Abstract
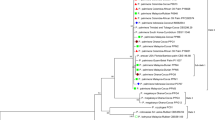
Apple chlorotic leaf spot virus (ACLSV) is one of the latent viruses that occur in apple orchards worldwide but usually without visible symptoms. In 2010–2012, a total of 550 apple leaf samples from 12 different major apple-producing areas in Shaanxi, China, were tested by serological assay for ACLSV; the results revealed an infection level of 51.5 %. Because of the known variability in the putative amino acid sequences of the coat protein (CP), and thus the potential for non-detection by serological assay, the molecular variability of isolates of ACLSV collected in Shaanxi was analyzed using PCR and compared with isolates from the rest of the world. Sequences of 504 nt corresponding to 87 % of the CP gene of 12 isolates were acquired by RT-PCR and deposited in GenBank with the accession numbers KF134387–KF134298. Comparisons of the partial CP gene sequences of these 12 isolates as well as isolates previously reported in the world revealed the pairwise identities ranging from 68.9–99.8 % and 73.8–100 % at the nucleotide and amino acid level, respectively. Phylogenetic analysis based on these nucleotide sequences showed that the 72 isolates deposited in GenBank fell into three groups (P205, B6 and Ta Tao 5 Group). Our 12 ACLSV isolates were separated into the P205 and B6 groups, respectively. Multiple alignment analysis of the amino acid sequences of CP revealed that there was a combination of six amino acids at positions 40, 59, 75, 86, 130 and 184 in isolates from each group that could be used to distinguish among the three groups. Two recombination events were identified from all isolates by recombination analysis, and three ACLSV isolates collected in this study participated in these two events. Our results show that molecular variation was present in isolates of ACLSV collected in Shaanxi province and this may reflect introductions of the virus associated with different sources of germplasm.




Similar content being viewed by others
References
Al Rwahnih, M., Turturo, C., Minafra, A., Saldarelli, P., Myrta, A., Pallás, V., et al. (2004). Molecular variability of Apple chlorotic leaf spot virus in different hosts and geographical regions. Journal of Plant Pathology, 86, 117–122.
Asif, M., Trivedi, P., Solomos, T., & Tucker, M. (2006). Isolation of high-quality RNA from apple (Malus domestica) fruit. Journal of Agricultural and Food Chemistry, 54, 5227–5229.
Candresse, T., Lanneau, M., Revers, F., Grasseau, N., Macquaire, G., German, S., et al. (1995). An immunocapture PCR assay adapted to the detection and the analysis of the molecular variability of apple chlorotic leaf spot virus. Acta Horticulturae, 386, 136–147.
Carstens, E. (2010). Ratification vote on taxonomic proposals to the International Committee on Taxonomy of Viruses (2009). Archives of Virology, 155, 133–146.
Corpet, F. (1988). Multiple sequence alignment with hierarchical clustering. Nucleic Acids Research, 16, 10881–10890.
Desvignes, J., & Boyé, R. (1989). Different diseases caused by the chlorotic leaf spot virus on the fruit trees. Acta Horticulturae, 235, 31–38.
Desvignes, J. C., Boyé, R., Cornaggia, D., Grasseau, N., Hurtt, S., & Waterworth, H. (1999). Virus diseases of fruit trees. (pp. 239–252). Paris, France: Centre Technique Interprofessionnel des Fruits et Légumes (CTIFL).
Dhir, S., Zaidi, A. A., & Hallan, V. (2013). Molecular characterization and recombination analysis of the complete genome of Apple chlorotic leaf spot virus. Journal of Phytopathology, 161, 704–712.
Drummon, A., Oliver, G., & Rambaut, A. (2003). Inference of viral evolutionary rates from molecular sequences. Advances in Parasitology, 54, 331–358.
Dunez, J., Marenaud, C., Delbos, R., & Lansac, M. (1972). Variability of symptoms induced by the apple chlorotic leaf spot (CLSV) – a type of CLSV probably responsible for bark split disease of prune trees. Plant Disease Reporter, 56, 293–295.
Fu, Y.-X., & Li, W.-H. (1993). Statistical tests of neutrality of mutations. Genetics, 133, 693–709.
Gagarinova, A. G., Babu, M., Strömvik, M. V., & Wang, A. (2008). Recombination analysis of Soybean mosaic virus sequences reveals evidence of RNA recombination between distinct pathotypes. Virology Journal, 5, 143.
García-Arenal, F., Fraile, A., & Malpica, J. M. (2001). Variability and genetic structure of plant virus populations. Annual Review of Phytopathology, 39, 157–186.
German, S., Candresse, T., Lanneau, M., Huet, J., Pernollet, J., & Dunez, J. (1990). Nucleotide sequence and genomic organization of apple chlorotic leaf spot closterovirus. Virology, 179, 104–112.
German-Retana, S., Bergey, B., Delbos, R., Candresse, T., & Dunez, J. (1997). Complete nucleotide sequence of the genome of a severe cherry isolate of apple chlorotic leaf spot trichovirus (ACLSV). Archives of Virology, 142, 833–841.
Jelkmann, W., & Kunze, L. (1995). Plum pseudopox in German plum after infection with an isolate of apple chlorotic leaf spot virus causing plum line pattern. Acta Horticulturae, 386, 122–125.
Karan, M., Harding, R. M., & Dale, J. L. (1994). Evidence for two groups of banana bunchy top virus isolates. Journal of General Virology, 75, 3541–3546.
Li, R., Mock, R., Huang, Q., Abad, J., Hartung, J., & Kinard, G. (2008). A reliable and inexpensive method of nucleic acid extraction for the PCR-based detection of diverse plant pathogens. Journal of Virological Methods, 154, 48–55.
Librado, P., & Rozas, J. (2009). DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics, 25, 1451–1452.
Liu, N., Niu, J., & Zhao, Y. (2012). Complete genomic sequence analyses of Apple stem pitting virus isolates from China. Virus Genes, 44, 124–130.
Liu, P., Zhang, L., Zhang, H., Jiao, H., & Wu, Y. (2013). Detection and molecular variability of Apple stem grooving virus in Shaanxi, China. Journal of Phytopathology, 161, 445–449.
Marini, D., Gibson, P., & Scott, S. (2008). The complete nucleotide sequence of an isolate of Apple chlorotic leaf spot virus from peach (Prunus persica (L.) Batch). Archives of Virology, 153, 1003–1005.
Martelli, G., Candresse, T., & Namba, S. (1994). Trichovirus, a new genus of plant viruses. Archives of Virology, 134, 451–455.
Martelli, G. P., Adams, M. J., Kreuze, J. F., & Dolja, V. V. (2007). Family Flexiviridae: a case study in virion and genome plasticity. Annual Review of Phytopathology, 45, 73–100.
Martin, D., & Rybicki, E. (2000). RDP: detection of recombination amongst aligned sequences. Bioinformatics, 16, 562–563.
Martin, D. P., Lemey, P., Lott, M., Moulton, V., Posada, D., & Lefeuvre, P. (2010). RDP3: a flexible and fast computer program for analyzing recombination. Bioinformatics, 26, 2462–2463.
Menzel, W., Jelkmann, W., & Maiss, E. (2002). Detection of four apple viruses by multiplex RT-PCR assays with coamplification of plant mRNA as internal control. Journal of Virological Methods, 99, 81–92.
Muhire, B., Martin, D. P., Brown, J. K., Navas-Castillo, J., Moriones, E., Zerbini, F. M., et al. (2013). A genome-wide pairwise-identity-based proposal for the classification of viruses in the genus Mastrevirus (family Geminiviridae). Archives of Virology, 1–14.
Nemeth, M. (1986). Virus, mycoplasma and rickettsia diseases of fruit trees. Budapest, Hungary: Akademiai Kiado
Niu, F., Pan, S., Wu, Z., Jiang, D., & Li, S. (2012). Complete nucleotide sequences of the genomes of two isolates of apple chlorotic leaf spot virus from peach (Prunus persica) in China. Archives of Virology, 157, 783–786.
Pasquini, G., Faggioli, F., Pilotti, M., Lumia, V., & Barba, M. (1998). Characterization of Apple chlorotic leaf spot virus isolates from Italy. Acta Horticulturae, 472, 195–199.
Peña-Iglesias, A., & Ayuso Gonzalez, P. (1975). Preliminary identification of the viruses producing Spanish apricot pseudopox (viruela) and apricot mosaic diseases. Acta Horticulturae, 44, 255–268.
Ragozzino, A., & Pugliano, G. (1974). La butteratura delle albicocche. Indagini preliminari sulla eziologia. Rivista dell’Ortoflorofrutticoltura Italiana, 58, 136–145.
Revers, F., Le Gall, O., Candresse, T., Le Romancer, M., & Dunez, J. (1996). Frequent occurrence of recombinant potyvirus isolates. Journal of General Virology, 77, 1953–1965.
Rico, P., Ivars, P., Elena, S. F., & Hernández, C. (2006). Insights into the selective pressures restricting Pelargonium flower break virus genome variability: evidence for host adaptation. Journal of Virology, 80, 8124–8132.
Sato, K., Yoshikawa, N., & Takahashi, T. (1993). Complete nucleotide sequence of the genome of an apple isolate of apple chlorotic leaf spot virus. Journal of General Virology, 74, 1927.
Schroeder, M., & Weidemann, H. L. (1989). Simplified application of return gel electrophoresis for the routine detection of potato spindle tuber viroid. EPPO Bulletin, 19, 661–665.
Seo, J.-K., Ohshima, K., Lee, H.-G., So, M., Choi, H.-S., Lee, S.-H., et al. (2009). Molecular variability and genetic structure of the population of Soybean mosaic virus based on the analysis of complete genome sequences. Virology, 393, 91–103.
Tajima, F. (1983). Evolutionary relationship of DNA sequences in finite populations. Genetics, 105, 437–460.
Tajima, F. (1989). Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics, 123, 585–595.
Tamura, K., Peterson, D., Peterson, N., Stecher, G., Nei, M., & Kumar, S. (2011). MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Molecular Biology and Evolution, 28, 2731–2739.
Varsani, A., Shepherd, D. N., Monjane, A. L., Owor, B. E., Erdmann, J. B., Rybicki, E. P., et al. (2008). Recombination, decreased host specificity and increased mobility may have driven the emergence of Maize streak virus as an agricultural pathogen. Journal of General Virology, 89, 2063–2074.
Wang, G., Hong, N., Zhang, Z., Hu, S., & Dong, Y. (1994). Identification of virus species in pears cultivated in northern China. China Fruits, 2, 1–4.
Watterson, G. (1975). On the number of segregating sites in genetical models without recombination. Theoretical Population Biology, 7, 256–276.
Yaegashi, H., Isogai, M., Tajima, H., Sano, T., & Yoshikawa, N. (2007). Combinations of two amino acids (Ala40 and Phe75 or Ser40 and Tyr75) in the coat protein of Apple chlorotic leaf spot virus are crucial for infectivity. Journal of General Virology, 88, 2611–2618.
Yanase, H. (1974). Studies on apple latent viruses in Japan: The association of apple top-working disease with apple latent viruses. Bulletin of the Fruit Tree Research Station. Series C. Morioka. 47-109
Yang, Z., & Bielawski, J. P. (2000). Statistical methods for detecting molecular adaptation. Trends in Ecology & Evolution, 15, 496–503.
Yoshikawa, N., & Takahashi, T. (1988). Properties of RNAs and proteins of apple stem grooving and apple chlorotic leaf spot viruses. Journal of General Virology, 69, 241–245.
Zhang, X., Zhu, H., & Zhang, H. (2005). Present situation and developing thought of apple industry in Shaanxi. Journal of Northwest University(Philosophy and Social Sciences Edition), 35, 36–41.
Acknowledgments
This work was supported by the 111 project (Grant No. B07049) and the National High-tech R&D Program of China (2012AA101504). The authors are grateful to fruit growers for their cooperation during the surveys and to Tao Ye for careful reading of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOC 40 kb)
Rights and permissions
About this article
Cite this article
Liu, P., Li, Z., Song, S. et al. Molecular variability of Apple chlorotic leaf spot virus in Shaanxi, China. Phytoparasitica 42, 445–454 (2014). https://doi.org/10.1007/s12600-013-0381-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12600-013-0381-2