Abstract
Copy number variation (CNV) is an essential genetic driving factor of cancer formation and progression, making intelligent classification based on CNV feasible. However, there are a few challenges in the current machine learning and deep learning methods, such as the design of base classifier combination schemes in ensemble methods and the selection of layers of neural networks, which often result in low accuracy. Therefore, an adaptive bilinear dynamic cascade model (Adap-BDCM) is developed to further enhance the accuracy and applicability of these methods for intelligent classification on CNV datasets. In this model, a feature selection module is introduced to mitigate the interference of redundant information, and a bilinear model based on the gated attention mechanism is proposed to extract more beneficial deep fusion features. Furthermore, an adaptive base classifier selection scheme is designed to overcome the difficulty of manually designing base classifier combinations and enhance the applicability of the model. Lastly, a novel feature fusion scheme with an attribute recall submodule is constructed, effectively avoiding getting stuck in local solutions and missing some valuable information. Numerous experiments have demonstrated that our Adap-BDCM model exhibits optimal performance in cancer classification, stage prediction, and recurrence on CNV datasets. This study can assist physicians in making diagnoses faster and better.
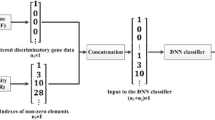
Graphical Abstract












Similar content being viewed by others
Code Availability
The source codes for the Adap-BDCM model and the CNV dataset for cancer classification have been archived in the GitHub repository (https://github.com/junhonga/Adap-BDCM).
References
Jin J, Wu X, Yin J et al (2019) Identification of genetic mutations in cancer: challenge and opportunity in the new era of targeted therapy. Front Onco 9:263. https://doi.org/10.3389/fonc.2019.00263
Poduri A, Evrony GD, Cai X et al (2013) Somatic mutation, genomic variation, and neurological disease. Science 341(6141):1237758. https://doi.org/10.1126/science.1237758
Redon R, Ishikawa S, Fitch KR et al (2006) Global variation in copy number in the human genome. Nature 444(7118):444–454. https://doi.org/10.1038/nature05329
Buchynska LG, Brieiev OV, Iurchenko NP (2019) Assessment of HER-2/neu, c-MYC and CCNE1 gene copy number variations and protein expression in endometrial carcinomas. Exp Oncol 41(2):138–143. https://doi.org/10.32471/exp-oncology.2312-8852.vol-41-no-2.12973
Tian T, Bi H, Liu Y et al (2020) Copy number variation of ubiquitin-specific proteases genes in blood leukocytes and colorectal cancer. Cancer Biol Ther 21(7):637–646. https://doi.org/10.1080/15384047.2020.1750860
Zhang N, Wang M, Zhang P et al (1860) (2016) Classification of cancers based on copy number variation landscapes. Bba-gen Subjects 11:2750–2755. https://doi.org/10.1016/j.bbagen.2016.06.003
Liang Y, Wang H, Yang J et al (2020) A deep learning framework to predict tumor tissue-of-origin based on copy number alteration. Front Bioeng Biotech 8:701. https://doi.org/10.3389/fbioe.2020.00701
Wu Q, Li D (2022) CRIA: an interactive gene selection algorithm for cancers prediction based on copy number variations. Front Plant Sci 13:839044. https://doi.org/10.3389/fpls.2022.839044
Zhou ZH, Feng J (2019) Deep forest. Natl Sci Rev 6(1):74–86. https://doi.org/10.1093/nsr/nwy108
Guo Y, Liu S, Li Z et al (2018) BCDForest: a boosting cascade deep forest model towards the classification of cancer subtypes based on gene expression data. BMC Bioinform 19(5):1–13. https://doi.org/10.1186/s12859-018-2095-4
El-Nabawy A, Belal NA, El-Bendary N (2021) A cascade deep forest model for breast cancer subtype classification using multi-omics data. Mathematics 9(13):1574. https://doi.org/10.3390/math9131574
Zhong L, Meng Q, Chen Y (2021) A cascade flexible neural forest model for cancer subtypes classification on gene expression data. Comput Intel Neurosc 2021:1–11. https://doi.org/10.1155/2021/6480456
Shaaban MA, Hassan YF, Guirguis SK (2022) Deep convolutional forest: a dynamic deep ensemble approach for spam detection in text. Complex Intell Syst 8(6):4897–4909. https://doi.org/10.1007/s40747-022-00741-6
Tenenbaum JB, Freeman WT (2000) Separating style and content with bilinear models. Neural Comput 12(6):1247–1283. https://doi.org/10.1162/089976600300015349
Lin TY, RoyChowdhury A, Maji S (2015) Bilinear CNN models for fine-grained visual recognition. In: Proceedings of the IEEE International Conference on computer vision, pp 1449–1457. https://doi.org/10.1109/ICCV.2015.170
Gao Y, Beijbom O, Zhang N et al (2016) Compact bilinear pooling. In: Proceedings of the IEEE Conference on computer vision and pattern recognition, pp 317–326. https://doi.org/10.1109/CVPR.2016.41
Kim JH, On KW, Lim W et al (2016) Hadamard product for low-rank bilinear pooling. arXiv. https://doi.org/10.48550/arXiv.1610.04325
Li Y, Wang N, Liu J et al (2017) Factorized bilinear models for image recognition. In: Proceedings of the IEEE International Conference on Computer Vision, pp 2079–2087. https://doi.org/10.1109/ICCV.2017.229
Li E, Samat A, Du P et al (2020) Improved bilinear CNN model for remote sensing scene classification. IEEE Geosci Remote Sens Lett 19:1–5. https://doi.org/10.1109/LGRS.2020.3040153
Yu Z, Yu J, Fan J et al (2017) Multi-modal factorized bilinear pooling with co-attention learning for visual question answering. In: Proceedings of the IEEE International Conference on computer vision, pp 1821–1830. https://doi.org/10.1109/ICCV.2017.202
Gao C, Chen Y, Jiang X et al (2023) Bi-STAN: bilinear spatial-temporal attention network for wearable human activity recognition. Int J Mach Learn Cyb 14(7):2545–2561. https://doi.org/10.1007/s13042-023-01781-1
Wang Z, Li R, Wang M et al (2021) GPDBN: deep bilinear network integrating both genomic data and pathological images for breast cancer prognosis prediction. Bioinformatics 37(18):2963–2970. https://doi.org/10.1093/bioinformatics/btab185
Li R, Wu X, Li A et al (2022) HFBSurv: hierarchical multimodal fusion with factorized bilinear models for cancer survival prediction. Bioinformatics 38(9):2587–2594. https://doi.org/10.1093/bioinformatics/btac113
Qiu L, Khormali A, Liu K (2023) Deep biological pathway informed pathology-genomic multimodal survival prediction. arXiv. https://doi.org/10.48550/arXiv.2301.02383
Dudoit S, Fridlyand J (2003) Bagging to improve the accuracy of a clustering procedure. Bioinformatics 19(9):1090–1099. https://doi.org/10.1093/bioinformatics/btg038
Wang A, Liu H, Yang J et al (2022) Ensemble feature selection for stable biomarker identification and cancer classification from microarray expression data. Comput Biol Med 142:105208. https://doi.org/10.1016/j.compbiomed.2021.105208
Elmi J, Eftekhari M, Mehrpooya A et al (2023) A novel framework based on the multi-label classification for dynamic selection of classifiers. Int J Mach Learn Cyb 14(6):2137–2154. https://doi.org/10.1007/s13042-022-01751-z
Hashemi A, Dowlatshahi MB, Nezamabadi-pour H (2022) Ensemble of feature selection algorithms: a multi-criteria decision-making approach. Int J Mach Learn Cyb 13(1):49–69. https://doi.org/10.1007/s13042-021-01347-z
Ferreira AJ, Figueiredo MAT (2012) Boosting algorithms: a review of methods, theory, and applications. In: Zhang C, Ma Y (eds) Ensemble machine learning: methods and applications. Spring, New York, pp 35–85. https://doi.org/10.1007/978-1-4419-9326-7_2
Wang FY, Zhou DW, Ye HJ et al (2022) Foster: Feature boosting and compression for class-incremental learning In: European Conference on Computer Vision, pp 398–414. https://doi.org/10.1007/978-3-031-19806-9_23
Mostafaei SH, Tanha J (2023) OUBoost: boosting based over and under sampling technique for handling imbalanced data. Int J Mach Learn Cyb 14(10):3393–3411. https://doi.org/10.1007/s13042-023-01839-0
Roshan S, Tanha J, Hallaji F et al (2023) IMBoost: a new weighting factor for boosting to improve the classification performance of imbalanced data. Complexity 2023:2176891. https://doi.org/10.1155/2023/2176891
Liong VE, Lu J, Wang G (2013) Face recognition using deep PCA. In: 2013 9th International Conference on Information, Communications & Signal Processing, pp 1–5. https://doi.org/10.1109/ICICS.2013.6782777
Chan TH, Jia K, Gao S et al (2015) PCANet: a simple deep learning baseline for image classification? IEEE T Image Process 4(12):5017–5032. https://doi.org/10.1109/TIP.2015.2475625
Wang W, Dai QY, Li F et al (2021) MLCDForest: multi-label classification with deep forest in disease prediction for long non-coding RNAs. Brief Bioinform 22(3):bbaa104. https://doi.org/10.1093/bib/bbaa104
Peng L, Tan J, Tian X et al (2022) EnANNDeep: an ensemble-based lncRNA–protein interaction prediction framework with adaptive k-nearest neighbor classifier and deep models. Interdiscip Sci 14(1):209–232. https://doi.org/10.1007/s12539-021-00483-y
Muthukrishnan R, Rohini R (2016) LASSO: A feature selection technique in predictive modeling for machine learning. In: 2016 IEEE International Conference on advances in computer applications (ICACA), pp 18–20. https://doi.org/10.1109/ICACA.2016.7887916
Arevalo J, Solorio T, Montes-y-Gómez M et al (2017) Gated multimodal units for information fusion. arXiv. https://doi.org/10.48550/arXiv.1702.01992
Zhu T, Lin Y, Liu Y (2017) Synthetic minority oversampling technique for multiclass imbalance problems. Pattern Recogn 72:327–340. https://doi.org/10.1016/j.patcog.2017.07.024
Cerami E, Gao J, Dogrusoz U et al (2012) The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov 2(5):401–404. https://doi.org/10.1158/2159-8290.CD-12-0095
Gao JJ, Aksoy BA, Dogrusoz U et al (2013) Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal 6(269):l1. https://doi.org/10.1126/scisignal.2004088
Mermel CH, Schumacher SE, Hill B et al (2011) GISTIC2. 0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers. Genome Biol 12:1–14. https://doi.org/10.1186/gb-2011-12-4-r41
Ciriello G, Miller ML, Aksoy BA et al (2013) Emerging landscape of oncogenic signatures across human cancers. Nat Genet 45(10):1127–1133. https://doi.org/10.1038/ng.2762
Li J, Cheng K, Wang S et al (2017) Feature selection: a data perspective. ACM Comput Surv (CSUR) 50(6):1–45. https://doi.org/10.1145/3136625
Fisher A, Rudin C, Dominici F (2019) All models are wrong, but many are useful: learning a variable’s importance by studying an entire class of prediction models simultaneously. J Mach Learn Res 20(177):1–81. https://doi.org/10.1016/j.petsci.2022.09.003
Pan X, Hu XH, Zhang YH et al (2019) Identification of the copy number variant biomarkers for breast cancer subtypes. Mol Genet Genom 294:95–110. https://doi.org/10.1007/s00438-018-1488-4
Huang T, Chen C, Du J et al (2023) A tRF-5a fragment that regulates radiation resistance of colorectal cancer cells by targeting MKNK1. J Cell Mol Med 27(24):4021–4033. https://doi.org/10.1111/jcmm.17982
Fernandez-Rozadilla C, Cazier JB, Tomlinson IP et al (2013) A colorectal cancer genome-wide association study in a Spanish cohort identifies two variants associated with colorectal cancer risk at 1p33 and 8p12. BMC Genom 14:1–11. https://doi.org/10.1186/1471-2164-14-55
Kim S, Kim JM, Lee HJ et al (2020) Alteration of CYP4A11 expression in renal cell carcinoma: diagnostic and prognostic implications. J Cancer 11(6):1478. https://doi.org/10.7150/jca.36438
Lee K, Jeong H, Lee S et al (2019) CPEM: Accurate cancer type classification based on somatic alterations using an ensemble of a random forest and a deep neural network. Sci Rep-UK 9(1):16927. https://doi.org/10.1038/s41598-019-53034-3
Shen J, Shi J, Luo J et al (2022) Deep learning approach for cancer subtype classification using high-dimensional gene expression data. BMC Bioinform 23(1):1–17. https://doi.org/10.1186/s12859-022-04980-9
Acknowledgements
In this paper, the work is supported by the Fundamental Research Program of Shanxi Province (General program) (Grant No. 202303021211082, 202303021211025).
Author information
Authors and Affiliations
Contributions
Concept and design: Liancheng Jiang and Liye Jia, data collection and analysis: Yizhen Wang; drafting of the article: Liancheng Jiang, Liye Jia and Junhong Yue; critical revision of the article for important content: Liancheng Jiang, Liye Jia, Yizhen Wang, Yongfei Wu, Junhong Yue, All the authors approved the final article.
Corresponding author
Ethics declarations
Competing Interest
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Ethics statement
Not applicable.
Informed consent
Not applicable.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Jiang, L., Jia, L., Wang, Y. et al. Adap-BDCM: Adaptive Bilinear Dynamic Cascade Model for Classification Tasks on CNV Datasets. Interdiscip Sci Comput Life Sci (2024). https://doi.org/10.1007/s12539-024-00635-w
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12539-024-00635-w