Abstract
Complex by DDMC/sncRNAs(a-miR-155, piR-30074, and miR-125b) have shown the full recovery of mice from virus-induced sarcoma after treatment by changing to normal cells from cancer cells. With kinetic of inhibiting tumor growth, the difference from control \(\varvec{(mm^2)}\) of one intravenous injection and two intravenous injections is the same curvature and rate so that this is a soliton wave having a permanent form (velocity, shape) such as one faster soliton overcoming another one without any changing shape in the case of two intravenous injections. Signal transduction for this DDS must be Hill-type sigmoids following the nonlinear Schrödinger model by transfer of energy, sine-Gordon soliton model by momentum, and Fisher KPP soliton model by mass transfer. We find out that the cell outlet/inlet response of DDMC/sncRNAs is more dependent on the soliton signal not losing energy and shape without jamming communication. The endoplasmic reticulum-mitochondrial \(\varvec{Ca^{2+}}\) fluxes induced by soliton to Chromosome in nuclear will take place epigenetic modifications on 5-position of cytosine (5mC) by TET enzymes and thymine DNA glycosylase (TDG) as an overall intracellular reaction. By soliton flow, the result shows the equations of quantum mechanics can be related to the epigenetic control with \(\varvec{Ca^{2+}}\) fluxes followed by “induced fit model” Hill equation S-shaped.



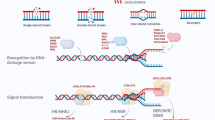
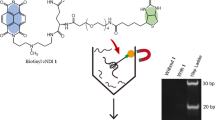
Similar content being viewed by others
Data Availability
The data that support the findings of this study are available within the article.
References
Doi, T., Kuroda, S., Michikawa, T., et al. (2005). Inositol 1,4,5-trisphosphate-dependent Ca2+ threshold dynamics detect spike timing in cerebellar Purkinje cells. The Journal of Neuroscience, 25, 950–961.
Englander, S. W., Kallenbach, J. A., & Litwin, S. (1980). Nature of the open state in long polynucleotide double helices: Possibility of soliton excitations. PNAS, 77, 7222–7226.
Onishi, Y., Ji, R., Kobayashi, T., et al. (2022). Soliton-based signaling communication and supermolecular nano-complex by ddmc/ptx in tumor micro-environment. BioNanoScience
Onishi, Y., Eshitai, Y., Ji, R. C., et al. (2018). A robust control system for targeting melanoma by a supermolecular DDMC/paclitaxel complex. Integrative Biology, 10, 549–554.
Klimenko, O. V., & Sidorov, A. (2019). The full recovery of mice (Mus Musculus c57bl/6 strain) from virus-induced sarcoma after treatment with a complex of DDMC delivery system and sncRNAs. Non-coding RNA Research, 4, 69–78.
Li, B. Q., Ma, Y. L., Li-Po, et al. (2017). Computers and mathematics with applications the n-loop soliton solutions for-dimensional Vakhnenko equation. Computers and Mathematics with Applications, 74, 504–512.
Onorato, M., Proment, D., Clauss, G., et al. (2013). Rogue waves from nonlinear Schrödinger breather solutions to sea-keeping test. PLOS ONE, 8, e54,629.
Onishi, Y., Eshita, Y., Murashita, A., et al. (2005). Synthesis and characterization of 2-diethylaminoethyl(DEAE)-dextran-MMA graft copolymer for non-viral gene delivery vector. Journal of Applied Polymer Science, 98, 9–14.
Onishi, Y., Eshita, Y., Murashita, A., et al. (2007). Characteristics of 2-diethylaminoethyl(DEAE)-dextran-MMA graft copolymer as a non-viral gene carrier. Nanomedicine: Nanotech, Biology and Medicine, 3, 184–191.
Zhang, M., Wang, J., Zhang, K., et al. (2021). Ten-eleven translocation 1 mediated-DNA hydroxymethylation is required for myelination and remyelination in the mouse brain. Nature Communications, 12, 5091.
Zhang, M., Wang, J., Zhang, K., et al. (2019). Homeostasis regulates myelination and synaptic functions. bioRxiv preprint.
Kohli, R. M., & Zhang, Y. (2013). TET enzymes, TDG and the dynamics of DNA demethylation. Nature, 502, 472–479.
Luan, F., Guerrero, R. C., Zhong, N., et al. (2014). Tetmediated formation of 5-hydroxymethylcytosine in RNA. Journal of the American Chemical Society, 136, 11582–11585.
Pecorino, L. (2017). Molecular biology of cancer: Mechanisms, targets, and therapeutics. Tokyo: Medical Sciences International.
Craig, J. M. (2011). Epigenetics: A reference manual. Caister Academic Press, Victoria.
Yomosa, S. (1974). Theory of the excited state of molecular complex in solution. Journal of the Physical Society of Japan, 36, 1655–1660.
Klimenko, O. V., & Onishi, Y. (2018). Disappeared cancer cell by sncRNAs: Application of DDMC vector/sncRNAs complex for transformation of cancer cells into non-cancerous cells. Journal of Nanomedicine and Biotherapeutic Discovery, 8, 1.
Matsumura, Y., & Maeda, H. (1986). A new concept for macromolecular therapeutics in cancer chemotherapy: Mechanism of tumoritropic accumulation of proteins and the antitumor agent smancs. Cancer Research, 12 Pt 1, 6387–6392.
Allen, T. M., & Chonn, A. (1987). Large unilamellar liposomes with low uptake into the reticuloendothelial system. FEBS Letters, 223, 42–46.
Oku, N., Namba, Y., & Okada, S. (1992). Tumor accumulation of novel res-avoiding liposomes. Biochimica et Biophysica, 1126, 255–260.
Barros, M. T. (2017). Ca2+-signaling-based molecular communication systems: Design and future research directions. Nano Communication Networks, 11, 103–113.
Huang, X., Schwind, S., Yu, B., et al. (2013). Targeted delivery of microrna-29b by transferrin-conjugated anionic lipopolyplex nanoparticles: A novel therapeutic strategy in acute myeloid leukemia. Clinical Cancer Research, 19, 2355–23,567.
Singh, R., & Saini, N. X. (2012). Downregulation of bcl2 by miRNAs augments druginduced apoptosis-a combined computational and experimental approach. Journal of Cell Science, 125, 1568–1578.
Cimmino, A., Calin, G. A., Fabbri, M., et al. (2005). miR-15 and miR-16 induce apoptosis by targeting bcl2. Proceedings of the National Academy of Sciences of the United States of America, 102, 13,944-13,949.
Bouchie, A. (2013). First microRNA mimic enters clinic. Nature Biotechnology, 31, 577.
Daige, C. L., Wiggins, J. F., Priddy, L., et al. (2014). Systemic delivery of a mir34a mimic as a potential therapeutic for liver cancer. Molecular Cancer Therapeutics, 13, 2352–2360.
Agostini, M., & Knight, R. A. (2014). miR-34: From bench to bedside. Oncotarget, 5, 872–881.
Pileczki, V., Cojocneanu-Petric, R., Maralani, M., et al. (2016). Sandulescu, r. micrornas as regulators of apoptosis mechanisms in cancer. Clujul Medical, 89, 50–55.
Klimenko, O. V. (2017). Joint action of the nano-sized system of small noncoding RNAs with DDMC vector and recombinant IL-7 reprograms A-549 lung adenocarcinoma cells into cd4+ cells. Immunotherapy (Los Angel), 3, 137.
Liu, X., Li, J., Qin, F., et al. (2016). miR-152 as a tumor suppressor microRNA: Target recognition and regulation in cancer. Oncology Letters, 11, 3911–3916.
Zhang, Y. J., Liu, X. C., Du, J., et al. (2015). miR-152 regulates metastases of non-small cell lung cancer cells by targeting neuropilin-1. International Journal of Clinical and Experimental Pathology, 8, 14,235-1424.
Klimenko, O. V. (2017). Small non-coding RNAs as regulators of structural evolution and carcinogenesis. Non-coding RNA Research, 2, 88–92.
Pal-Bhadra, M., Leibovitch, B. A., Gandhi, S. G., et al. (2004). Heterochromatic silencing and HP1 localization in drosophila are dependent on the RNAi machinery. Science, 303, 669–672.
Brower-Toland, B., Findley, S. D., Jiang, L., et al. (2007). Drosophila PIWI associates with chromatin and interacts directly with HP1a. Genes and Development, 21, 2300–2310.
Yin, H., & Lin, H. (2007). An epigenetic activation role of PIWI and a PIWI-associated piRNA in Drosophila melanogaster. Nature, 450, 304–308.
Tian, Y., Guo, R., Shi, B., et al. (2016). MicroRNA-30a promotes chondrogenic differentiation of mesenchymal stem cells through inhibiting delta-like 4 expression. Life Sciences, 148, 220–228.
Otto, T., Candido, S. V., Pilarz, M. S., et al. (2017). Cell cycle-targeting microRNAs promote differentiation by enforcing cell-cycle exit. Proceedings of the National Academy of Sciences of the United States of America, 114, 10,660-10,665.
Jin, M., Wu, Y., Wang, Y., et al. (2016). MicroRNA-29a promotes smooth muscle cell differentiation by targeting YY1. Stem Cell Research, 17, 277–284.
Chen, C. H., Lu, H. T., Tsuang, Y. H., et al. (2017). MicroRNA-215 promotes proliferation and differentiation of osteoblasts by regulation of c-fos. International Journal of Clinical and Experimental Pathology, 10, 6536–6543.
Le, M. T. N., Xie, H., Zhou, B., et al. (2009). MicroRNA-125b promotes neuronal differentiation in human cells by repressing multiple targets. Molecular Cell Biology, 29, 5290–5305.
Fujii, T., Shimada, K., Tatsumi, Y., et al. (2015). MicroRNA-145 promotes differentiation in human urothelial carcinoma through down-regulation of syndecan-1. BMC Cancer, 15, 818.
Mai, H., Lini, Y., Zhaoi, Z. A., et al. (2016). MicroRNA-127 promotes mesendoderm differentiation of mouse embryonic stem cells by targeting leftright determination factor 2. Journal of Biological Chemistry, 291, 12,126-12,135.
Klimenko, O. V., & Shtilman, M. I. (2013). Transfection of Kasumi-1 cells with a new type of polymer carriers loaded with miR-155 and antago-miR-155. Cancer Gene Therapy, 20, 237–241.
Yoshikawa, Y., Tsumoto, K., & Yoshikawa, K. (2002). Switching of higher-order structure of DNA and gene expression. Seibutsu Butsuri, 42, 179–184.
Klimenko, O. V. (2017). Toxicity and transfection efficiency of new non-viral delivery systems for small non-coding RNAs: Amphiphilic poly(n-vinylpyrrolidone) and diethylaminoethyl-dextran-methacrylic acid methyl ester copolymer. Advanced Science, Engineering and Medicine, 9, 426–431.
Suzuki, H., Yamagata, K., Sugimoto, K., et al. (2009). Modulation of microRNA processing by p53. Nature, 460, 529–533.
Filippov, A. (2000). The versatile soliton: Modern Birkhauser classics. Boston: Birkhauser.
Braun, O. M., & Kivshar, Y. (2004). The Frenkel-Kontorova model: Concepts, methods, and applications. Berlin: Springer-Verlag.
Novikov, S. P., Manakov, S. V., Pitaevskiii, L. P., et al. (1984). Theory of solitons: The inverse scattering method. Berlin: Springer-Verlag.
Helmholtz, H. (1850). Messungen uber den zeitlichen verlauf der zuchung animalisher muskeln und die fortplanzungsgeschwindig-keit der reizung in der nerven. Archives of Anatomy and Physiology, 276, 71–73.
Hodgkin, A. L., & Huxley, A. F. (1952). A quantitative description of membrane current and its application to conduction and excitation in nerve. Journal of Physiology, 117, 500–544.
Fitzhugh, R. (1969). Biological engineering. New York: McGraw-Hill.
Nagumo, J., Arimoto, S., & Yoshizawa, S. (1962). An active pulse transmission line simulating nerve axon. Proceedings of the IRE, 50, 2061–2070.
Scott, A. (1979). Sine-gordon breather dynamics. Physica Scripta, 20, 395–401.
Frank, S. A. (2013). Input-output relations in biological systems: Measurement, information and the hill equation. Biology Direct, 8, 1–25.
Toda, M. (1967). Vibration of a chain with a non-linear interaction. Journal of the Physical Society of Japan, 22, 431–436.
Warburg, O., Wind, F., & Negelein, E. (1927). The metabolism of tumors in the body. Journal of General Physiology, 8, 519–530.
Avalle, L., Camporeale, A., Morciano, G., et al. (2019). Stat3 localizes to the er, acting as a gatekeeper for er-mitochondrion ca2+ fluxes and apoptotic responses. Cell Death and Differentiation, 26, 932–942.
Berridge, M. J., Bootman, M. D., & Roderick, H. L. (2003). Calcium signalling: Dynamics, homeostasis and remodelling. Nature Reviews. Molecular Cell Biology, 4, 517–529.
Hideshima, T., & Kato, Y. (2006). Oscillatory reaction of catalase wrapped by liposome. Biophysical Chemistry, 124, 100–105.
Savic, R., Luo, L., Eisenberg, A., et al. (2003). Micellar nanocontainers distribute to defined cytoplasmic organelles. Science, 300, 615–618.
Nagahama, K., Sano, Y., Inui, M., et al. (2020). Bioinspired cell nuclear nanotransporters generated by self-assembly of amphiphilic polysaccharide-amino acid derivatives conjugates. Advanced Biosystems, 4, e1900,189.
Acknowledgements
We are deeply grateful to the late Prof. Terukiyo Hanafusa of Hiroshima University for his discussion of DDMC.
Funding
Not applicable
Author information
Authors and Affiliations
Contributions
YO designed and performed the data analysis, and wrote the manuscript. OK conducted the experiments, performed the data analysis, and wrote the manuscript. RJ, TK, MM, JY, Nk, and UE reviewed the manuscript. MO supervised the project.
Corresponding author
Ethics declarations
Informed Consent
Not applicable
Conflict of Interest
The authors declare no competing interests.
Research Involving Humans and Animals Statement
Not applicable
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Klimenko, O.V., Ji, RC., Kobayashi, T. et al. Soliton Dynamics and DDMC/sncRNAs Complex for Epigenetic Change to Normal Cells in TME. BioNanoSci. (2023). https://doi.org/10.1007/s12668-023-01258-z
Accepted:
Published:
DOI: https://doi.org/10.1007/s12668-023-01258-z