Abstract
Different methods and formulae have been suggested to estimate effective population size based on pedigree data. These methods vary in their sensitivity to various sources of bias related to heterogenous pedigree knowledge or pedigree structure. We propose here to adapt a pre-existing method estimating coancestry rate for the specific purpose of monitoring genetic variability within livestock and captive populations. Coancestry rate is computed by averaging coancestries between pairs of individuals corrected by their equivalent numbers of generations, while restricting pedigree information to a maximum number of generations. Simulation demonstrated that restricting the number of generations allows a much clearer observation of the impact of recent events on genetic variability. Restricting the number of generations for the calculation of coancestry also has less bias related to incomplete pedigree, although it may overestimate effective population size due to non-independence in family sizes across generations. This strategy was tested on the Norwegian Nordland Lyngen horse, the Colblood Trotter horse, the French Avranchin sheep, and Bresse chicken, illustrating the applications of the approach for the monitoring of genetic variability.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The monitoring of genetic variability is an important component of any process for conserving and using genetic resources in a sustainable manner. The assessment of changes both in terms of demographics and genetic variability is indeed the key to understanding the temporal dynamics of populations, evaluating threats and management effectiveness, and detecting drivers of changes (Gibbs 2000). In livestock, the development of international technical standards and protocols for characterization, inventory, and monitoring of trends and associated risks is one of the strategic priorities of the Global Plan of Action for Animal Genetic Resources (FAO 2007). Therefore, there is a need to develop appropriate metrics to measure current trends and loss of genetic variability. These indicators would be great tools to support decision making in population management, for example, for use in early warning systems.
In domestic animal species and captive populations, genealogical and demographic data remain the main source of information for the monitoring of genetic variability despite increasing development of genomic tools (Lacy 2013; Leroy et al. 2017). When available, following FAO recommendations (FAO 2013), national early warning systems have integrated metrics based on pedigree information to assess the level of endangerment of a given breed (Berger and Fischerleitner 2001; Verrier et al. 2015; Duruz et al. 2017). Effective population size (Ne), which is defined as the size of an idealized population that would produce the same genetic variation as the population under study (Wright 1931), is a key parameter for that purpose.
Different methods and formulae have been proposed to estimate Ne (Leroy et al. 2013). Among the most frequently used methods is the regression of Identity by Descent (IBD) coefficients (inbreeding or coancestry) over time or generations, with \({N_e}=1/2\Delta IBD\) (Falconer and Mackay 1996). In practice, however, heterogeneous pedigree information may result in negative average IBD rates, making this approach unfitted in numerous cases (see Leroy et al. 2013; Mokhtari et al. 2014; Lewis et al. 2015 for instance). One of the most recent approaches, developed by Gutierrez et al. (2008) and improved by Cervantes et al. (2011), estimates individual IBD rates, combining individual IBD coefficients with total equivalent complete generations (EqG). This metric measures the pedigree knowledge for a given individual as the sum of the proportion of known ancestors over generations (Maignel et al. 1996). The advantage of using average coancestry rather than inbreeding as an IBD indicator for the estimate of Ne estimate is related to the fact that (i) the former uses more information than the latter (n × (n − 1)/2 coefficients versus n, with n being the number of individuals within the reference population) thus improving the precision of the estimate, and (ii) contrary to inbreeding, coancestry is not affected by pedigree structure. The method of Cervantes et al. (2011), which is adapted to pedigrees with overlapping generations, has also been shown to be more robust than other approaches considering variations due to bias in pedigree knowledge across time and populations (Leroy et al. 2013). Nevertheless, as it considers the whole pedigree information, this approach does not compute Ne on a predetermined time scale. Also, because EqG is an average of pedigree knowledge over generations and ancestral pathways, in some cases pedigree information may be highly detailed for one given parental pathway, while being unknown in the other one. This will result in an increase in EqG, while in the same time IBD is expected to stay stable. One may therefore hypothesize that an increased proportion of unknown pedigree may lead to bias in Ne estimation in models based on individual rate of IBD.
The goal of this study is to propose an improved genealogical metric of Ne, considering a restricted number of generations. This metric was tested on both a simulated data set and real data from four breeds, and discussed in relation to practical population monitoring.
Materials and methods
Approach considered
The individual coancestry rate model NeC (Cervantes et al. 2011) proposes to approximate coancestry rate ∆Cij between two individuals i and j by using the following formula, where Cij is the coancestry coefficient between those two individuals, and EqGi and EqGj are their respective equivalent complete generations:
Coancestry rates are averaged over individual pairs in a reference population (here considering sampling of 10,000 random pairs from the reference population, see Leroy et al. 2013). The Ne can then be computed using the formula \({{\text{N}}_{eC}}=1/2\overline {{\Delta C}}.\)
The restricted individual coancestry rate model for estimating effective population size (NeR) considers the same formula with changes in estimating Cij and respective EqG. Considering a restriction over G generations, coancestries CijR are estimated by calculating the inbreeding coefficient of a hypothetical offspring of individuals i and j with the method of VanRaden (1992). EqGiR and EqGjR are computed considering the most recent G generations only.
Simulation data
The simulation considered a population consisting of 1000 males and 1000 females over 10 non-overlapping generations (Supplementary File 1). Dams were sampled randomly (with replacement) from the 1000 females, and lists of 100 standard sires and 10 popular sires (sires producing a greater proportion of offspring) were sampled from the 1000 males.
Five different scenarios were considered (Table 1). In scenario a, male reproducers were randomly chosen within the sire lists, with popular sires producing 70% of the offspring. In scenario b, to simulate a selection process involving non-independence in family size across generations (Hill 1979), selection on a given trait was considered, assuming an additive polygenic model (see Supplementary File 1). Lists of popular sires and standard sires were determined at each generation by selecting the 10 + 100 males with the greatest estimated breeding values (EBVs). Scenario c was similar to scenario b, but popular sires produced 50% of the offspring to simulate a less intensive selection. In scenario d, the percentage of offspring produced by popular sires dropped from 70% (b) to 50% (c) only from generation 6 onward, to mimic a decrease in selection intensity related to improvement in genetic variability management. Scenario e was the same as scenario b, but indicators of genetic variability were estimated based on incomplete pedigree knowledge, with 5, 10% or 20% of sires being considered randomly as unknown.
Each of the scenarios was simulated 100 times. The EqG, Cij and Ne, were estimated for each replicate, considering both the individual coancestry rate model and the restricted individual coancestry rate model with G = 2–4. The simulations and computations were based on Fortran routines developed mostly based on Pedig (Boichard 2002) modules. The Fortran script used to compute NeR based on pedigree file using Pedig format is provided as Supplementary File 2.
Case studies
Four breeds were used to evaluate the methodology on real data sets: the Norwegian Nordland Lyngen horse breed (4360 individuals), the Norwegian Coldblood Trotter horse breed (78,433 individuals), the French Avranchin sheep breed (24,895 individuals), and the French Bresse chicken breed (79,463 individuals). The three mammalian breeds are raised by breeders, whereas the chicken breed has been managed since 1990 in a breeding centre (Centre de Sélection de Béchanne). Reference populations were defined as animals born over 4-year intervals between 1988 and 2015 for horse and sheep breeds, and as animals born each year between 1990 and 2015 for the chicken breed.
Results
Simulations
When considering scenario a, average Cij increased in a linear manner over generations, providing an estimate of Ne based on coancestry regression around 81. Similar to Cij, CijR increased until the number of simulated generations (i.e. the EqG) exceeded the respective threshold in number of generations for coancestry computations (see Fig. 1). NeC and NeR metrics were stable over time and were very close to the Ne estimated by regression, i.e. 78.
Trends in coancestry C and effective population size Ne using various restrictions over ten generations, based on simulated data considering no selection on a given trait (scenario a) or selection on a given trait with different selection intensities (scenario b and c) (standard error over simulations indicated)
In scenarios b and c, average Cij increased over time, at different rates, and in a nonlinear manner in the first generations (Fig. 1). This result could be related to the fact that non-independence in family size across generation brought substantial increase in IBD that could not be taken into account during the first generations of simulation. Consequently, in scenarios b and c, NeC decreased from 79 to 141, respectively, in generation 0 to asymptotic values close to 59 and 94, respectively. These estimates have been considered here as the actual values of Ne. As restricted CijR were stabilised when the number of simulated generations (i.e. the EqG) exceeded the respective generation threshold, NeR estimates also stabilized to values higher than actual Ne. The difference was all the more important since the generation threshold was low. In scenario b, the differences were relatively limited, with NeR ranging from 63 (four generations considered) to 69 (two generations considered), while these differences appeared higher in scenario c, with NeR ranging from 108 (four generations considered) to 120 (two generations considered).
When considering scenario d, with selection changing at generation 6 from the configuration described in scenario b to the one described in scenario c, the increase in Cij was proportionally reduced, while CijR decreased and then stabilized after a number of generations corresponding to their generation threshold (Fig. 2). As a consequence, NeR reached an equilibrium corresponding to the new selection intensity much more quickly than when taking all the generations into account (NeC), even though, NeR estimates at generation 10 showed the very same bias already illustrated in scenario b.
As expected, increasing rates of unknown parents reduced the increase in pedigree knowledge per generation (see Supplementary Fig. 1): at generation 10, EqG reached on average 7.6, 5.9, and 3.5 generations for 5, 10, and 20% of unknown parents, respectively. Similarly, the increase in coancestry was reduced with increasing rate of unknown parents. For 10% and 20% of unknown parents, it actually stabilized after several generations, while EqG continued to increase. Therefore, when considering all the pedigree knowledge, the NeC increased (by construction) over time, even if actual Ne was supposed to be stable (Fig. 3). Actually, all the different Ne metrics showed an overestimation of Ne, which was particularly inflated when both the percentage of unknown pedigree was high and the number of generations included in the computation of Ne was high. With 5% of unknown sires the overestimation of Ne was relatively limited when the number of generations was restricted; the NeR stabilized at around 78–80 (actual Ne = 59). When restricting the number of generations for Ne computation, the difference between NeR and actual Ne also stabilized after a few generations. However, when considering all the pedigree knowledge (NeC), this bias increased with generations, reaching 162 in generation 10 with 20% of unknown sires.
Case studies
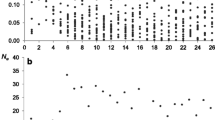
The four breeds illustrate the practical interest of NeR for population monitoring (Fig. 4a). Both Avranchin and Nordland Lyngen populations showed an increase in population size from the 1990s to the 2000s (especially for Avranchin breed) and then a decrease over the recent years. Coldblood Trotter showed a decrease in population size over the period, while the Bresse chicken showed fluctuations in terms of number of registrations. In terms of pedigree knowledge (Fig. 4b), for the Bresse chicken (maintained as a nucleus flock raised in non-overlapping generations in a breeding center), the EqG increased by 1 each year/generation over the period from 1990 to 2015, reaching 26 in 2015. Among the three mammalian breeds, Nordland Lyngen and Coldblood Trotter breeds had more complete pedigree information (EqG increasing from 6.7 to 9.3, and from 8.7 to 11, respectively between 1988 and 2015, respectively) compared to Avranchin (EqG increasing from 3.3 to 4.8 between 1988 and 2015). Also, in the horse breeds, the pedigree information was almost complete for the fifth and closer generations for all periods (see Supplementary Fig. 2). By contrast, in Avranchin breed, regardless of the period considered, the percentage of known parents (i.e. first generation) ranged from 67 to 89%.
Compared to the two other breeds, the different Ne metrics showed relatively similar trends over time for Coldblood Trotter and Avranchin breeds. In the Coldblood Trotter, where no specific effort had been implemented in terms of management of genetic variability, the different Ne metrics showed decreasing trends, probably reflecting the demographic ones. In the Avranchin breed, Ne metrics showed relatively similar trends over time, with slightly larger values for NeC compared to NeR metrics, probably in relation to pedigree information being largely unbalanced.
The, Ne metrics showed contrasting results in Nordland Lyngen. The NeC showed a worrying situation, being very low, although slightly increasing over time, from 24.5 (1988–1991) to 33.7 (2012–2015) (Fig. 4c). On the basis of NeC alone, it was not possible to conclude if this increase was related to improvement in management or to bias related to the metric itself (i.e. unbalanced pedigree from generation 6). Indeed, coancestry was stable over the period, while EqG increased. The discrepancy between coancestry and EqG, as illustrated in the simulations (Fig. 3), could explain such an increase. Given that the pedigree information was complete up to the fifth generation, it is clear that NeR metric were not biased by an unbalanced pedigree. Trends in NeR metrics, therefore, probably confirm an improvement of the genetic management, with Ne reaching values between 121 and 176 in 2012–2015 according to the number of generations considered. It may eventually be hypothesized that the decrease in NeR (G = 2) over the last period is related to the demographic decrease occurring at the same time.
The changes in Ne metrics for Bresse breed were particularly informative of the management history of the population. Indeed, over the 1990–1996 period, the breed was selected without a formal plan for management of genetic variability, but the number of sires and dams used was increasing in this period, explaining the overall positive trend in Ne metrics (data not shown). From 1997 on, the population was grouped into families of sires and dams, with a breeding scheme keeping at least a single male and a single female per sire and dam family, respectively, as contributing parents of the next generation (within-family selection). As a consequence, once the generation lag was integrated into the different NeR metrics, those indicators gave very similar estimates over the 2000–2012 period, reflecting the absence of family variance within and across generations. In 2012, a supplementary constraint was added in the breeding scheme to minimize inbreeding rate at a fixed rate of genetic gain (Chapuis et al. 2016). The corresponding selection algorithm used coancestry values to compute mating design, with those values largely depending on the number of generations considered (at least for the first generations of this scheme). Therefore, it may be hypothesized that the more contrasted results in NeR metrics observed after 2012 could be linked to some underestimation of Ne for the metrics with limited number of generations considered. These changes in breeding management were not reflected in any case in NeC values, which displayed much lower value and steady increase over the period.
Discussion
For population monitoring, breed managers require information on how external drivers (e.g. demographic changes, implementation of conservation measures) impact genetic variability in a determined (and generally recent) time scale. The aim of this study was to propose an alternate Ne model based on the change in the individual rate of coancestry, considering a restricted number of pedigree generations since pedigree knowledge is often unbalanced in the different ancestral pathways.
Considering all the pedigree information available may be a suboptimal solution for population monitoring, as the metrics of genetic variability may be influenced by the impacts of both ancient and recent events on genetic variability. The simulation study undertaken by Gutiérrez et al. (2008) already underlined that as approaches based on individual IBD rates estimate a Ne averaged over the genealogy of the population, they converge slowly to theoretical Ne in case of change in effective population size. This work and other studies (Cervantes et al. 2011) also underlined that the NeC metric seems to be an adequate method to measure Ne in breeds with structured populations and overlapping generations. Our simulations also investigated how those estimates can be biased by two other phenomena that are common in pedigreed populations, namely the non-independence in family size across generations and unbalanced pedigree knowledge.
Non-independence in family size across generations is an inherent factor in selection, which results from the fact that offspring of reproducers of interest are expected to be themselves of interest, as they are likely to inherit alleles giving them reproductive advantages (for natural or artificial selection). For example, the sons of popular sires have a greater probability to become popular sires than under random conditions, which therefore has an impact on genetic variability. The issue is related to the fact that such a phenomenon cannot be detected in the earliest generations in pedigree. This bias, however, tends to decrease as a larger number of generations is considered in the Ne metric.
The second bias is related to the existence of discrepancies in pedigree knowledge over ancestral pathways. In theory, variation in pedigree knowledge across individuals should not affect the Ne metrics based on individual IBD rates, as long as the pedigree knowledge is the same in the different ancestral pathways. Discrepancies between equivalent complete generations and IBD coefficients, in relation to unbalanced pedigree knowledge across ancestral pathways, are however frequently observed in livestock genealogies (Menéndez et al. 2016; IDELE 2016). Our simulations have shown that a random percentage of unknown sires may result in strong overestimation of Ne (Fig. 3). Those biases are all the more important since pedigree information by generation are incomplete, while Ne metrics based on restricted number of generations (NeR) are less affected. In livestock species, such unbalanced pedigree knowledge is frequent in breeds with a low percentage of AI and large herd or flock sizes. In these situations, the percentage of unknown sires is usually greater than the percentage of unknown dams. In the Avranchin breed, when considering animals born in the period 1988–2015, a much larger proportion of sires than dams were unknown (25% and 13.5% respectively). A potential solution to decrease the proportion of incomplete pedigrees relies on the probabilistic assignation of parental contributions for individuals with unknown sires or dams (Lacy 2012). However, these solutions require additional information (e.g. lists of possible parents for each animal) to infer potential parents, which are not always simple to obtain.
The four case studies illustrate quite well these different issues. In the Nordland Lyngen horse (and to a lesser extent Bresse chicken), strong founder effects, with an effective number of ancestors around seven in the case of Nordland Lyngen breed (Olsen et al. 2010) resulted in a low Ne, when considering the whole pedigree information (NeC = 33.7 for 2012–2017). However Ne estimates on a restricted number of generations (NeR) provided a much less negative picture of the situation of Nordland Lyngen. It can be underlined that in the three mammalian breeds, NeR metrics and especially NeR (G = 2), probably provide overestimated values of Ne, because non-independence in family size across generations are imperfectly taken into account. Nevertheless, the increasing difference between NeR metrics and NeC in Nordland Lyngen are probably related to improvements in genetic management over recent generations. Indeed, substantial progress has been made in the last 30 years both in terms of population size and conservation measures, either through European financial support or application of a mating plan implemented since 1995. However the future of the breed is still not bright since registrations have dropped since the 2008–2011 period in relation to the worldwide financial crisis. In the case of Bresse chicken, similar trends between NeR metrics and NeC were observed, but the comparison between the NeR variants also reflected changes in management practices that were not detected when considering the complete pedigree information. In the Avranchin breed, trends in Ne clearly reflected the changes in coancestry over time. Given the incomplete pedigree over generations, it is expected that metrics of NeR and especially NeC provide overestimated values of Ne. The fact that in the first periods (1988–2000) NeR estimates were lower when a smaller number of generations was considered, while the inverse tendency was observed later, probably indicated a change in the way those metrics were affected by the different types of bias present in different stages of the population’s history. It is however difficult to quantify the extent of these effects.
Our results show that restricting the number of generations in the calculation of Ne can be a good option for population monitoring, as it allows (i) the identification of recent changes affecting genetic variability, (ii) more precise determination of the time scale considered for Ne computation and (iii) reduction in the bias related to disequilibrium in pedigree knowledge across ancestral pathways. An important limitation of this approach is related to the fact that considering a very limited number of generations does not fully account for non-independence of family sizes across generations. The choice of the number of generations to be considered should depend on different factors (Table 2). For breeds with relatively complete pedigree records, the estimation of NeC may be of interest to provide an evaluation of changes in genetic variability over a (relatively) long period, while restricted NeR estimates should reflect more recent changes. Breed managers may also have to decide on the most relevant time scale for monitoring. Knowing the number of generations to be monitored, the time scale can be approximated by multiplying the number of generations by the generation interval. This is especially important for species with long generation intervals such as horses, which have generation intervals close to 10 years. In addition, non-independence in family size across generations and unbalanced pedigree knowledge may lead to an overestimation of the Ne estimate. In the first case, it is difficult to quantify the level of overestimation, however, restricting the number of considered generations is expected to increase the bias. In the latter case, recent genealogical studies have shown wide ranges in amount of pedigree knowledge and unbalanced pedigree, with EqG and percent of unknown parents ranging from < 2 to more than 10 generations, and from 0 to more than 30% respectively (Leroy et al. 2006, 2013; Bartolomé et al. 2011; Santana et al. 2016; Addo et al. 2017; Kumpulainen et al. 2017). From our simulations, genealogies with more than 10% of unknown parents are expected to provide Ne estimates with substantial overestimations regardless of the metrics chosen, and we may, nevertheless, recommend to restrict the number of generations considered for Ne estimation in order to limit the extent of bias.
Monitoring changes in Ne over time would allow evaluation of how the genetic variability within a livestock breed or a captive population has been impacted by recent changes in population size or population management. However this requires regular update (yearly or at least every generation) in the provision of metrics to the population managers and/or local early warning systems (FAO 2011). Having an overview of those changes should allow decision makers to better characterise drivers and threats to genetic variability, and also to identify the best measures to be undertaken for the conservation of genetic variability.
References
Addo S, Schäler J, Hinrichs D, Thaller G (2017) Genetic diversity and ancestral history of the German Angler and the red-and-white dual-purpose cattle breeds assessed through pedigree analysis. Agric Sci 8:1033
Bartolomé E, Cervantes I, Valera M, Gutiérrez JP (2011) Influence of foreign breeds on the genetic structure of the Spanish Sport Horse population. Livest Sci 142:70–79
Berger B, Fischerleitner F (2001) Conservation of AnGR—the Austrian way. https://www.rfp-europe.org/fileadmin/SITE_ERFP/country_reports/Austria/CR_Austria_2001.doc
Boichard D (2002) PEDIG: a fortran package for pedigree analysis suited for large populations. In: Proceedings of the 7th world congress of genetics applied to livestock production: 19–23 August 2002, Montpellier
Cervantes I, Goyache F, Molina A, Valera M, Gutiérrez JP (2011) Estimation of effective population size from the rate of coancestry in pedigreed populations. J Anim Breed Genet 128:56–63
Chapuis H, Pincent C, Colleau JJ (2016) Optimizing selection with several constraints in poultry breeding. J Anim Breed Genet 133:3–12
Duruz S, Flury C, Matasci G, Joerin F, Widmer I, Joost S (2017) A WebGIS platform for the monitoring of Farm Animal Genetic Resources (GENMON). PLoS ONE 12:e0176362
Falconer DS, Mackay TFC (eds) (1996) Introduction to quantitative genetics, 4th edn. Longman, White Plains, New York
FAO (2007) Global plan of action for animal genetic resources. FAO, Rome
FAO (2011) Surveying and monitoring of animal genetic resources. FAO animal production and health guidelines. No. 7. FAO, Rome
FAO (2013) In vivo conservation animal genetic resources. FAO animal production and health guidelines. No. 14. FAO, Rome
Gibbs JP (2000) Monitoring populations. In: Boitani L, Fuller TK (eds) Research techniques in animal ecology, controversies and consequences. Columbia University Press, New York, pp 213–252
Gutiérrez JP, Cervantes I, Molina A, Valera M, Goyache F (2008) Individual increase in inbreeding allows estimating effective sizes from pedigrees. Genet Sel Evol 40:359–378
Hill WG (1979) A note on effective population size with overlapping generations. Genetics 92:317–322
IDELE (2016) VARUME—Résultats 2016. http://idele.fr/presse/publication/idelesolr/recommends/varume-resultats-2016-1.html. Accessed 26 Nov 2017
Kumpulainen M, Anderson H, Svevar T, Kangasvuo I, Donner J, Pohjoismäki J (2017) Founder representation and effective population size in old versus young breeds—genetic diversity of Finnish and Nordic Spitz. J Anim Breed Genet 134:422–433
Lacy RC (2012) Extending pedigree analysis for uncertain parentage and diverse breeding systems. J Hered 103:197–205
Lacy RC (2013) Achieving true sustainability of zoo populations. Zoo Biol 32:19–26
Leroy G, Rognon X, Varlet A, Joffrin C, Verrier E (2006) Genetic variability in French dog breeds assessed by pedigree data. J Anim Breed Genet 123:1–9
Leroy G, Mary-Huard T, Verrier E, Danvy S, Charvolin E, Danchin-Burge C (2013) Methods to estimate effective population size using pedigree data: examples in dog, sheep, cattle and horse. Genet Sel Evol 45:1
Leroy G, Carroll EL, Bruford MW, DeWoody JA, Strand A, Waits L, Wang J (2017) Next-generation metrics for monitoring genetic erosion within populations of conservation concern. Evol Appl. https://doi.org/10.1111/eva.12564
Lewis TW, Abhayaratne BM, Blott SC (2015) Trends in genetic diversity for all Kennel Club registered pedigree dog breeds. Canine Genet Epidemiol 2:13
Maignel L, Boichard D, Verrier E (1996) Genetic variability of French dairy breeds estimated from pedigree information. Interbull Bull 14:49–54
Menéndez J, Álvarez I, Fernández I, Goyache F (2016) Genealogical analysis of the Gochu Asturcelta pig breed: insights for conservation. Czech J Anim Sci 61:140–143
Mokhtari MS, Shahrbabak MM, Esmailizadeh AK, Shahrbabak HM, Gutierrez JP (2014) Pedigree analysis of Iran-Black sheep and inbreeding effects on growth and reproduction traits. Small Rumin Res 116:14–20
Olsen HF, Klemetsdal G, Ruane J, Helfjord T (2010) Pedigree structure and genetic variation in the two endangered Norwegian horse breeds: Dole and Nordland/Lyngen. Acta Agric Scand A 60(1):13–22
Santana ML, Pereira RJ, Bignardi AB, Ayres DR, Menezes GRO, Silva LOC et al (2016) Structure and genetic diversity of Brazilian Zebu cattle breeds assessed by pedigree analysis. Livest Sci 187:6–15
VanRaden PM (1992) Accounting for inbreeding and crossbreeding in genetic evaluation of large populations. J Dairy Sci 75:3136–3144
Verrier E, Audiot A, Bertrand C, Chapuis H, Charvolin E, Danchin-Burge C et al (2015) Assessing the risk status of livestock breeds: a multi-indicator method applied to 178 French local breeds belonging to ten species. Anim Genet Res 57:105–118
Wright S (1931) Evolution in Mendelian populations. Genetics 16:97–159
Acknowledgements
The authors would like to thank the French Government for providing the secondment of Dr Gregoire Leroy to FAO.
Disclaimer
The views expressed in this information product are those of the authors and do not necessarily reflect the views or policies of FAO.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Authors have no conflict of interest to declare.
Additional information
The views expressed in this publication are those of the author(s) and do not necessarily reflect the views or policies of FAO.
Electronic supplementary material
Below is the link to the electronic supplementary material.
12686_2019_1092_MOESM1_ESM.pdf
Supplementary Figure 1 Trends in coancestries C and equivalent complete generations EqG over 10 generations, based on simulated data considering various percentage of unknown pedigree (PDF 298 KB)
12686_2019_1092_MOESM2_ESM.pdf
Supplementary Figure 2 Percentage of ancestor known in Nordland Lyngen, Coldblood Trotter and Avranchin breeds according to generations and period considered (PDF 522 KB)
Rights and permissions
The opinions expressed in this publication are those of the authors/editors and do not necessarily reflect the views of the [FAO], its Board of Directors, or the countries they represent.
Open Access This book is licensed under the terms of the Creative Commons Attribution 3.0 IGO License (https://creativecommons.org/licenses/by/3.0/igo/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the [FAO], provide a link to the Creative Commons licence and indicate if changes were made.
The use of the [FAO]’s name, and the use of the [FAO]’s logo, shall be subject to a separate written licence agreement between the [FAO] and the user and is not authorized as part of this CC-IGO licence. Note that the link provided above includes additional terms and conditions of the licence.
The images or other third party material in this book are included in the book’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the book’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-sa/3.0/igo/.
About this article
Cite this article
Leroy, G., Gicquel, E., Boettcher, P. et al. Coancestry rate’s estimate of effective population size for genetic variability monitoring. Conservation Genet Resour 12, 275–283 (2020). https://doi.org/10.1007/s12686-019-01092-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12686-019-01092-0