Abstract
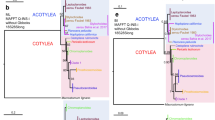
Mimopidae is a monotypic family of scolopendromorph centipedes known only by its holotype until the recent rediscovery of Mimops orientalis from Qinling Mountain, China. Here we generated novel transcriptomic data for M. orientalis and analyzed them in conjunction with other scolopendromorph and centipede transcriptomes. We used a diversity of approaches, including analyzing three matrices with different occupancy thresholds and a diversity of phylogenetic methods, including concatenation and coalescent-based approaches. All our analyses supported a phylogenetic position of Mimopidae as sister group to all other scolopendromorphs with maximal support. This position contrasts with previous Sanger data for Mimopidae but provide stability to the scolopendromorph phylogeny, showing that the loss of eyes occurred in the common ancestor of the clade leading to Cryptopidae, Plutoniumidae and Scolopocryptopidae. Our analysis thus resolves a longstanding question in centipede evolution while providing a stable relationship for the least studied family of centipedes to date.


Similar content being viewed by others
Data availability
Raw reads are publicly available in SRA archives (https://www.ncbi.nlm.nih.gov/sra); assembled transcriptomes, data matrices, analysis files and resulting tree files are available in the Harvard Dataverse (https://dataverse.harvard.edu/dataset.xhtml?persistentId=doi:10.7910/DVN/IJYRNB).
References
Altenhoff, A. M., Levy, J., Zarowiecki, M., Tomiczek, B., Vesztrocy, A. W., Dalquen, D. A., Müller, S., Telford, M. J., Glover, N. M., Dylus, D., & Dessimoz, C. (2019). OMA standalone: Orthology inference among public and custom genomes and transcriptomes. Genome Research, 29(7), 1152–1163. https://doi.org/10.1101/gr.243212.118
Ballesteros, J. A., & Sharma, P. P. (2019). A critical appraisal of the placement of Xiphosura (Chelicerata) with account of known sources of phylogenetic error. Systematic Biology, 68(6), 896–917. https://doi.org/10.1093/sysbio/syz011
Borucki, H. (1996). Evolution und Phylogenetisches System der Chilopoda (Mandibulata, Tracheata). Verhandlungen Des Naturwissenschaftlichen Vereins in Hamburg, 35, 95–226.
Crotty, S. M., Minh, B. Q., Bean, N. G., Holland, B. R., Tuke, J., Jermiin, L. S., & von Haeseler, A. (2020). GHOST: Recovering historical signal from heterotachously-evolved sequence alignments. Systematic Biology, 69(2), 249–264. https://doi.org/10.1093/sysbio/syz051
Dohle, W. (1985). Phylogenetic pathways in the Chilopoda. Bijdragen Tot De Dierkunde, 55(1), 55–66.
Edgecombe, G. D., & Bonato, L. (2011). Chilopoda – Taxonomic overview. Order Scolopendromorpha. In A. Minelli (Ed.), Treatise on Zoology - Anatomy, Taxonomy, Biology. The Myriapoda, Volume 1 (Brill ed., pp. 392–407): Brill.
Edgecombe, G. D., & Giribet, G. (2002). Myriapod phylogeny and the relationships of Chilopoda. In J. E. Llorente Bousquets, & J. J. Morrone (Eds.), Biodiversidad, taxonomía y biogeografía de artrópodos de México: Hacia una síntesis de su conocimiento (pp. 143–168). Mexico D.F.: Prensas de Ciencias, Universidad Nacional Autónoma de México.
Edgecombe, G. D., & Giribet, G. (2004). Adding mitochondrial sequence data (16S rRNA and cytochrome c oxidase subunit I) to the phylogeny of centipedes (Myriapoda, Chilopoda): An analysis of morphology and four molecular loci. Journal of Zoological Systematics and Evolutionary Research, 42, 89–134.
Edgecombe, G. D., & Giribet, G. (2007). Evolutionary biology of centipedes (Myriapoda: Chilopoda). Annual Review of Entomology, 52, 151–170. https://doi.org/10.1146/annurev.ento.52.110405.091326
Edgecombe, G. D., Giribet, G., & Wheeler, W. C. (1999). Phylogeny of Chilopoda: Combining 18S and 28S rRNA sequences and morphology. Boletín De La Sociedad Entomológica Aragonesa, 26, 293–331.
Edgecombe, G. D., & Koch, M. (2008). Phylogeny of scolopendromorph centipedes (Chilopoda): Morphological analysis featuring characters from the peristomatic area. Cladistics, 24, 872–901.
Edgecombe, G. D., & Koch, M. (2009). The contribution of preoral chamber and foregut morphology to the phylogenetics of Scolopendromorpha (Chilopoda). Soil Organisms, 81(3), 295–318.
Fernández, R., Edgecombe, G. D., & Giribet, G. (2016). Exploring phylogenetic relationships within Myriapoda and the effects of matrix composition and occupancy on phylogenomic reconstruction. Systematic Biology, 65(5), 871–889. https://doi.org/10.1093/sysbio/syw041
Fernández, R., Edgecombe, G. D., & Giribet, G. (2018). Phylogenomics illuminates the backbone of the Myriapoda Tree of Life and reconciles morphological and molecular phylogenies. Scientific Reports, 8(1), 83. https://doi.org/10.1038/s41598-017-18562-w
Fernández, R., Laumer, C. E., Vahtera, V., Libro, S., Kaluziak, S., Sharma, P. P., Pérez-Porro, A. R., Edgecombe, G. D., & Giribet, G. (2014). Evaluating topological conflict in centipede phylogeny using transcriptomic data sets. Molecular Biology and Evolution, 31(6), 1500–1513. https://doi.org/10.1093/molbev/msu108
Fu, L. M., Niu, B. F., Zhu, Z. W., Wu, S. T., & Li, W. Z. (2012). CD-HIT: Accelerated for clustering the next-generation sequencing data. Bioinformatics, 28(23), 3150–3152. https://doi.org/10.1093/Bioinformatics/Bts565
Giribet, G., Carranza, S., Riutort, M., Baguñà, J., & Ribera, C. (1999). Internal phylogeny of the Chilopoda (Myriapoda, Arthropoda) using complete 18S rDNA and partial 28S rDNA sequences. Philosophical Transactions of the Royal Society of London b: Biological Sciences, 354(1380), 215–222. https://doi.org/10.1098/rstb.1999.0373
Giribet, G., & Edgecombe, G. D. (2006). Conflict between datasets and phylogeny of centipedes: An analysis based on seven genes and morphology. Proceedings of the Royal Society b: Biological Sciences, 273(1586), 531–538. https://doi.org/10.1098/rspb.2005.3365
Haas, B. J., Papanicolaou, A., Yassour, M., Grabherr, M., Blood, P. D., Bowden, J., Couger, M. B., Eccles, D., Li, B., Lieber, M., Macmanes, M. D., Ott, M., Orvis, J., Pochet, N., Strozzi, F., Weeks, N., Westerman, R., William, T., Dewey, C. N., & Regev, A. (2013). De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nature Protocols, 8(8), 1494–1512. https://doi.org/10.1038/nprot.2013.084
Hoang, D. T., Chernomor, O., von Haeseler, A., Minh, B. Q., & Vinh, L. S. (2018). UFBoot2: Improving the Ultrafast Bootstrap approximation. Molecular Biology and Evolution, 35(2), 518–522. https://doi.org/10.1093/molbev/msx281
Jiang, C., Bai, Y., Shi, M., & Liu, J. (2020). Rediscovery and phylogenetic relationships of the scolopendromorph centipede Mimops orientalis Kraepelin, 1903 (Chilopoda): A monotypic species of Mimopidae endemic to China, for more than one century. ZooKeys, 932, 75–91. https://doi.org/10.3897/zookeys.932.51461
Katoh, K., & Standley, D. M. (2013). MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Molecular Biology and Evolution, 30(4), 772–780. https://doi.org/10.1093/Molbev/Mst010
Koch, M., Edgecombe, G. D., & Shelley, R. M. (2010). Anatomy of Ectonocryptoides (Scolopocryptopidae: Ectonocryptopinae) and the phylogeny of blind Scolopendromorpha (Chilopoda). International Journal of Myriapodology, 3(1), 51–81. https://doi.org/10.1163/187525410X12578602960344
Koch, M., Pärschke, S., & Edgecombe, G. D. (2009). Phylogenetic implications of gizzard morphology in scolopendromorph centipedes (Chilopoda). Zoologica Scripta, 38(3), 269–288. https://doi.org/10.1111/j.1463-6409.2008.00372.x
Kraepelin, K. (1903). Revision der Scolopendriden. Kommissionsverlag von Lucas Gräfe & Sillem.
Langmead, B., & Salzberg, S. L. (2012). Fast gapped-read alignment with Bowtie 2. Nature Methods, 9(4), 357–359. https://doi.org/10.1038/nmeth.1923
Lartillot, N., & Philippe, H. (2004). A Bayesian mixture model for across-site heterogeneities in the amino-acid replacement process. Molecular Biology and Evolution, 21(6), 1095–1109. https://doi.org/10.1093/molbev/msh112
Lartillot, N., Rodrigue, N., Stubbs, D., & Richer, J. (2013). PhyloBayes MPI: Phylogenetic reconstruction with infinite mixtures of profiles in a parallel environment. Systematic Biology, 62(4), 611–615. https://doi.org/10.1093/Sysbio/Syt022
Laumer, C. E., Bekkouche, N., Kerbl, A., Goetz, F., Neves, R. C., Sørensen, M. V., Kristensen, R. M., Hejnol, A., Dunn, C. W., Giribet, G., & Worsaae, K. (2015). Spiralian phylogeny informs the evolution of microscopic lineages. Current Biology, 25(15), 2000–2006. https://doi.org/10.1016/j.cub.2015.06.068
Lewis, J. G. E. (2006). On the scolopendromorph centipede genus Mimops Kraepelin, 1903, with a description of a new family (Chilopoda: Scolopendromorpha). Journal of Natural History, 40(19–20), 1231–1239. https://doi.org/10.1080/00222930600861231
Minh, B. Q., Schmidt, H. A., Chernomor, O., Schrempf, D., Woodhams, M. D., von Haeseler, A., & Lanfear, R. (2020). IQ-TREE 2: New models and efficient methods for phylogenetic inference in the genomic era. Molecular Biology and Evolution, 37(5), 1530–1534. https://doi.org/10.1093/molbev/msaa015
Murienne, J., Edgecombe, G. D., & Giribet, G. (2010). Including secondary structure, fossils and molecular dating in the centipede tree of life. Molecular Phylogenetics and Evolution, 57(1), 301–313. https://doi.org/10.1016/j.ympev.2010.06.022
Sayyari, E., & Mirarab, S. (2016). Fast coalescent-based computation of local branch support from quartet frequencies. Molecular Biology and Evolution, 33(7), 1654–1668. https://doi.org/10.1093/molbev/msw079
Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V., & Zdobnov, E. M. (2015). BUSCO: Assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics, 31(19), 3210–3212. https://doi.org/10.1093/bioinformatics/btv351
Smith, S. A., & Dunn, C. W. (2008). Phyutility: A phyloinformatics tool for trees, alignments and molecular data. Bioinformatics, 24(5), 715–716. https://doi.org/10.1093/bioinformatics/btm619
Smith, J. J., & Undheim, E. A. B. (2018). True lies: Using proteomics to assess the accuracy of transcriptome-based venomics in centipedes uncovers false positives and reveals startling intraspecific variation in Scolopendra subspinipes. Toxins, 10(3), 96, https://doi.org/10.3390/toxins10030096
Song, L., & Florea, L. (2015). Rcorrector: Efficient and accurate error correction for Illumina RNA-seq reads. GigaScience, 4, 48. https://doi.org/10.1186/s13742-015-0089-y
Stoev, P., Komerički, A., Akkari, N., Liu, S., Zhou, X., Weigand, A. M., Hostens, J., Porco, D., & Penev, L. (2013). Transcriptomic, DNA barcoding, and micro-CT imaging data from an advanced taxonomic description of a novel centipede species (Eupolybothrus cavernicolus Komerički & Stoev, sp. n.). GigaScience Database. https://doi.org/10.5524/100063
Szucsich, N. U., Bartel, D., Blanke, A., Böhm, A., Donath, A., Fukui, M., Grove, S., Liu, S., Macek, O., Machida, R., Misof, B., Nakagaki, Y., Podsiadlowski, L., Sekiya, K., Tomizuka, S., Von Reumont, B. M., Waterhouse, R. M., Walzl, M., Meng, G., & Meusemann, K. (2020). Four myriapod relatives – but who are sisters? No end to debates on relationships among the four major myriapod subgroups. BMC Evolutionary Biology, 20(1), 144. https://doi.org/10.1186/s12862-020-01699-0
Vahtera, V., Edgecombe, G. D., & Giribet, G. (2012a). Evolution of blindness in scolopendromorph centipedes (Chilopoda: Scolopendromorpha): Insight from an expanded sampling of molecular data. Cladistics, 28(1), 4–20. https://doi.org/10.1111/j.1096-0031.2011.00361.x
Vahtera, V., Edgecombe, G. D., & Giribet, G. (2012b). Spiracle structure in scolopendromorph centipedes (Chilopoda: Scolopendromorpha) and its contribution to phylogenetics. Zoomorphology, 131(3), 225–248. https://doi.org/10.1007/s00435-012-0157-0
Vahtera, V., Edgecombe, G. D., & Giribet, G. (2013). Phylogenetics of scolopendromorph centipedes: Can denser taxon sampling improve an artificial classification? Invertebrate Systematics, 27(5), 578–602. https://doi.org/10.1071/IS13035
Zhang, C., Sayyari, E., & Mirarab, S. (2017). ASTRAL-III: Increased scalability and impacts of contracting low support branches. In J. Meidanis, & L. Nakhleh (Eds.). RECOMB International Workshop on Comparative Genomics, 10562, 53–75. https://doi.org/10.1007/978-3-319-67979-2_4
Acknowledgements
We thank Mr Jiazhou Lu for help in fieldwork. Collections were made under the permit of the Fourth National Survey Chinese Material Medical Resources. Greg Edgecombe and an anonymous reviewer provided thorough comments that helped us refine this manuscript.
Funding
This work has been supported by the National Natural Science Foundation of China (Grant #82073972) and by internal funds from the Museum of Comparative Zoology and the Harvard Faculty of Arts and Sciences.
Author information
Authors and Affiliations
Contributions
All authors designed the study; CJ collected specimens and generated novel transcriptomic data; LRB analyzed the data; GG and LRB wrote the first draft; All authors revised the final draft.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Benavides, L.R., Jiang, C. & Giribet, G. Mimopidae is the sister group to all other scolopendromorph centipedes (Chilopoda, Scolopendromorpha): a phylotranscriptomic approach. Org Divers Evol 21, 591–598 (2021). https://doi.org/10.1007/s13127-021-00502-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13127-021-00502-2