Abstract
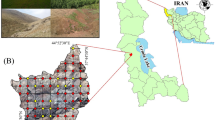
Mapping the spatial distribution of soil classes is useful for proper soil and land-use management. This study investigates the ability of different digital soil mapping (DSM) approaches to predict taxonomic classes up to the family level in the Shahrekord plain of Chaharmahal-Va-Bakhtiari province, Iran. A total of 120 pedons were dug at various map units of a semi-detailed soil map with 750-m intervals. After pedons description, soil samples were taken from different genetic horizons. Based on the pedon descriptions and soil analytical data, pedons were classified up to the family level. Different machine learning techniques such as artificial neural networks, boosted regression tree, random forest and multinomial logistic regression were used to test the predictive power for mapping the soil classes. Overall accuracy (OA), adjusted kappa index and brier scores (BS) were used to determine the accuracy of the prediction. The model with the highest OA (i.e., the highest adjusted kappa) and the lowest BS values was considered as the most accurate model for each soil taxonomic level. Results showed that the different models had the same ability for the prediction of the soil classes across all taxonomic levels while a considerable decreasing trend was observed for their accuracy at subgroup and family levels. The terrain attributes were the most important environmental covariates to predict the soil classes in all taxonomic levels, but they could not display the soil variation entirely. This shows that the unexplained variations are controlled by unobserved variations in environment, which can be due to management over the time. Results suggest that the DSM approaches have not enough prediction accuracy for the soil classes at lower taxonomic levels that focus on the soil properties affecting land use and management. Further studies may still be required to distinguish new environmental covariates and introduce new tools to capture the complex nature of soils.




Similar content being viewed by others
References
Behrens T, Forster H, Scholten T, Steinrucken U, Spies ED, Goldschmitt M (2005) Digital soil mapping using artificial neural networks. J Plant Nutr Soil Sci 168:21–33
Behrens T, Zhu AX, Schmidt K, Scholten T (2010) Multi-scale digital terrain analysis and feature selection for digital soil mapping. Geoderma 155:175–185
Boettinger JL, Ramsey RD, Bodily JM, Cole NJ, Kienast-Brown S, Nield SJ, Saunders AM, Stum AK (2008) Landsat spectral data for digital soil mapping. In: Hartemink AE (ed) Digital soil mapping with limited data. Springer, Berlin, pp 193–203
Breiman L (2001) Random forests. Mach Learn 45:5–32
Brungard CW, Boettinger JL, Duniway MC, Wills SA, EdwardsJr TC (2015) Machine learning for predicting soil classes in three semi-arid landscapes. Geoderma 239–240:68–83
Bui EN, Moran CJ (2003) A strategy to fill gaps in soil survey over large spatial extents: an example from the Murray–Darling basin of Australia. Geoderma 111:21–44
Byrt T, Bishop J, Carling JB (1993) Bias, prevalence and kappa. J Clin Epidemiol 46:423–429
Carre F, McBratney A, Mayr T, Montanarella L (2007) Digital soil assessments: beyond DSM. Geoderma 142:69–79
Congalton R (1991) A review of assessing the accuracy of classifications of remotely sensed data. Remote Sens Environ 37:35–46
Congalton RG, Green K (1998) Assessing the accuracy of remotely sensed data: principles and practices. CRC/Taylor & Francis, Boca Raton
Elith J, Leathwick JR, Hastie T (2008) A working guide to boosted regression trees. J Anim Ecol 77:802–813
Finke PA (2012) On digital soil assessment with models and the pedometrics agenda. Geoderma 171–172:3–15
Friedman JH (2001) Greedy function approximation: a gradient boosting machine. Ann Stat 29:1189–1232
Gee GW, Bauder JW (1986) Particle size analysis. In: Klute A (ed) Methods of soil analysis. Am Soc Agron, Madison, pp 383–411
U.S. Geology Survey (2014) Geology.com/news/2010/free-lansat-images-from-USGS-2. http://glovis.usgs.gov
Hengl T, Toomanian N, Reuter HI, Malakouti MJ (2007) Methods to interpolate soil categorical variables from profile observations: lessons from Iran. Geoderma 140:417–427
Jafari A, Finke PA, Van deWauw J, Ayoubi S, Khademi H (2012) Spatial prediction of USDA-great soil groups in the arid Zarand region, Iran: comparing logistic regression approaches to predict diagnostic horizons and soil types. Eur J Soil Sci 63:284–298
Jafari A, Ayoubi S, Khademi H, Finke PA, Toomanian N (2013) Selection of a taxonomic level for soil mapping using diversity and map purity indices: a case study from an Iranian arid region. Geomorphology 201:86–97
Kempen B, Brus DJ, Heuvelink GBM, Stoorvogel JJ (2009) Updating the 1:50,000 Dutch soil map using legacy soil data: a multinomial logistic regression approach. Geoderma 151:311–326
Kovacevic M, Bajat B, Gajic B (2010) Soil type classification and estimation of soil properties using support vector machines. Geoderma 154:340–347
Kuhn M (2008) Building predictive models in R using the caret package. J Stat Soft 28:1–26
Looney SW (2002) Biostatistical methods. Humana Press, Totowa
Malone BP, Minasny B, McBratney AB (2017) Using digital soil mapping to update, harmonize and disaggregate legacy soil maps. In: Malone BP (ed) Using R for Digital Soil Mapping. Springer, Switzerland, pp 221–230
McBratney AB, Mendonça Santos ML, Minasny B (2003) On digital soil mapping. Geoderma 117:3–52
Mosleh Z, Salehi MH, Jafari A, Esfandiarpoor Borujeni I, Mehnatkesh A (2016) The effectiveness of digital soil mapping to predict soil properties over low-relief areas. Environ Monit Assess 188:1–13
Nelson RE (1982) Carbonate and gypsum. In: Page AL (ed) Methods of soil analysis. American Society of Agronomy, Madison, pp 181–197
Olaya VF (2004) A gentle introduction to Saga GIS. User Manual. The SAGA user group, Gottingen
Pahlavan Rad MR, Toomanian N, Khormali F, Brungard CW, Komaki CB, Bogaert P (2014) Updating soil survey maps using random forest and conditioned Latin hypercube sampling in the loess derived soils of northern Iran. Geoderma 232–234:97–106
Peters J, De Baets B, Verhoest NE, Samson R, Degroeve S, Becker PD, Huybrechts W (2007) Random forests as a tool for ecohydrological distribution modeling. Ecol Model 207:304–318
Phillips JD (2016) Identifying sources of soil landscape complexity with spatial adjacency graphs. Geoderma 267:58–64
Ranst E, Tang H, Groenemmans R, Sinthurahat S (1996) Application of fuzzy logicto land suitability for rubber production in peninsular Thailand. Geoderma 70:1–19
Richardson AJ, Wiegand CL (1977) Distinguishing vegetation from soil background information. Photogramm Eng Remote Sens 43:1541–1552
Rossiter DG (2000) Methodology for soil resource inventories. Soil Science Division, International institute for Aerospace Survey and Earth Science (ITC). 2nd revised version
Rouse JW, Haas RH, Schell JA, Deering DW (1973) Monitoring vegetation systems in the Great plain with ERTS. In: Third ERTS Symposium, NASA, Scientific and Technical Information Office
Schaetzl RJ, Anderson S (2005) Soils genesis and geomorphology. Cambridge University Press, New York
Schoeneberger PJ, Wysocki DA, Benham EC, Soil Survey Staff (2012) Field book for describing and sampling soils, 3nd version. Natural Resources Conservation Service. National Soil Survey Center, Lincoln
Scull P, Franklin J, Chadwick OA (2005) The application of classification tree analysis to soil type prediction in a desert landscape. Ecol Model 181:1–15
Soil Survey Staff (2014) Keys to soil taxonomy. 12th edn. USDA-Natural Resources Conservation Service, Washington, DC
Stum AK, Boettinger JL, White MA, Ramsey RD (2010) Random forests applied as a soil spatial predictive model in Arid Utah. In: Boettinger JL (ed) Digital soil mapping: bridging research, environmental application and operation. Springer, Dordrecht, pp 179–190
Sumner ME, Miller WP (1996) Cation exchange capacity and exchange coefficients. In: Bartels JM (ed) Methods of soil analysis. American Society of Agronomy, Madison, pp 1201–1231
Taghizadeh-Mehrjard R, Minasny B, McBratney AB, Triantafilis J, Sarmadian F, Toomanian N (2012) Digital soil mapping of soil classes using decision trees in central Iran. In: Minasny B (ed) Digital soil assessments and beyond: proceedings of the 5th global workshop on digital soil mapping. CRC Press, Sydney, pp 197–202
Taghizadeh-Mehrjardi R, Nabiollahi K, Minasny B, Triantafilis J (2015) Comparing data mining Classifiers to predict spatial distribution of USDA-family soil groups in Baneh region, Iran. Geoderma 253–254:67–77
R Core Team (2012) R: a language and environment for statistical computing. R foundation for statistical computing, Vienna. http://www.R-project.org/
Viloria JA, Viloria-Botello A, Pineda MC, Valera A (2016) Digital modelling of landscape and soil in a mountainous region: a neuro-fuzzy approach. Geomorphology 253:199–207
Walkley A, Black IA (1934) An examination of Degtjareff method for determining soil organic matter and a proposed modification of chromic acid in soil analysis. Soil Sci Soc Am J 79:459–465
Wu J, Ransom M, Kluitenberg G, Nellis M, Seyler H (2001) Land-use management using a soil survey geographic database for Finney County, Kansas. Soil Sci Soc Am J 65:169–177
Zeraatpisheh M, Ayoubi S, Jafari A, Finke P (2017) Comparing the efficiency of digital and conventional soil mapping to predict soil types in a semi-arid region in Iran. Geomorphology 285:186–204
Zinck JA (1989) Physiography and soils. Lecture notes for soil students. Soil Science Division, Soil survey courses subject matter. Enschede
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mosleh, Z., Salehi, M.H., Jafari, A. et al. Identifying sources of soil classes variations with digital soil mapping approaches in the Shahrekord plain, Iran. Environ Earth Sci 76, 748 (2017). https://doi.org/10.1007/s12665-017-7100-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12665-017-7100-0