiler J, Lowe EW (2013) Computational methods in drug discovery. Pharmacol Rev 66:334–395
Trempolec N, Dave-Coll N, Nebreda AR (2013) SnapShot: p38 MAPK substrates. Cell. https://doi.org/10.1016/j.cell.2013.01.047
Tusar M, Minovski N, Fjodorova N, Novic M (2012) In silico assessment of adverse effects of a large set of 6-fluoroquinolones obtained from a study of tuberculosis chemotherapy. Curr Drug Saf 7:313–320
Xu L, Chen S, Bergan RC (2006) MAPKAPK2 and HSP27 are downstream effectors of p38 MAP kinase-mediated matrix metalloproteinase type 2 activation and cell invasion in human prostate cancer. Oncogene 25:2987–2998
Acknowledgements
K. Kranthi Kumar (F1-17.1/2013-14/RGNF-2013-14-SC-AND-40670/(SAIII/Website)) and B. Uma Devi (F1–17.1/2012–13/RGNF-2012–13-SC-AND-32761/(SA-III/WEBSITE)) are thankful to University Grants Commission (UGC) New Delhi, India, for awarding research fellowship, Rajiv Gandhi National Fellowship (RGNF/SRF), to carry out this work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Konidala, K.K., Bommu, U.D., Yeguvapalli, S. et al. In silico insights into prediction and analysis of potential novel pyrrolopyridine analogs against human MAPKAPK-2: a new SAR-based hierarchical clustering approach. 3 Biotech 8, 385 (2018). https://doi.org/10.1007/s13205-018-1405-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-018-1405-x