Abstract
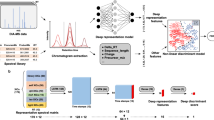
Mass spectrometry is crucial in proteomics analysis, particularly using Data Independent Acquisition (DIA) for reliable and reproducible mass spectrometry data acquisition, enabling broad mass-to-charge ratio coverage and high throughput. DIA-NN, a prominent deep learning software in DIA proteome analysis, generates peptide results but may include low-confidence peptides. Conventionally, biologists have to manually screen peptide fragment ion chromatogram peaks (XIC) for identifying high-confidence peptides, a time-consuming and subjective process prone to variability. In this study, we introduce SeFilter-DIA, a deep learning algorithm, aiming at automating the identification of high-confidence peptides. Leveraging compressed excitation neural network and residual network models, SeFilter-DIA extracts XIC features and effectively discerns between high and low-confidence peptides. Evaluation of the benchmark datasets demonstrates SeFilter-DIA achieving 99.6% AUC on the test set and 97% for other performance indicators. Furthermore, SeFilter-DIA is applicable for screening peptides with phosphorylation modifications. These results demonstrate the potential of SeFilter-DIA to replace manual screening, providing an efficient and objective approach for high-confidence peptide identification while mitigating associated limitations.
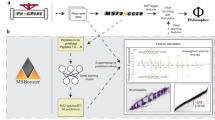
Graphical Abstract









Similar content being viewed by others
References
Zhang Y, Fonslow BR, Shan B et al (2013) Protein analysis by shotgun/bottom-up proteomics. Chem Rev 113:2343–2394. https://doi.org/10.1021/cr3003533
Gillet LC, Navarro P, Tate S et al (2012) Targeted data extraction of the ms/ms spectra generated by data-independent acquisition: a new concept for consistent and accurate proteome analysis. Mol Cell Proteom 11(O111):016717. https://doi.org/10.1074/mcp.O111.016717
Li X, Zhong C, Wu R et al (2021) RIP1-dependent linear and nonlinear recruitments of caspase-8 and RIP3 respectively to necrosome specify distinct cell death outcomes. Protein Cell 12:858–876. https://doi.org/10.1007/s13238-020-00810-x
Li X, Zhong C, Yin Z et al (2020) Data-driven modeling identifies TIRAP-independent MyD88 activation complex and myddosome assembly strategy in LPS/TLR4 signaling. Int J Mol Sci 21:3061. https://doi.org/10.3390/ijms21093061
Röst HL, Rosenberger G, Navarro P et al (2014) OpenSWATH enables automated, targeted analysis of data-independent acquisition MS data. Nat Biotechnol 32:219–223. https://doi.org/10.1038/nbt.2841
Keller A, Bader SL, Shteynberg D et al (2015) Automated validation of results and removal of fragment ion interferences in targeted analysis of data-independent acquisition Mass Spectrometry (MS) using SWATHProphet. Mol Cell Proteom 14:1411–1418. https://doi.org/10.1074/mcp.O114.044917
Peckner R, Myers SA, Jacome ASV et al (2018) Specter: linear deconvolution for targeted analysis of data-independent acquisition mass spectrometry proteomics. Nat Methods 15:371–378. https://doi.org/10.1038/nmeth.4643
Tsou C, Avtonomov D, Larsen B et al (2015) DIA-Umpire: comprehensive computational framework for data-independent acquisition proteomics. Nat Methods 12:258–264. https://doi.org/10.1038/nmeth.3255
Li Y, Zhong C, Xu X et al (2015) Group-DIA: analyzing multiple data-independent acquisition mass spectrometry data files. Nat Methods 12:1105–1106. https://doi.org/10.1038/nmeth.3593
Meyer JG, Mukkamalla S, Steen H et al (2017) PIQED: automated identification and quantification of protein modifications from DIA-MS data. Nat Methods 14:646–647. https://doi.org/10.1038/nmeth.4334
Bruderer R, Bernhardt OM, Gandhi T et al (2015) Extending the limits of quantitative proteome profiling with data-independent acquisition and application to acetaminophen-treated three-dimensional liver microtissues. Mol Cell Proteom 14:1400–1410. https://doi.org/10.1074/mcp.M114.044305
Ting YS, Egertson JD, Bollinger JG et al (2017) PECAN: library-free peptide detection for data-independent acquisition tandem mass spectrometry data. Nat Methods 14:903–908. https://doi.org/10.1038/nmeth.4390
Sinitcyn P, Hamzeiy H, Salinas Soto F et al (2021) MaxDIA enables library-based and library-free data-independent acquisition proteomics. Nat Biotechnol 39:1563–1573. https://doi.org/10.1038/s41587-021-00968-7
Qian X, Qiu Y, He Q et al (2021) A review of methods for sleep arousal detection using polysomnographic signals. Brain Sci 11:1274. https://doi.org/10.3390/brainsci11101274
Hu H, Feng Z, Lin H et al (2023) Modeling and analyzing single-cell multimodal data with deep parametric inference. Brief Bioinform 24:bbad005. https://doi.org/10.1093/bib/bbad005
Wang W, Zhang L, Sun J et al (2022) Predicting the potential human lncRNA–miRNA interactions based on graph convolution network with conditional random field. Brief Bioinform 23:bbac463. https://doi.org/10.1093/bib/bbac463
Zhao J, Sun J, Shuai SC et al (2023) Predicting potential interactions between lncRNAs and proteins via combined graph auto-encoder methods. Brief Bioinform 24:bbac527. https://doi.org/10.1093/bib/bbac527
Zhong J, Song Z, Zhang L et al (2022) Assembly of guanine crystals as a low-polarizing broadband multilayer reflector in a spider, phoroncidia rubroargentea. ACS Appl Mater Interfaces 14:32982–32993. https://doi.org/10.1021/acsami.2c09546
Chen X, Zhu R, Zhong J et al (2022) Mosaic composition of RIP1–RIP3 signalling hub and its role in regulating cell death. Nat Cell Biol 24:471–482. https://doi.org/10.1038/s41556-022-00854-7
Wang J, Chen F, Ma Y et al (2023) XBound-former: toward cross-scale boundary modeling in transformers. IEEE Trans Med Imaging 42:1735–1745. https://doi.org/10.1109/tmi.2023.3236037
Gessulat S, Schmidt T, Zolg DP et al (2019) Prosit: proteome-wide prediction of peptide tandem mass spectra by deep learning. Nat Methods 16:509–518. https://doi.org/10.1038/s41592-019-0426-7
Tran NH, Qiao R, Xin L et al (2019) Deep learning enables de novo peptide sequencing from data-independent-acquisition mass spectrometry. Nat Methods 16:63–66. https://doi.org/10.1038/s41592-018-0260-3
Tiwary S, Levy R, Gutenbrunner P et al (2019) High-quality MS/MS spectrum prediction for data-dependent and data-independent acquisition data analysis. Nat Methods 16:519–525. https://doi.org/10.1038/s41592-019-0427-6
Zhou X, Zeng W, Chi H et al (2017) pDeep: predicting MS/MS spectra of peptides with deep learning. Anal Chem 89:12690–12697. https://doi.org/10.1021/acs.analchem.7b02566
Yang Y, Liu X, Shen C et al (2020) In silico spectral libraries by deep learning facilitate data-independent acquisition proteomics. Nat Commun 11:146. https://doi.org/10.1038/s41467-019-13866-z
He Q, Zhong C, Li X et al (2023) Dear-DIAXMBD: deep autoencoder enables deconvolution of data-independent acquisition proteomics. Research 6:0179. https://doi.org/10.34133/research.0179
Gao M, Yang W, Li C et al (2021) Deep representation features from DreamDIAXMBD improve the analysis of data-independent acquisition proteomics. Commun Biol 4:1190. https://doi.org/10.1038/s42003-021-02726-6
Demichev V, Messner CB, Vernardis SI et al (2020) DIA-NN: neural networks and interference correction enable deep proteome coverage in high throughput. Nat Methods 17:41–44. https://doi.org/10.1038/s41592-019-0638-x
MacLean B, Tomazela DM, Shulman N et al (2010) Skyline: an open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics 26:966–968. https://doi.org/10.1093/bioinformatics/btq054
Sturm M, Kohlbacher O (2009) TOPPView: an open-source viewer for mass spectrometry data. J Proteome Res 8:3760–3763. https://doi.org/10.1021/pr900171m
Li Y, He Q, Guo H et al (2022) MSSort-DIAXMBD: A deep learning classification tool of the peptide precursors quantified by OpenSWATH. J Proteomics 259:104542. https://doi.org/10.1016/j.jprot.2022.104542
Gupta S, Sing J, Mahmoodi A et al (2020) DrawAlignR: an interactive tool for across run chromatogram alignment visualization. Proteomics 20:1900353. https://doi.org/10.1002/pmic.201900353
Tatjana V, Domitille S, Jean-Charles S (2021) Paraquat-induced cholesterol biosynthesis proteins dysregulation in human brain microvascular endothelial cells. Sci Rep 11:18137. https://doi.org/10.1038/s41598-021-97175-w
Midha MK, Kusebauch U, Shteynberg D et al (2020) A comprehensive spectral assay library to quantify the Escherichia coli proteome by DIA/SWATH-MS. Sci Data 7:389. https://doi.org/10.1038/s41597-020-00724-7
Navarro P, Kuharev J, Gillet LC et al (2016) A multicenter study benchmarks software tools for label-free proteome quantification. Nat Biotechnol 34:1130–1136. https://doi.org/10.1038/nbt.3685
Muntel J, Kirkpatrick J, Bruderer R et al (2019) Comparison of protein quantification in a complex background by DIA and TMT workflows with fixed instrument time. J Proteome Res 18:1340–1351. https://doi.org/10.1021/acs.jproteome.8b00898
Chambers MC, Maclean B, Burke R et al (2012) A cross-platform toolkit for mass spectrometry and proteomics. Nat Biotechnol 30:918–920. https://doi.org/10.1038/nbt.2377
Reiter L, Rinner O, Picotti P et al (2011) mProphet: automated data processing and statistical validation for large-scale SRM experiments. Nat Methods 8:430–435. https://doi.org/10.1038/nmeth.1584
Röst HL, Liu Y, D’Agostino G et al (2016) TRIC: an automated alignment strategy for reproducible protein quantification in targeted proteomics. Nat Methods 13:777–783. https://doi.org/10.1038/nmeth.3954
Eng JK, Jahan TA, Hoopmann MR (2013) Comet: An open-source MS/MS sequence database search tool. Proteomics 13:22–24. https://doi.org/10.1002/pmic.201200439
Craig R, Beavis RC (2004) TANDEM: matching proteins with tandem mass spectra. Bioinformatics 20:1466–1467. https://doi.org/10.1093/bioinformatics/bth092
Keller A, Nesvizhskii AI, Kolker E et al (2002) Empirical statistical model to estimate the accuracy of peptide identifications made by MS/MS and database search. Anal Chem 74:5383–5392. https://doi.org/10.1021/ac025747h
Shteynberg D, Deutsch EW, Lam H et al (2011) iProphet: multi-level integrative analysis of shotgun proteomic data improves peptide and protein identification rates and error estimates. Mol Cell Proteom. https://doi.org/10.1074/mcp.M111.007690
Lam H, Deutsch EW, Eddes JS et al (2007) Development and validation of a spectral library searching method for peptide identification from MS/MS. Proteom 7:655–667. https://doi.org/10.1002/pmic.200600625
Shi X, Chen Z, Wang H et al (2015) Convolutional LSTM Network: a machine learning approach for precipitation nowcasting. Proceed Int Conf Neural Inform Process Syst 1:802–810. https://doi.org/10.5555/2969239.2969329
He K, Zhang X, Ren S et al (2016) Deep Residual Learning for Image Recognition. IEEE Conference on Computer Vision and Pattern Recognition (CVPR) 2016:770–778. https://doi.org/10.1109/CVPR.2016.90
Hu J, Shen L, Albanie S et al (2020) Squeeze-and-Excitation Networks. IEEE Trans Pattern Anal Mach Intell 42:2011–2023. https://doi.org/10.1109/TPAMI.2019.2913372
Bekker-Jensen DB, Bernhardt OM, Hogrebe A et al (2020) Rapid and site-specific deep phosphoproteome profiling by data-independent acquisition without the need for spectral libraries. Nat Commun 11:787. https://doi.org/10.1038/s41467-020-14609-1
Zhou Q, Meng Q, Tan X et al (2021) Protein phosphorylation changes during systemic acquired resistance in Arabidopsis thaliana. Front Plant Sci. https://doi.org/10.3389/fpls.2021.748287
Li X, Zhang P, Yin Z et al (2022) Caspase-1 and gasdermin d afford the optimal targets with distinct switching strategies in NLRP1b inflammasome-induced cell death. Research 2022:9838341. https://doi.org/10.34133/2022/9838341
Xu F, Miao D, Li W et al (2023) Specificity and competition of mRNAs dominate droplet pattern in protein phase separation. Phys Rev Res 5:023159. https://doi.org/10.1103/PhysRevResearch.5.023159
Funding
This work is supported by the Ministry of Science and Technology of the People's Republic of China (STI2030-Major Projects2021ZD0201900), the National Natural Science Foundation of China under Grant 12090052 and 11874310, the Natural Science Foundation of Fujian Province of China (Grant No. 2023J05002), and the Fundamental Research Funds for the Central Universities (Grant No. 20720230017).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declare that they have no conflicts of interest.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
He, Q., Guo, H., Li, Y. et al. SeFilter-DIA: Squeeze-and-Excitation Network for Filtering High-Confidence Peptides of Data-Independent Acquisition Proteomics. Interdiscip Sci Comput Life Sci (2024). https://doi.org/10.1007/s12539-024-00611-4
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12539-024-00611-4