Abstract
Self-assembly of biomolecules has gained increasing attention as it generates various supramolecular structural assemblies having potential applications principally in biomedical sciences. Here, we show that amino acid like tryptophan or tyrosine readily aggregates as nanotubes via a simple self-assembly process. These were characterized by FTIR, scanning electron microscopy, and by fluorescence microscopy. Nanotubes prepared from tryptophan are having ~200 nm inner diameter and those from tyrosine are having the same around ~50 nm diameter.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The molecular aggregation of amino acids in creating all the essential proteins are dictated by DNA using basic coded assemblies under RNA scaffold. Nevertheless, while continuing such faultless assemblies, on aging, sometimes the damaged DNA and also the RNA result in the production of non-functional protein aggregates. Such age-related or environmentally damaged proteins normally introduce diseases with abnormal function by themselves and also influencing normal proteins by their antagonistic interaction. As specific shape is very much related to selective functioning of a complex biomolecule, the possibility of self-assembly of amino acids even under non-coded form is worth exploring. This could be related to exploring the bio world in relevance to its benign utilization as carriers for drug and for fluorescent species to be utilized for bio-imaging. Furthermore, such possible assemblies may address the role of amyloidal proteins which invite various diseases, disrupting normal physiological functions leading to disrupt the healthy function of tissues and organs (Pulawski et al. 2012). General investigation where chances to explore various nano-shaped aggregates prepared by self-assembly of simple building blocks have attracted immense attention in recent years. Such aggregation have been shown to be extremely useful in the domain of nanoscience and nanotechnology because of their uses in casting metal nanowires (Carny et al. 2006; Reches and Gazit 2003), vessels for nanomaterials (Shimizu 2006; Zhao et al. 2006), nanofluidic devices (Sott et al. 2006) and for drug delivery (Masaru et al. 2011; Raviv et al. 2005). The various forces that control the molecular assembly or organization of biomolecules in forming nanoshapes are the hydrogen-bonding interactions, van der Waals attractions, steric repulsions, and capillary forces during solvent evaporation. In 1993, Ghadiri et al. (1993) prepared organic nanotubes from cyclic D,L-peptide rings. Other routes involved cyclic peptides of all β-amino acids for the formation of nanotubes (Clark et al. 1998; Seebach et al. 1997). Serine-based aromatic cyclodepsipeptide has been transformed into crystalline cylindrical assemblies (Ranganathan et al. 1998), and to cyclodextrin (Harada et al. 1993) or to cyclic d,l-rhamnopyranose (Ashton et al. 1996). Reches and Gazit (2003) showed a simple self-assembled system for the preparation of fiber structure using a dipeptide (phenylalanine–phenylalanine, FF). Ryan et al. (2010) detailed the using of an FMOC-protected simpler biomolecule phenylalanine (Fmoc-F5-Phe) to form hollow tubular structures. In 2012, scientist used unprotected phenylaniline-made fibrous structures (Adler-Abramovich et al. 2012). So far, only derivatized amino acids were used to synthesize nanostructured materials especially nanotubes via self-assembly through peptide formation. Perween et al. (2013) used single amino acid (phenylalanine, tyrosine and glycine) for the preparation of febrile structure under neutral, aqueous conditions. The nature of such interactions was simply dominated by non-covalent π–π interactions. A stable structure in nanodomain using single amino acid is rare. Therefore, in this communication, we describe the synthesis of stable nanotube type aggregates from individual amino acids like tryptophan or tyrosine retaining characteristic fluorescence.
Experimental
Materials
All chemicals used in the synthesis were purchased from Sigma-Aldrich and were used without any further purification.
Synthesis of nanotubes from tryptophan and tyrosine
In a typical sample preparation for nanotubes of tryptophan or tyrosine, a stock solution of each was prepared in ethanol with concentration of 50 mg/mL. Each stock solution was used fresh to avoid undesirable reactions in the aging process. The stock solution was diluted to the final concentration of 0.5 mg/mL before use. The solution was sonicated for 10 min and 5 μL of such a solution was deposited onto brass stub in each case separately. The samples were dried for 3 h under a table lamp (60 W tungsten) and then subjected to field emission scanning electron microscopic (FESEM) analysis. Sequential gold sputtering was done before imaging.
Macroscopic and spectroscopic characterization
Infrared spectroscopy
The infrared spectroscopic measurements of nanotubes were recorded on a Bruker Vertex 70, FT-IR spectrophotometer in KBr phase.
Field emission scanning electron microscopy (FESEM)
SUPRA 40VP field emission scanning electron microscope (Carl ZeissNTS GmbH, Oberkochen, Germany) equipped with energy-dispersive X-ray (EDAX), in high vacuum mode at 10 kV, was used for the visualization of nanostructures of amino acids tryptophan and tyrosine.
Fluorescence microscopy
The fluorescence images of nanotubes of tryptophan and tyrosine were performed on a Leica inverted microscope (Leica DC200, Leica microscopy system Ltd., CH-9435, Heerbrugg) equipped with an RS PhotometricsSensys camera, KAF1401E G1.
Results and discussion
Amino acid nanotubes
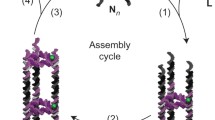
Molecular interactions play very important role in self-assembly to form aggregate nanotubular structures. The polymer assembly process could be visualized as shown in Scheme 1. This is assumed on the basis of previous reports (Ghadiri et al. 1993; Khazanovich et al. 1994). Here, once stacking took place to get a flat structure then the drive for minimum surface area spontaneously made it to a cylindrical shape. The formation of self-assembled structures, here, are favored by aromatic ring stacking (π–π) of amino acids as envisioned earlier with aromatic functionalities (Mishra and Chauhan 2011; Pandit et al. 2008). Along with such interaction, there are other possible interactions like van der Walls and hydrogen bonding like C–H–π as discussed earlier (Meital and Ehud 2006; Mishra and Chauhan 2011; Parween et al. 2014; Reches and Gazit 2006). In addition, hydrogen bonding of other type may occur via nitrogen of amine of tryptophan to the carboxylic hydrogen end of the adjoining molecule of tryptophan. Similar interactions may be invoked for the other amino acid like tyrosine. In tyrosine, the presence of –OH group may also participate in the formation hydrogen bonding. They may also involve the backbone-backbone-type, intermolecular hydrogen-bonding interactions. These interactions may be parallel or antiparallel (Granja and Ghadiri 1994; Hartgerink et al. 1996; Kobayashi et al. 1995). In addition, these molecules are self-assembled using co-operative force of water molecules present in the solvent like ethanol and those were used in the assembly utilizing hydrogen bonding. The role of water molecule in such self-assembly of nanotubes is reported (Fujibayashi et al. 2008; Begum et al. 2014). Water also plays important role in the stacking process. It interacts with formed nanotubes wall and also with the other water molecules with the help of hydrogen bonding that may lead to the stabilization of achieved structure (Andrade-Filho et al. 2016). By changing the concentration of water, Kim et al. (2010) showed such a variation may dictate the change in the shape of the final product from nanotube to nanowire. The self-assembly of peptides to nanotubes are fashioned via a network of hydrogen bonds between the backbone of the peptide and water molecules, which can enter the channels of polypeptide chain due to its hydrophilic nature (Mao et al. 2000). The use of amino acid side chain and charged termini also may play crucial role in initiating self-assembled processes. Therefore, in the present case stacking interaction followed by self-assembly readily drive to attain minimum energy on the surface leading to tube form of the amino acids under nanodomain (Scheme 1).
FTIR study
The FTIR spectra of the tryptophan and nanotubes of tryptophan are shown in Fig. 1. The black lined corresponds to the starting material tryptophan and red lined spectrum shown in Fig. 1, corresponds to the nanotubes. There is no makeable difference in both the spectra displaying that the integrity of the amino acid is retained. It is only to be noted that in nanotube the intensity of a particular vibration is damped and that could be due to the movement of big mass associated with the aggregated assembly in nanotube structure The peak at 1668 cm−1 corresponds to the C=O stretching of the tryptophan (Tsuguo 1972) and the deformation vibrations lie in between 700 and 500 cm−1 (Carubelli et al. 1997). The peak at 1586 cm−1 is due to C=C stretching in the aromatic ring. The peak around 1358 cm−1 corresponds to the stretching vibration of C=C in indole ring (Cao and Fischer 1999). The peaks in between 1230 cm−1 and 1000 cm−1 are due to the in-plane deformation of C–H in indole ring (Ma et al. 2009). The intense peak at around 3399 cm−1 corresponds to the NH stretching vibrations of indole ring (Ivanova 2006). As per the present discussion, most of these vibrations of monomeric tryptophan get broadened with the loss in intensity. The vibrations around 3041 cm−1 which is related to aromatic CH vibrations of tryptophan responds to further splitting indicative of (π–π) and C–H–π interactions in tryptophan nanotube. The drastic change in the shape of deformation vibrations around 700–500 cm−1 supports such interactions.
Similarly, Fig. 2 shows the FTIR spectrum-free tyrosine (black line) and tyrosine nanotubes as shown by red line. Again, we observed that both the spectra are almost identical with respect to their relative peak intensities. The peak at 3205 cm−1 corresponds to the OH stretching vibration. Similarly, the broad absorption centered at 2965 cm−1 is the composite vibrations arising of ν(NH), ν(CH) stretching. In the nanoaggregate of tyrosine, these peaks retained the profile but some distinctive changes occur in the ring deformation region around 840–500 cm−1 region suggesting similar π–π and C–H–π type interactions. The peak at 1609 cm−1 belongs to to C=O stretching frequency and the peak at 1592 cm−1 corresponds to the C=C stretching in the aromatic ring (Arp et al. 2001; Encinar et al. 2002; Reid et al. 2003). Therefore, to attain a new complex shape these molecules almost retain their individual IR signature.
FESEM study
The spontaneous self-assembly of amino acids like tryptophan or tyrosine in ethanol can be viewed in the nanodomain by FESEM imaging. The FESEM analysis of nanotubes prepared from tryptophan or tyrosine are shown in Fig. 3. The low-resolution FESEM images of tryptophan and tyrosine nanotubes are presented in Fig. 3a, b, respectively, and high-resolution FESEM images of tryptophan nanotubes are shown in Fig. 3c, d. In Fig. 3a there are some fibre type structures along with the nanotubes. The fibres may correspond to amyloid structures (Meital and Ehud 2006). Figure 3b shows only the tyrosine nanotubes, which are uniform in nature. These FESEM images clearly indicate that formed nanotube have wide diversity in their diameter in the case of tryptophan nanotubes. The inner diameter is ~200 nm and the length lies between 4 and 6 μm. High-resolution FESEM images clearly indicate that the formed nanotubes from tyrosine (Fig. 3b) are uniform, having nearly same length and diameter. The inner diameter of the nanotube is ~50 nm and outer is ~150 nm.
Fluorescence study
Tryptophan and tyrosine are amino acids having aromatic side chains which are fluorescent constituents in proteins. Tryptophan and tyrosine respond to fluorescence emission around 348 and 303 nm (Keleti 1970; Teale and Weber 1957). There are very fewer reports about the construction of fluorescent peptide nanotubes. Fluorescence images of tryptophan and tyrosine nanotubes are shown in Fig. 4. In this panel (a) corresponds to the tryptophan nanotubes and (b) for the tyrosine nanotubes. These nanotubes show fluorescence at different excitation wavelength viz. 385 (blue), 488 (yellow) and 561 (red) nm. This shows drastic changes in the fluorescence properties of these aggregates which could be a measure for their aggregation. Such shift in fluorescence in the visible region is due to the quantum confined phenomenon in the nanotube assembly (Amdursky et al. 2009; Fan et al. 2016).
Conclusion
The amino acid nanotubes are synthesized using simple building block such as tryptophan and tyrosine through self-assembly using ethanol as a solvent. These nanotubes synthesized are homogeneous in nature. These nanotubes show good fluorescence in varied excitation lines used. This simple strategy described herein would allow synthesizing a varied range of nanotubes from single or mixture of amnioacids and also to use these as drug carriers. The toxicity study of these aggregates may encourage such study in future.
References
Adler-Abramovich L et al. (2012) Phenylalanine assembly into toxic fibrils suggests amyloid etiology in phenylketonuria. Nat Chem Biol 8:701–706. http://www.nature.com/nchembio/journal/v8/n8/abs/nchembio.1002.html#supplementary-information
Amdursky N, Molotskii M, Aronov D, Adler-Abramovich L, Gazit E, Rosenman G (2009) Blue luminescence based on quantum confinement at peptide nanotubes. Nano Lett 9:3111–3115. doi:10.1021/nl9008265
Andrade-Filho T, Martins TC, Ferreira FF, Alves WA, Rocha AR (2016) Water-driven stabilization of diphenylalanine nanotube structures. Theor Chem Acc 135:185. doi:10.1007/s00214-016-1936-3
Arp Z, Autrey D, Laane J, Overman SA, Thomas GJ (2001) Tyrosine raman signatures of the filamentous virus Ff are diagnostic of non-hydrogen-bonded phenoxyls: demonstration by Raman and infrared spectroscopy of p-cresol vapor. Biochemistry 40:2522–2529. doi:10.1021/bi0023753
Ashton PR, Brown CL, Menzer S, Nepogodiev SA, Stoddart JF, Williams DJ (1996) Synthetic cyclic oligosaccharides-syntheses and structural properties of a cyclo[(1 → 4)-α-L-rhamnopyranosyl-(1 → 4)-α-D-mannopyranosyl]trioside and -tetraoside. Chem A Eur J 2:580–591. doi:10.1002/chem.19960020518
Begum A, Tripathi KM, Sarkar S (2014) Water induced formation, characterization, photoluminescence of carbon nanotube based composites of gadolinium(III)- and platinum(II)- dithiolenes. Chem Eur J 20:1–6. doi:10.1002/chem.201404461
Cao X, Fischer G (1999) Infrared spectral, structural, and conformational studies of zwitterionic l-tryptophan. J Phys Chem A 103:9995–10003. doi:10.1021/jp992421c
Carny O, Shalev DE, Gazit E (2006) Fabrication of coaxial metal nanocables using a self-assembled peptide nanotube scaffold. Nano Lett 6:1594–1597. doi:10.1021/nl060468l
Carubelli CR, Massabni A, Leite SRA (1997) Study of the binding of Eu3 + and Tb3 + to l-phenylalanine and l-tryptophan. J Braz Chem Soc 8:597–602
Clark TD, Buehler LK, Ghadiri MR (1998) Self-assembling cyclic β3-peptide nanotubes as artificial transmembrane ion channels. J Am Chem Soc 120:651–656. doi:10.1021/ja972786f
Encinar JA et al (2002) Tyrosine phosphorylation of the inactivating peptide of the shaker B potassium channel: a structural—functional correlate. Biochemistry 41:12263–12269. doi:10.1021/bi020188u
Fan Z, Sun L, Huang Y, Wang Y, Zhang M (2016) Bioinspired fluorescent dipeptide nanoparticles for targeted cancer cell imaging and real-time monitoring of drug release. Nat Nano 11:388–394 doi:10.1038/nnano.2015.312. http://www.nature.com/nnano/journal/v11/n4/abs/nnano.2015.312.html#supplementary-information
Fujibayashi K, Hariadi R, Park SH, Winfree E, Murata S (2008) Toward reliable algorithmic self-assembly of DNA tiles: a fixed-width cellular automaton pattern. Nano Lett 8:1791–1797. doi:10.1021/nl0722830
Ghadiri MR, Granja JR, Milligan RA, McRee DE, Khazanovich N (1993) Self-assembling organic nanotubes based on a cyclic peptide architecture. Nature 366:324–327
Granja JR, Ghadiri MR (1994) Channel-mediated transport of glucose across lipid bilayers. J Am Chem Soc 116:10786
Harada A, Li J, Kamachi M (1993) Synthesis of a tubular polymer from threaded cyclodextrins. Nature 364:516–518
Hartgerink JD, Granja JR, Milligan RA, Ghadiri MR (1996) Self-assembling peptide nanotubes. J Am Chem Soc 118:43–50. doi:10.1021/ja953070s
Ivanova BB (2006) IR-LD spectroscopic characterization of l-Tryptophan containing dipeptides. Spectrochim Acta Part A Mol Biomol Spectrosc 64:931–938. doi:10.1016/j.saa.2005.08.022
Keleti T (1970) The excimer fluorescence of tryptophan, tyrosine and d-glyceraldehyde-3-phosphate dehydrogenase. FEBS Lett 7:280–282. doi:10.1016/0014-5793(70)80181-8
Khazanovich N, Granja JR, McRee DE, Milligan RA, Ghadiri MR (1994) Nanoscale tubular ensembles with specified internal diameters. Design of a self-assembled nanotube with a 13-[angstrom] pore. J Am Chem Soc 116:6012
Kim J et al (2010) Role of water in directing diphenylalanine assembly into nanotubes and nanowires. Adv Mater 22:583–587. doi:10.1002/adma.200901973
Kobayashi K, Granja JR, Ghadiri MR (1995) The structural and thermodynamic basis for the formation of self‐assembled peptide nanotubes. Angew Chem Int Ed Engl 34:98
Ma L, Li Y, Li L, Wu Y, Buchet R, Ding Y (2009) Clarification of the binding model of lead(II) with a highly sensitive and selective fluoroionophore sensor by spectroscopic and structural study. Spectrochim Acta Part A Mol Biomol Spectrosc 72:306–311. doi:10.1016/j.saa.2008.09.014
Mao C, LaBean TH, Reif JH, Seeman NC (2000) Logical computation using algorithmic self-assembly of DNA triple-crossover molecules. Nature 407:493–496. http://www.nature.com/nature/journal/v407/n6803/suppinfo/407493a0_S1.html
Masaru M, Masaru A, Hiroyuki M, Masumi A, Toshimi S, Masaki K (2011) A simple N-Acyl-L-amino acid constructed metal-complexed organic nanotube having an inner diameter below 10 nm. Chem Lett 40:218–220. doi:10.1246/cl.2011.218
Meital R, Ehud G (2006) Designed aromatic homo-dipeptides: formation of ordered nanostructures and potential nanotechnological applications. Phys Biol 3:S10
Mishra A, Chauhan VS (2011) Probing the role of aromaticity in the design of dipeptide based nanostructures. Nanoscale 3:945–949. doi:10.1039/C0NR00691B
Pandit A et al (2008) Self-assembly of the octapeptide lanreotide and lanreotide-based derivatives: the role of the aromatic residues. J Pept Sci 14:66–75. doi:10.1002/psc.913
Parween S, Misra A, Ramakumar S, Chauhan VS (2014) Self-assembled dipeptide nanotubes constituted by flexible [small beta]-phenylalanine and conformationally constrained [small alpha],[small beta]-dehydrophenylalanine residues as drug delivery system. J Mater Chem B 2:3096–3106. doi:10.1039/C3TB21856B
Perween S, Chandanshive B, Kotamarthi HC, Khushalani D (2013) Single amino acid based self-assembled structure. Soft Matter 9:10141–10145. doi:10.1039/C3SM51054A
Pulawski W, Ghoshdastider U, Andrisano V, Filipek S (2012) Ubiquitous amyloids. Appl Biochem Biotechnol 166(7):1626–1643. doi:10.1007/s12010-012-9549-3
Ranganathan D, Haridas V, Gilardi R, Karle IL (1998) Self-assembling aromatic-bridged serine-based cyclodepsipeptides (Serinophanes): a demonstration of tubular structures formed through aromatic π–π interactions. J Am Chem Soc 120:10793–10800. doi:10.1021/ja982244d
Raviv U, Needleman DJ, Li Y, Miller HP, Wilson L, Safinya CR (2005) Cationic liposome–microtubule complexes: pathways to the formation of two-state lipid–protein nanotubes with open or closed ends. Proc Natl Acad Sci USA 102:11167–11172. doi:10.1073/pnas.0502183102
Reches M, Gazit E (2003) casting metal nanowires within discrete self-assembled peptide nanotubes. Science 300:625
Reches M, Gazit E (2006) Molecular self-assembly of peptide nanostructures: mechanism of association and potential uses. Curr Nanosci 2:105–111. doi:10.2174/157341306776875802
Reid PJ, Loftus C, Beeson CC (2003) Evaluating the potential of fluorinated tyrosines as spectroscopic probes of local protein environments: a UV resonance Raman study. Biochemistry 42:2441–2448. doi:10.1021/bi0202676
Ryan DM, Anderson SB, Senguen FT, Youngman RE, Nilsson BL (2010) Self-assembly and hydrogelation promoted by F5-phenylalanine. Soft Matter 6:475–479. doi:10.1039/B916738B
Seebach D, Matthews JL, Meden A, Wessels T, Baerlocher C, McCusker LB (1997) Cyclo-β-peptides: structure and tubular stacking of cyclic tetramers of 3-aminobutanoic acid as determined from powder diffraction data. Helv Chim Acta 80:173–182. doi:10.1002/hlca.19970800116
Shimizu T (2006) Self-assembled lipid nanotube hosts: the dimension control for encapsulation of nanometer-scale guest substances. J Polym Sci Part A Polym Chem 44:5137–5152. doi:10.1002/pola.21619
Sott K, Lobovkina T, Lizana L, Tokarz M, Bauer B, Konkoli Z, Orwar O (2006) Controlling enzymatic reactions by geometry in a biomimetic nanoscale network. Nano Lett 6:209–214. doi:10.1021/nl052078p
Teale FWJ, Weber G (1957) Ultraviolet fluorescence of the aromatic amino acids. Biochem J 65:476–482
Tsuguo T (1972) The syntheses of 3-(Carboxyalkyl)salicylaldehyde derivatives and their copper chelates. Bull Chem Soc Jpn 45:2113–2120. doi:10.1246/bcsj.45.2113
Zhao Y, Mahaja N, Fang J (2006) Self-assembled cylindrical lipid tubules with a birefringent core. Small 2:364–367. doi:10.1002/smll.200500430
Acknowledgements
DGB thanks, IIT Kanpur for a Senior Research Fellowship and SS thanks the SERB-DST, New Delhi for funding this work.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Babar, D.G., Sarkar, S. Self-assembled nanotubes from single fluorescent amino acid. Appl Nanosci 7, 101–107 (2017). https://doi.org/10.1007/s13204-017-0551-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13204-017-0551-5