Abstract
Flooding stress causes severe yield reduction in soybean worldwide. The development of stress-tolerant cultivars could be an effective measure to reduce the negative effects of flooding stress. Molecular information on the gene expression pattern of tolerant and susceptible genotypes under flooding stress could be valuable to improve the flooding tolerance in soybean. The objective of this study was to analyze the differentially expressed genes (DEGs) revealed by RNA sequencing in the soybean leaf tissues of tolerant (‘Paldalkong’ and ‘Danbaekkong’) and susceptible (‘NTS1116’) cultivars under flooding stress. Seedlings were grown in a well-watered condition up to the V1–V2 stage and flood-stressed by inundating ~ 10-cm water for 14 days. A total of 22,468 genes were differentially expressed in flood-stressed condition compared to the well-watered control condition, out of which 13,729, 13,405, and 13,160 were differentially expressed in ‘Paldalkong’, ‘Danbaekkong’, and ‘NTS1116’, respectively. A higher number of some of the flooding tolerance-related genes such as lipoxygenase, expansin, glutathione S-transferase, and sugar efflux transporter were up-regulated in the tolerant cultivars than in the susceptible cultivar. The number of some abscisic acid-related transcription factors of basic leucine zipper domain and myeloblastosis families was also higher in the tolerant cultivars than in the susceptible cultivar. The molecular information about the DEGs of tolerant and susceptible cultivars obtained in the present study could be valuable to improve the flooding tolerance in soybeans.
Similar content being viewed by others
Introduction
Considering the multiple uses of soybean as sources of food, feed, biodiesel, and other industrial products, extensive efforts have been made to increase its yield worldwide. The rise in commercial value of and demand for soybean in national and international markets has increased the soybean cultivation area. Although soybean is an upland crop, the cultivation of soybean in converted paddy fields has been increased for a few years owing to comparative economic benefit, nutrient management, and government policy (Nishida et al. 2013; Singh 2010). Soybean crop is generally sensitive to flooding stress which hampers crop growth and causes significant yield reduction (Ahmed et al. 2013; Komatsu et al. 2009). Flooding stress results from heavy rainfall and/or excessive irrigation coupled with poor drainage. This situation is further worsened under paddy soils which are specially developed to hold water for longer durations.
Plants, under flooding stress, make efforts to overcome the stress by adopting various mechanisms, including flooding avoidance, flooding tolerance, and flooding adaptation (Bailey-Serres et al. 2012; Mustroph 2018; Yamauchi et al. 2018). Various adaptive mechanisms, as well as physiological, biochemical, and molecular responses under drought and flooding stresses, help to survive and retain normal growth in plants (Dat et al. 2004). Plants perceive environmental signals that are subsequently transmitted to molecular signaling as the initial step in a stress response. Among various signal compounds in several pathways of stress responses, abscisic acid (ABA) dependent and independent responses include transcription factors (TFs). Regular transcription and its regulation rely on specific protein factors, TFs, which bind to certain DNA sequences in gene regulatory regions and control their transcription (Latchman 1993). A number of TFs of diverse families account for approximately 10% of the genes in soybean that are participated in many biotic and abiotic stress responses (http://planttfdb.cbi.pku.edu.cn/index.php?sp=Gma).
A number of efforts have been made to unravel the complex molecular mechanisms underlying various biotic and abiotic stresses in soybean. Molecular techniques such as microarray profiling have been conducted to produce gene expression data of soybeans and stored in a public database (https://www.ncbi.nlm.nih.gov/geo/). The microarray platforms have several drawbacks, such as limited sensitivity, cross- and non-specific hybridization, and limited usefulness under scarce gene information, especially for soybean in which gene models have not been well characterized. RNA sequencing (RNA-Seq), a sequencing-based recently advanced techniques, can overcome the limitations of microarray techniques (Trapnell et al. 2013). Several RNA-Seq studies have been conducted to investigate the gene expression in numerous tissues of soybean under biotic and abiotic stresses, such as drought (Vidal et al. 2012), drought and flooding (Chen et al. 2016), common cutworm attack (Du et al. 2019), salt (Zeng et al. 2019), and seed-flooding (Sharmin et al. 2020). The objective of this study was to analyze the differentially expressed genes (DEGs) revealed by RNA-Seq in soybean leaf tissue of tolerant and susceptible cultivars under flooding stress.
Materials and methods
Plant material and growing condition
Two flood-tolerant (‘Paldalkong’ and ‘Danbaekkong’) and one flood-susceptible (‘NTS1116’) soybean cultivars were selected on the basis of a previous study (Koo et al. 2014). Plants were grown in round-bottomed pots of 20 × 30 cm dimension (top and bottom diameters × height), kept in the greenhouse of Department of Southern Area Crop Science, National Institute of Crop Science, Miryang, Republic of Korea in April of 2019. The pots were filled with the soils prepared by mixing upland soil, compost, and nursery soil at 1:1:0.75 ratio. Five seeds were sown in three replicates for each cultivar and treatment condition. Seedlings were thinned by the first trifoliate (V1) stage to keep three plants in each pot. The plants were grown in a well-watered condition up to the V1–V2 stage and flood-stressed by inundating ~ 10-cm water for 14 days. The control-designated pots were grown in the well-watered condition during the period. Leaf samples were collected at 14 days after flooding (DAF) in 1.5-mL tubes, immediately kept into liquid nitrogen, and stored at – 80 °C until RNA extraction.
Measurement of flooding tolerance
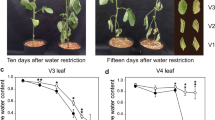
Leaf chlorophyll content (CC) was measured on the second trifoliate leaves of the control and flooded plants at 2–3-day intervals using a chlorophyll meter (SPAD-502Plus, Minolta Camera Co., Osaka, Japan). Leaf chlorophyll index (CCI) was calculated as the ratio of mean CC values obtained under flooded to control conditions (Dhungana et al. 2019). Three replications were maintained for each cultivar and treatment condition and the mean values of three replicates were reported.
RNA extraction and library preparation
Total RNA was extracted using an RNA extraction kit (RNeasy Plant Mini Kit, Qiagen, Hilden, Germany) following the manufacturer’s instructions. The concentration of RNA was measured with a NanoDrop 2000 spectrophotometer (Thermo Scientific, Waltham, MA, USA). Pooled samples of three biological replicates were prepared for individual cultivar and control/flooding treatment and thus six RNA samples were applied for RNA-Seq and DEGs analysis. The possible contaminated DNAs in the RNA samples were removed using DNase. Total RNA was further purified using a ribo-zero rRNA removal kit. The cDNA libraries were constructed using a TruSeq RNA Sample Preparation v2 Guide, Part # 15026495 Rev. F.
Quality filtering and mapping of RNA-Seq reads
The paired-end RNA-Seq reads were generated on the Illumina Genome Analyzer platform and the quality of the trimmed reads was evaluated with FastQC (Bolger et al. 2014). The RNA-Seq reads were aligned to the Glycine max reference genome (Gmax2.1version) using Bowtie2 aligner. The read mapping was performed using the HISAT2 program (Kim et al. 2015).
Sequence assembly and differential counting
After read mapping, transcript assembly was performed through the StringTie program (Pertea et al. 2015). The estimated gene abundance of the samples was determined in terms of the fragment per kilobase of transcript per million mapped reads (FPKM). The DEGs between the flood-stressed and control samples of three cultivars were identified using the StringTie-e option. The significantly up- and down-regulated genes of each cultivar were obtained for the flooding stress. The genes with a log2 fold change ≥ + 1.5 and ≤ − 1.5, and a false discovery rate adjusted P ≤ 0.05 were identified as DEGs.
Functional annotation and gene ontology enrichment
The DEGs were annotated for gene ontology (GO) terms and grouped into three categories: molecular function (MF), cellular component (CC), and biological process (BP). The GO enrichment analysis was done using gProfileR (Raudvere et al. 2019). The DEGs were also mapped to the Kyoto Encyclopedia of Genes and Genomes (KEGG) database of soybean to find the genes associated with different KEGG pathways. The TFs and transcription-related genes were identified using the plant transcription factor database (http://planttfdb.cbi.pku.edu.cn/index.php?sp=Gma) to analyze their potential roles in flooding tolerance.
Screening of DEGs using the QTL results for flooding tolerance
To identify the DEGs for their functions related to flooding stress response, the physical location of some DEGs in the genome was compared with the former mapping results of quantitative trait loci (QTL) associated with flooding tolerance in soybean. The recombinant inbred line (RIL) populations for QTL were developed using the tolerant (‘Paldalkong’ and ‘Danbaekkong’) and susceptible (‘NTS1116’) cultivars as parents which were also used for RNA-Seq in this study. The QTL result of one of the RIL populations was published (Dhungana et al. 2020). QTL map-based identification of DEGs may allow us to find the putative genes related to flooding tolerance and to verify the results of RNA-Seq study.
Results
Flooding tolerance
The highest CCI value at 14 DAF was found in ‘Danbaekkong’ (0.84) followed by ‘Paldalkong’ (0.72) and ‘NTS1116’ (0.67) as shown in Table 1. The CCI value of the susceptible cultivar ‘NTS1116’ comparatively decreased 7 DAF onwards.
Mapping and analysis of DEGs under flooding stress
The average number of reads was 65.3, 67.5, 69.5, 60.1, 58.5, and 69.5 million for ‘Paldalkong’ control, ‘Paldalkong’ flooding, ‘Danbaekkong’ control, ‘Danbaekkong’ flooding, ‘NTS1116’ control, and ‘NTS1116’ flooding, respectively. Over 98.8% of the raw reads from each library retained after quality control and approximately 97% of reads in each library were mapped (Table 2). A summary of the workflow for RNA-Seq analysis is shown in Supplementary Fig. S1.
In total, 22,468 genes were differentially expressed under flooding stress as compared to the control condition (Supplementary Table S1). The core set of DEGs was analyzed in three combinations, namely ‘Paldalkong’ flooding vs. ‘Paldalkong’ control, ‘Danbaekkong’ flooding vs. ‘Danbaekkong’ control, and ‘NTS1116’ flooding vs. ‘NTS1116’ control. The DEGs specific to and common between the three combinations were also determined (Fig. 1; Supplementary Table S2).
Some of the stress-related DEGs common to both tolerant cultivars ‘Paldalkong’ and ‘Danbaekkong’ such as lipoxygenase (LOC100785480) were significantly up-regulated only in the tolerant cultivars but not in the susceptible cultivar, ‘NTS1116’. Two phosphofructokinases (LOC100795344 and LOC100798766) were up-regulated and other two (LOC100782166 and LOC100820495) were down-regulated in the tolerant cultivars but were not significantly regulated in the susceptible cultivar. Four glutathione S-transferase (GST) (LOC100306119, LOC100796296, LOC106796199, and LOC100793025) were up-regulated and one LOC100775492 was down-regulated in the tolerant cultivars; however, none of these five genes were significantly regulated in the susceptible cultivar. Similarly, two sugar efflux transporter (SWEET) genes (GMSWEET30 and GMSWEET40) were up-regulated in the tolerant cultivars but not in the susceptible cultivar.
The analysis of stress-related DEGs common to tolerant and susceptible cultivars showed mixed results, that is some of the DEGs were significantly up-regulated and others were down-regulated in the tolerant as well as in the susceptible cultivars. Lipoxygenase LOC100811820 was up-regulated in ‘Paldalkong’ and down-regulated in ‘NTS1116’; LOC100793614 was down-regulated in ‘Danbaekkong’ but up-regulated in ‘NTS1116’; whereas, LOC100802887 and LOC100800451 were down-regulated in ‘Paldalkong’ and ‘NTS1116’. Expansin-coding gene LOC100799370 was up-regulated in ‘Paldalkong’ but down-regulated in ‘NTS1116’; LOC100807807 was up-regulated in ‘Danbaekkong’ but down-regulated in ‘Paldalkong’ and ‘NTS1116’; whereas LOC100807183, EXP2, and EXPB8 were down-regulated in ‘Paldalkong’ and ‘NTS1116’. Phosphofructokinase LOC100815278 was up-regulated in ‘Paldalkong’ but down-regulated in ‘NTS1116’; LOC100818755 and LOC100803139 were down-regulated in ‘Paldalkong’ but up-regulated in ‘NTS1116’; whereas, LOC100780164 was up-regulated in ‘Danbaekkong’ and ‘NTS1116’. Three GST were up-regulated in ‘Paldalkong’, out of which only one GSTU58 was up-regulated and the other two LOC100805827 and LOC548030 were down-regulated in ‘NTS1116’; whereas, EF1BGAMMA3 and GSTZ3 were down-regulated in the tolerant as well as susceptible cultivars. Three SWEET genes GMSWEET21, GMSWEET29, and GMSWEET42 were down-regulated in ‘Danbaekkong’ and ‘NTS1116’.
A total of 13,729, 13,405, and 13,160 genes in ‘Paldalkong’, ‘Danbaekkong’, and ‘NTS1116’, respectively, showed significant changes (P ≤ 0.05, fold change ≥ + 1.5 or ≤ − 1.5) under the control and flooded conditions. The number of up- and down-regulated genes found in ‘Paldalkong’, ‘Danbaekkong’, and ‘NTS1116’ were 6654 and 7075; 6017 and 7388; and 6623 and 6537, respectively (Fig. 2).
Functional annotation and gene ontology (GO) enrichment
The GO analysis was performed to observe the gene expression pattern based on three categories: BP, MF, and CC. The DEGs were functionally annotated to investigate their GO (Supplementary Fig. S2) and were also mapped to KEGG pathways to analyze their functional enrichment. The GO functional annotations indicated that the DEGs were involved in a series of biological processes, such as photosynthesis, light reaction, biosynthetic process, metabolic process, and cell wall biogenesis. The DEGs were also related to the following molecular functions: binding, transporter, and signal receptor. In addition, the DEGs were components of photosystem, membranes, cells, and organelles. The KEGG pathway analysis revealed that some of the major pathways were biosynthesis of secondary metabolites, carbon metabolism, biosynthesis of amino acids, glycolysis/gluconeoprotein, starch and sucrose metabolism, and oxidative phosphorylation (Supplementary Fig. S3).
A total of 462, 464, and 437 differentially expressed TFs were identified in ‘Paldalkong’, ‘Danbaekkong’, and ‘NTS1116’, respectively. The RNA-Seq analysis showed that 308 and 154; 244 and 220; and 286 and 151 TFs were up- and down-regulated in ‘Paldalkong’, ‘Danbaekkong’, and ‘NTS1116’, respectively (Fig. 3). The number of the up-regulated TFs was higher than that of the down-regulated TFs in all cultivars. The bHLH, bZIP, ERF, MYB, and WRKY families, some of the TF families related to stress tolerance, represented the majority of the differentially expressed TFs in all cultivars. The number of up-regulated TFs in bHLH, ERF, and WRKY was higher in the susceptible cultivar; whereas that of bZIP and MYB was higher in the tolerant cultivars. Another stress-related TF family G2-like also consisted of a higher number of up-regulated TF in the tolerant cultivars than in the susceptible cultivar.
The number of transcription factor (TF) of different families identified in DEGs of three cultivars. A total of 462 (308 up- and 154 down-regulated) in ‘Paldalkong’ (a), 464 (244 up- and 220 down-regulated) in ‘Danbaekkong’ (b), and 437 (286 up- and 151 down-regulated) in ‘NTS1116’ (c) TFs were differentially expressed
Differentially expressed genes in the QTL region
Several abiotic stress-related DEGs including GST (GLYMA_11G216400), xyloglucan endotransglucosylase/hydrolase (LOC100799976), polygalacturonase (GLYMA_07G066900), calmodulin-binding protein (GLYMA_07G008400), cysteine protease (GLYMA_12G039400), soybean NAC gene (GLYMA_07G050600), DEAD-box RNA helicase gene (GLYMA_07G056600), and dehydration-responsive element-binding (GLYMA_07G017300) reside in the QTL associated with flooding tolerance in soybean. Moreover, all of these genes (except GST) were found in the QTL hotspots which were the regions containing relatively more stable QTL with higher phenotypic variation. Some of these DEGs, which were found in the QTL and revealed by the RNA-Seq analysis, were up-regulated in the tolerant and down-regulated in the susceptible cultivar (Supplementary Table S1).
Discussion
The results indicated that the flooding tolerance level of susceptible cultivar reduces as the flooding period extends. Several genes are found to be differentially expressed in various abiotic stresses including flooding, drought, salinity, and low temperature (Deshmukh et al. 2014; Patil et al. 2016). Although several physiological and molecular mechanisms are involved in specific stress responses, the majority of genes and pathways are common across various stress conditions (Deshmukh et al. 2014). The RNA-Seq expression analysis in this study revealed the DEGs in flood-tolerant and flood-susceptible cultivars under flooding stress conditions. The results of the up- and down-regulated genes in the tolerant and susceptible cultivars could provide useful information for the comparative transcriptional analysis of cultivars with differential stress tolerance. Such transcriptional analyses are useful in collecting the information about candidate genes associated with flooding tolerance that would help develop flooding tolerant cultivars.
Plants undergo various adaptive mechanisms, including molecular responses by regulating several genes, to survive and retain normal growth under stress conditions. In this study, several genes were also found to be differentially expressed in tolerant and susceptible cultivars under flooding stress. The analysis of DEGs was basically focused on the genes that were related to stress tolerance mechanism. Relative expressions of genes such as lipoxygenase, expansin, phosphofructokinase, GST, and SWEET between the tolerant and susceptible cultivars were analyzed. Although some of the stress-related genes were differently expressed across cultivars, the overall analysis showed that a relatively higher number of genes were up-regulated in the tolerant than in the susceptible cultivars. Lipoxygenase was reported to play roles in the adaption of radish to biotic and abiotic stresses (Wang et al. 2019). Results with higher numbers of up-regulated expansin-coding genes were also found in a seed-flooding tolerant soybean (Sharmin et al. 2020). The roles of expansin on germination, root length, and the number of lateral roots under abiotic stresses have been reported (Lü et al. 2013; Marowa et al. 2016). Increased expression of phosphofructokinase, a glycolytic enzyme involved in the glycolytic pathway, is associated with adaptation to flooding and drought stresses in soybean (Oh and Komatsu 2015). The GST is believed to play a protective role against flooding and drought stresses in soybean (Chen et al. 2016; Oh and Komatsu 2015). The GSTs scavenge the reactive oxygen species, which may be produced in high concentration due to stresses (Klok et al. 2002) leading to cell death (Clement et al. 2008; Wrzaczek et al. 2011), and protect plants from oxidative damage. Up-regulations of several SWEET genes were also observed in tolerant soybean lines under drought and flooding stresses (Syed et al. 2015). It is implied that the regulation of SWEET genes affects sugar balancing under flooding stress condition (Mutava et al. 2015). The role of SWEET gene on plant development and stress tolerance has also been reported in Arabidopsis (Klemens et al. 2013).
As in a previous study (Sharmin et al. 2020), some cell wall-related genes such as GLYMA_08G249800 and GLYMA_18G272100 associated with COBRA-like protein were significantly up- and down-regulated in tolerant and susceptible cultivars, respectively. However, a higher number of the genes related to proline-rich cell wall protein, which were up-regulated in tolerant genotype (Sharmin et al. 2020), were down-regulated in the present study. Sharmin et al. (2020) reported that these cell wall-related genes had an important function in retaining the cell wall flexibility under flooding stress condition.
Abscisic acid (ABA) and other upstream signals may regulate the downstream pathways in abiotic stress responses (Tuteja 2007). A few number of the ABA-related TFs such as AP2, bZIP, and MYB were up-regulated in ‘Paldalkong’ and/or ‘Danbaekkong’ than in ‘NTS1116’. Soybean AP2 TFs are believed to play a vital role in enhancing abiotic stress tolerance in Arabidopsis and soybean (Zhao et al. 2019). The TF families of bZIP and MYB had been also reported to have an important role in abiotic stress response in soybean, tobacco, and Arabidopsis (Cai et al. 2015; Shukla et al. 2015; Xu et al. 2016). The bZIP TFs may adjust regulation of a few stress-related genes that result into accumulation of proline (Hoang et al. 2017). Proline is supposed to contribute in stabilizing sub-cellular structures, scavenging free radicals, and exhibiting signals for cross tolerance to many stresses (Kaur and Asthir 2015). The numbers of up-regulated G2-like TF were higher in ‘Paldalkong’ (16) and ‘Danbaekkong’ (12) than in ‘NTS1116’ (7). Tahmasebi et al. (2019) implied that G2-like TFs contributed to enhance abiotic tolerance in tobacco since they participated in the formation and development of chloroplast (Liu et al. 2016).
The results of some stress-related DEGs by RNA-Seq in this study were comparatively analyzed with putative genes in the QTL associated with flooding tolerance (Dhungana et al. 2020). Several genes located in the QTL regions were also found by RNA-Seq to contribute to flooding and other abiotic tolerance. For example, the cell wall remodeling enzymes such as xyloglucan endotransglucosylase/hydrolase maintains cell wall stiffness and helps overcome stress (Tenhaken 2015). Polygalacturonase has been found responsive to flooding stress in soybean (Nanjo et al. 2013). The contribution of calmodulin-binding protein in biotic and abiotic stresses tolerance has also been reported (Wang et al. 2015). Cysteine protease shows a positive impact on abiotic stress tolerance in soybean (Mangena 2020). Overexpression of the soybean NAC gene enhanced root formation and abiotic stress tolerance in transgenic Arabidopsis (Yang et al. 2019). Tomato plant with the overexpressed DEAD-box RNA helicase gene showed drought and salt tolerance (Zhu et al. 2015). The dehydration-responsive element-binding gene improved heat and drought stress tolerance in Arabidopsis (Mizoi et al. 2013). These reports further support the contribution of DEGs to flooding tolerance and increase the usefulness of their genetic information for soybean breeding programs.
Conclusion
Flood-tolerant and -susceptible soybean cultivars were grown under flooding stress and control conditions to investigate the gene expression patterns in leaf tissues using RNA-Seq. The flooding tolerance level, calculated as the leaf chlorophyll index, of susceptible cultivar reduced as flooding period prolonged. The number of DEGs in the tolerant cultivars was higher than that in the susceptible cultivar. A higher number of various genes and TFs related to stresses, including flooding, were up-regulated in the tolerant cultivars than in the susceptible cultivar. The results of QTL associated with flooding tolerance that were obtained with the same genetic background of tolerant parents, ‘Paldalkong’ and ‘Danbaekkong’ could support the findings of putative genes by RNA-Seq in this study. The variation in expression of some genes in the tolerant and susceptible cultivars could provide useful information about candidate genes associated with flooding tolerance that may be valuable to develop flooding tolerant soybean cultivars.
References
Ahmed F, Rafii MY, Ismail MR, Juraimi AS, Rahim HA, Asfaliza R, Latif MA (2013) Waterlogging tolerance of crops: breeding, mechanism of tolerance, molecular approaches, and future prospects. BioMed Res. Int. 2013:963525
Bailey-Serres J, Lee SC, Brinton E (2012) Waterproofing crops: effective flooding survival strategies. Plant Physiol 160:1698–1709
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30:2114–2120
Cai H, Tian S, Dong H, Guo C (2015) Pleiotropic effects of TaMYB3R1 on plant development and response to osmotic stress in transgenic Arabidopsis. Gene 558:227–234
Chen W, Yao Q, Patil GB, Agarwal G, Deshmukh RK, Lin L, Wang B, Wang Y, Prince SJ, Song L, Xu D (2016) Identification and comparative analysis of differential gene expression in soybean leaf tissue under drought and flooding stress revealed by RNA-Seq. Front Plant Sci 7:1044
Clement M, Lambert A, Herouart D, Boncompagni E (2008) Identification of new up-regulated genes under drought stress in soybean nodules. Gene 426:15–22
Dat JF, Capelli N, Folzer H, Bourgeade P, Badot PM (2004) Sensing and signalling during plant flooding. Plant Physiol Biochem 42:273–282
Deshmukh R, Sonah H, Patil G, Chen W, Prince S, Mutava R, Vuong T, Valliyodan B, Nguyen HT (2014) Integrating omic approaches for abiotic stress tolerance in soybean. Front Plant Sci 5:244
Dhungana SK, Kim HS, Kang BK, Seo JH, Kim HT, Shin SO, Park CH, Kwak DY (2019) Evaluation of flooding tolerance of soybean (Glycine max L. Merr.) in greenhouse under upland and paddy soil conditions. J Crop Sci Biotech 22:283–290
Dhungana SK, Kim HS, Kang BK, Seo JH, Kim HT, Shin SO, Park CH, Kwak DY (2020) Quantitative trait loci mapping for flooding tolerance at an early growth stage of soybean recombinant inbred line population. Plant Breed 139:626–638
Du H, Li X, Ning L, Qin R, Du Q, Wang Q, Song H, Huang F, Wang H, Yu D (2019) RNA-Seq analysis reveals transcript diversity and active genes after common cutworm (Spodoptera litura Fabricius) attack in resistant and susceptible wild soybean lines. BMC Genom 20:237
Hoang XLT, Nhi DNH, Thu NBA, Thao NP, Tran LP (2017) Transcription factors and their roles in signal transduction in plants under abiotic stresses. Curr Genom 18:483–497
Kaur G, Asthir BJ (2015) Proline: a key player in plant abiotic stress tolerance. Biol Planta 59:609–619
Kim D, Langmead B, Salzberg SL (2015) HISAT: a fast spliced aligner with low memory requirements. Nat Methods 12:357–360
Klemens PA, Patzke K, Deitmer J, Spinner L, Le Hir R, Bellini C, Bedu M, Chardon F, Krapp A, Neuhaus HE (2013) Overexpression of the vacuolar sugar carrier AtSWEET16 modifies germination, growth, and stress tolerance in Arabidopsis. Plant Physiol 163:1338–1352
Klok EJ, Wilson IW, Wilson D, Chapman SC, Ewing RM, Somerville SC, Peacock WJ, Dolferus R, Dennis ES (2002) Expression profile analysis of the low-oxygen response in Arabidopsis root cultures. Plant Cell 14:2481–2494
Komatsu S, Yamamoto R, Nanjo Y, Mikami Y, Yunokawa H, Sakata K (2009) A comprehensive analysis of the soybean genes and proteins expressed under flooding stress using transcriptome and proteome techniques. J Proteome Res 8:4766–4778
Koo S, Kim H, Kang B, Lee Y, Oh K, Kim H, Baek I, Yun H, Choi M (2014) Screening of flooding tolerance in soybean germplasm collection. Korean J Breed Sci 46:129–135
Latchman DS (1993) Transcription factors: an overview. Int J Exp Pathol 74:417–422
Liu F, Xu Y, Han G, Zhou L, Ali A, Zhu S, Li X (2016) Molecular evolution and genetic variation of G2-like transcription factor genes in maize. PloS One 11:e0161763
Lü P, Kang M, Jiang X, Dai F, Gao J, Zhang C (2013) RhEXPA4, a rose expansin gene, modulates leaf growth and confers drought and salt tolerance to Arabidopsis. Planta 237:1547–1559
Mangena P (2020) Phytocystatins and their potential application in the development of drought tolerance plants in soybeans (Glycine max L.). Protein Peptide Lett 27:135–144
Marowa P, Ding A, Kong Y (2016) Expansins: roles in plant growth and potential applications in crop improvement. Plant Cell Rep 35:949–965
Mizoi J, Ohori T, Moriwaki T, Kidokoro S, Todaka D, Maruyama K, Kusakabe K, Osakabe Y, Shinozaki K, Yamaguchi-Shinozaki K (2013) GmDREB2A; 2, a canonical dehydration-responsive element-binding protein2-type transcription factor in soybean, is posttranslationally regulated and mediates dehydration-responsive element-dependent gene expression. Plant Physiol 161:346–361
Mustroph A (2018) Improving flooding tolerance of crop plants. Agronomy 8:160
Mutava RN, Prince SJ, Syed NH, Song L, Valliyodan B, Chen W, Nguyen HT (2015) Understanding abiotic stress tolerance mechanisms in soybean: a comparative evaluation of soybean response to drought and flooding stress. Plant Physiol Biochem 86:109–120
Nanjo Y, Nakamura T, Komatsu S (2013) Identification of indicator proteins associated with flooding injury in soybean seedlings under label-free quantitative proteomics. J Proteome Res 12:4785–4798
Nishida M, Sekiya H, Yoshida K (2013) Status of paddy soils as affected by paddy rice and upland soybean rotation in northeast Japan, with special reference to nitrogen fertility. Soil Sci Plant Nutr 59:208–217
Oh M, Komatsu S (2015) Characterization of proteins in soybean roots under flooding and drought stresses. J Proteom 114:161–181
Patil G, Do T, Vuong TD, Valliyodan B, Lee JD, Chaudhary J, Shannon JG, Nguyen HT (2016) Genomic-assisted haplotype analysis and the development of high-throughput SNP markers for salinity tolerance in soybean. Sci Rep 6:19199
Pertea M, Pertea GM, Antonescu CM, Chang TC, Mendell JT, Salzberg SL (2015) StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat Biotech 33:290
Raudvere U, Kolberg L, Kuzmin I, Arak T, Adler P, Peterson H, Vilo J (2019) g: profiler: a web server for functional enrichment analysis and conversions of gene lists (2019 update). Nucleic Acids Res 47:W191–W198
Sharmin RA, Bhuiyan MR, Lv W, Yu Z, Chang F, Kong J, Bhat JA, Zhao T (2020) RNA-Seq based transcriptomic analysis revealed genes associated with seed-flooding tolerance in wild soybean (Glycine soja Sieb. & Zucc.). Environ Exp Bot 171:103906
Shukla PS, Gupta K, Agarwal P, Jha B, Agarwal PK (2015) Overexpression of a novel SbMYB15 from Salicornia brachiata confers salinity and dehydration tolerance by reduced oxidative damage and improved photosynthesis in transgenic tobacco. Planta 242:1291–1308
Singh G (2010) Replacing rice with soybean for sustainable agriculture in the indo-gangetic plain of India: production technology for higher productivity of soybean. Int J Agric Res 5:259–267
Syed NH, Prince SJ, Mutava RN, Patil G, Li S, Chen W, Babu V, Joshi T, Khan S, Nguyen HT (2015) Core clock, SUB1, and ABAR genes mediate flooding and drought responses via alternative splicing in soybean. J Exp Bot 66:7129–7149
Tahmasebi A, Ashrafi-Dehkordi E, Shahriari AG, Mazloomi SM, Ebrahimie E (2019) Integrative meta-analysis of transcriptomic responses to abiotic stress in cotton. Prog Biophys Mol Biol 146:112–122
Tenhaken R (2015) Cell wall remodeling under abiotic stress. Front Plant Sci 7:771
Trapnell C, Hendrickson DG, Sauvageau M, Goff L, Rinn JL, Pachter L (2013) Differential analysis of gene regulation at transcript resolution with RNA-seq. Nat Biotech 31:46
Tuteja N (2007) Abscisic acid and abiotic stress signaling. Plant Signal Behav 2:135–138
Vidal RO, do Nascimento LC, Mondego JM, Pereira GA, Carazzolle MF (2012) Identification of SNPs in RNA-seq data of two cultivars of Glycine max (soybean) differing in drought resistance. Genet Mol Biol 35:331–334
Wang G, Zeng H, Hu X, Zhu Y, Chen Y, Shen C, Wang H, Poovaiah BW, Du L (2015) Identification and expression analyses of calmodulin-binding transcription activator genes in soybean. Plant Soil 386:205–221
Wang J, Hu T, Wang W, Hu H, Wei Q, Wei X, Bao C (2019) Bioinformatics analysis of the lipoxygenase gene family in radish (Raphanus sativus) and functional characterization in response to abiotic and biotic stresses. Int J Mol Sci 20:6095
Wrzaczek M, Vainonen JP, Gauthier A, Overmyer K, Kangasjärvi J (2011) Reactive oxygen in abiotic stress perception—from genes to proteins. In: Shanker A, Venkateswarlu B (eds) Abiotic stress response in plants—physiological, biochemical and genetic perspectives. InTech, Rijeka, pp 27–55
Xu Z, Ali Z, Xu L, He X, Huang Y, Yi J, Shao H, Ma H, Zhang D (2016) The nuclear protein GmbZIP110 has transcription activation activity and plays important roles in the response to salinity stress in soybean. Sci Rep 6:20366
Yamauchi T, Colmer TD, Pedersen O, Nakazono M (2018) Regulation of root traits for internal aeration and tolerance to soil waterlogging-flooding stress. Plant Physiol 176:1118–1130
Yang X, Kim MY, Ha J, Lee SH (2019) Overexpression of the soybean NAC gene GmNAC109 increases lateral root formation and abiotic stress tolerance in transgenic Arabidopsis plants. Front Plant Sci 10:1036
Zeng A, Chen P, Korth KL, Ping J, Thomas J, Wu C, Srivastava S, Pereira A, Hancock F, Brye K, Ma J (2019) RNA sequencing analysis of salt tolerance in soybean (Glycine max). Genomics 111:629–635
Zhao MJ, Yin LJ, Liu Y, Ma J, Zheng JC, Lan JH, Fu JD, Chen M, Xu ZS, Ma YZ (2019) The ABA-induced soybean ERF transcription factor gene GmERF75 plays a role in enhancing osmotic stress tolerance in Arabidopsis and soybean. BMC Plant Biol 19:506
Zhu M, Chen G, Dong T, Wang L, Zhang J, Zhao Z, Hu Z (2015) SlDEAD31, a putative DEAD-box RNA helicase gene, regulates salt and drought tolerance and stress-related genes in tomato. PLoS One 10:e0133849
Acknowledgements
This research was supported by the Rural Development Administration Agenda Project (no. PJ01186801), Republic of Korea.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dhungana, S.K., Kim, HS., Kang, BK. et al. Analysis of differentially expressed genes in soybean leaf tissue of tolerant and susceptible cultivars under flooding stress revealed by RNA sequencing. J. Crop Sci. Biotechnol. 24, 83–91 (2021). https://doi.org/10.1007/s12892-020-00061-6
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12892-020-00061-6