Abstract
Computational approaches employed for predicting potential microbe-disease associations often rely on similarity information between microbes and diseases. Therefore, it is important to obtain reliable similarity information by integrating multiple types of similarity information. However, existing similarity fusion methods do not consider multi-order fusion of similarity networks. To address this problem, a novel method of linear neighborhood label propagation with multi-order similarity fusion learning (MOSFL-LNP) is proposed to predict potential microbe-disease associations. Multi-order fusion learning comprises two parts: low-order global learning and high-order feature learning. Low-order global learning is used to obtain common latent features from multiple similarity sources. High-order feature learning relies on the interactions between neighboring nodes to identify high-order similarities and learn deeper interactive network structures. Coefficients are assigned to different high-order feature learning modules to balance the similarities learned from different orders and enhance the robustness of the fusion network. Overall, by combining low-order global learning with high-order feature learning, multi-order fusion learning can capture both the shared and unique features of different similarity networks, leading to more accurate predictions of microbe-disease associations. In comparison to six other advanced methods, MOSFL-LNP exhibits superior prediction performance in the leave-one-out cross-validation and 5-fold validation frameworks. In the case study, the predicted 10 microbes associated with asthma and type 1 diabetes have an accuracy rate of up to 90% and 100%, respectively.
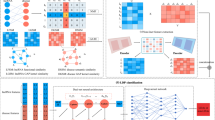
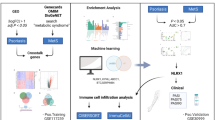
Graphic Abstract











Similar content being viewed by others
Data availability
All information of these data was collected from the public database HMDAD (http://www.cuilab.cn/hmdad) and Disbiome database (https://disbiome.ugent.be/home). The data and source code can be freely downloaded from: https://github.com/RuiBingo/MOSFL-LNP.
References
Morgan XC, Segata N, Huttenhower C (2013) Biodiversity and functional genomics in the human microbiome. Trends Genet 29(1):51–58. https://doi.org/10.1016/j.tig.2012.09.005
Ma W, Zhang L, Zeng P et al (2017) An analysis of human microbe-disease associations. Brief Bioinform 18(1):85–97. https://doi.org/10.1093/bib/bbw005
Puschhof J, Pleguezuelos-Manzano C, Clevers H (2021) Organoids and organs-on-chips: Insights into human gut-microbe interactions. Cell Host Microbe 29(6):867–878. https://doi.org/10.1016/j.chom.2021.04.002
Rook G, Bäckhed F, Levin BR et al (2017) Evolution, human-microbe interactions, and life history plasticity. Lancet 390(10093):521–530. https://doi.org/10.1016/S0140-6736(17)30566-4
Dedrick S, Sundaresh B, Huang Q et al (2020) The role of gut microbiota and environmental factors in type 1 diabetes pathogenesis. Front Endocrinol 11:78. https://doi.org/10.3389/fendo.2020.00078
Zhao Y, Wang C-C, Chen X (2021) Microbes and complex diseases: from experimental results to computational models. Brief Bioinform 22(3):158. https://doi.org/10.1093/bib/bbaa158
Chen X, Huang Y-A, You Z-H et al (2018) A novel approach based on katz measure to predict associations of human microbiota with non-infectious diseases. Bioinformatics 34(8):1440–1440. https://doi.org/10.1093/bioinformatics/btx773
Huang Z-A, Chen X, Zhu Z et al (2017) Pbhmda: path-based human microbe-disease association prediction. Front Microbiol 8:233. https://doi.org/10.3389/fmicb.2017.00233
Shokri Garjan H, Omidi Y, Poursheikhali Asghari M et al (2023) In-silico computational approaches to study microbiota impacts on diseases and pharmacotherapy. Gut Pathog 15(1):10. https://doi.org/10.1186/s13099-023-00535-2
Long Y, Luo J (2019) Wmghmda: a novel weighted meta-graph-based model for predicting human microbe-disease association on heterogeneous information network. BMC Bioinform 20(1):541. https://doi.org/10.1186/s12859-019-3066-0
Wen Z, Yan C, Duan G et al (2021) A survey on predicting microbe-disease associations: biological data and computational methods. Brief Bioinform 22(3):157. https://doi.org/10.1093/bib/bbaa157
Shen Z, Jiang Z, Bao W (2017) Cmfhmda: Collaborative matrix factorization for human microbe-disease association prediction, vol 10362. Springer, New York 261–269. https://doi.org/10.1007/978-3-319-63312-1_24
He B-S, Peng L-H, Li Z (2018) Human microbe-disease association prediction with graph regularized non-negative matrix factorization. Front Microbiol 9:2560. https://doi.org/10.3389/fmicb.2018.02560
Yang X, Kuang L, Chen Z et al (2021) Multi-similarities bilinear matrix factorization-based method for predicting human microbe-disease associations. Front Genet 12:754425. https://doi.org/10.3389/fgene.2021.754425
Xu D, Xu H, Zhang Y et al (2022) Novel collaborative weighted non-negative matrix factorization improves prediction of disease-associated human microbes. Front Microbiol 13:834982. https://doi.org/10.3389/fmicb.2022.834982
Wang L, Tan Y, Yang X et al (2022) Review on predicting pairwise relationships between human microbes, drugs and diseases: from biological data to computational models. Brief Bioinform 23(3):080. https://doi.org/10.1093/bib/bbac080
Luo J, Long Y (2018) Ntshmda: prediction of human microbe-disease association based on random walk by integrating network topological similarity. IEEE ACM Trans Comput Biol Bioinform 17(4):1341–1351. https://doi.org/10.1109/TCBB.2018.2883041
Yan C, Duan G, Wu F-X et al (2019) Brwmda: predicting microbe-disease associations based on similarities and bi-random walk on disease and microbe networks. IEEE ACM Trans Comput Biol Bioinform 17(5):1595–1604. https://doi.org/10.1109/TCBB.2019.2907626
Chen Q, Lai D, Lan W et al (2019) Ildmsf: inferring associations between long non-coding RNA and disease based on multi-similarity fusion. IEEE ACM Trans Comput Biol Bioinform 18(3):1106–1112. https://doi.org/10.1109/TCBB.2019.2936476
Jiang L, Ding Y, Tang J et al (2018) Mda-skf: similarity kernel fusion for accurately discovering miRNA-disease association. Front Genet 9:618. https://doi.org/10.3389/fgene.2018.00618
Xie G-B, Chen R-B, Lin Z-Y et al (2023) Predicting lncrna-disease associations based on combining selective similarity matrix fusion and bidirectional linear neighborhood label propagation. Brief Bioinform 24(1):595. https://doi.org/10.1093/bib/bbac595
Yin M-M, Liu J-X, Gao Y-L et al (2020) Ncplp: a novel approach for predicting microbe-associated diseases with network consistency projection and label propagation. IEEE Trans Cybern 52(6):5079–5087. https://doi.org/10.1109/TCYB.2020.3026652
Liu J-X, Yin M-M, Gao Y-L et al (2022) Msf-lrr: multi-similarity information fusion through low-rank representation to predict disease-associated microbes. IEEE ACM Trans Comput Biol Bioinform 20(1):534–543. https://doi.org/10.1109/TCBB.2022.3146176
Janssens Y, Nielandt J, Bronselaer A et al (2018) Disbiome database: linking the microbiome to disease. BMC Microbiol 18:50. https://doi.org/10.1186/s12866-018-1197-5
Van Laarhoven T, Nabuurs SB, Marchiori E (2011) Gaussian interaction profile kernels for predicting drug-target interaction. Bioinformatics 27(21):3036–3043. https://doi.org/10.1093/bioinformatics/btr500
Kamneva OK (2017) Genome composition and phylogeny of microbes predict their co-occurrence in the environment. PLOS Comput Biol 13(2):1005366. https://doi.org/10.1371/journal.pcbi.1005366
Zhang W, Qu Q, Zhang Y et al (2018) The linear neighborhood propagation method for predicting long non-coding rna-protein interactions. Neurocomputing 273:526–534. https://doi.org/10.1016/j.neucom.2017.07.065
Wang F, Zhang C (2007) Label propagation through linear neighborhoods. IEEE Trans Knowl Data Eng 20(1):55–67. https://doi.org/10.1109/TKDE.2007.190672
Long Y, Luo J, Zhang Y et al (2021) Predicting human microbe-disease associations via graph attention networks with inductive matrix completion. Brief Bioinform 22(3):146. https://doi.org/10.1093/bib/bbaa146
Xie G, Meng T, Luo Y et al (2019) Skf-lda: similarity kernel fusion for predicting lncrna-disease association. Mol Ther Nucleic 18:45–55. https://doi.org/10.1016/j.omtn.2019.07.022
Liu H, Bing P, Zhang M et al (2023) Mnnmda: predicting human microbe-disease association via a method to minimize matrix nuclear norm. Comput Struct Biotechnol J 21:1414–1423. https://doi.org/10.1016/j.csbj.2022.12.053
Wang F, Huang Z-A, Chen X et al (2017) Lrlshmda: laplacian regularized least squares for human microbe-disease association prediction. Sci Rep 7(1):7601. https://doi.org/10.1038/s41598-017-08127-2
Zou S, Zhang J, Zhang Z (2017) A novel approach for predicting microbe-disease associations by bi-random walk on the heterogeneous network. PLoS One 12(9):0184394. https://doi.org/10.1371/journal.pone.0184394
Maahs DM, West NA, Lawrence JM, Mayer-Davis EJ (2010) Epidemiology of type 1 diabetes. Endocrinol Metab Clin North Am 39(3):481–497. https://doi.org/10.1016/j.ecl.2010.05.011
Gillespie KM (2006) Type 1 diabetes: pathogenesis and prevention. Cmaj 175(2):165–170. https://doi.org/10.1503/cmaj.060244
Acharjee S, Ghosh B, Al-Dhubiab BE, Nair AB (2013) Understanding type 1 diabetes: etiology and models. Can J Diabetes 37(4):269–276. https://doi.org/10.1016/j.jcjd.2013.05.001
Lu X, Zhao C (2020) Exercise and type 1 diabetes. Adv Exp Med Biol 1228:107–121. https://doi.org/10.1007/978-981-15-1792-1_7
Vaarala O (2013) Human intestinal microbiota and type 1 diabetes. Curr Diabetes Rep 13:601–607. https://doi.org/10.1007/s11892-013-0409-5
Demirci M, Tokman HB, Taner Z et al (2020) Bacteroidetes and firmicutes levels in gut microbiota and effects of hosts tlr2/tlr4 gene expression levels in adult type 1 diabetes patients in istanbul, turkey. J. Diabetes Complicat 34(2):107449. https://doi.org/10.1016/j.jdiacomp.2019.107449
De Groot P, Nikolic T, Pellegrini S et al (2021) Faecal microbiota transplantation halts progression of human new-onset type 1 diabetes in a randomised controlled trial. Gut 70(1):92–105. https://doi.org/10.1136/gutjnl-2020-322630
Gans MD, Gavrilova T (2020) Understanding the immunology of asthma: pathophysiology, biomarkers, and treatments for asthma endotypes. Paediatr Respir Rev 36:118–127. https://doi.org/10.1016/j.prrv.2019.08.002
Ntontsi P, Photiades A, Zervas E et al (2021) Genetics and epigenetics in asthma. Int J Mol Sci 22(5):2412. https://doi.org/10.3390/ijms22052412
Ver Heul A, Planer J, Kau AL (2019) The human microbiota and asthma. Clin Rev Allergy IMMU 57(3):350–363. https://doi.org/10.1007/s12016-018-8719-7
Chen Y, Zhan X, Wang D (2022) Association between helicobacter pylori and risk of childhood asthma: a meta-analysis of 18 observational studies. J Asthma 59(5):890–900. https://doi.org/10.1080/02770903.2021.1892752
Guo M-Y, Chen H-K, Ying H-Z et al (2021) The role of respiratory flora in the pathogenesis of chronic respiratory diseases. BioMed Res Int 2021:6431862. https://doi.org/10.1155/2021/6431862
Aydin M, Weisser C, Rué O, Mariadassou M et al (2021) The rhinobiome of exacerbated wheezers and asthmatics: Insights from a german pediatric exacerbation network. Front Allergy 2:667562. https://doi.org/10.3389/falgy.2021.667562
Acknowledgements
This work is supported by the National Natural Science Foundation of China (62002070, 82001331) and the Science and Technology Plan Project of Guangzhou City (202102021236).
Author information
Authors and Affiliations
Contributions
Conceptualization: [Rui-Bin Chen, Guo-Bo Xie]; methodology: [Rui-Bin Chen, Guo-Bo Xie]; formal analysis and investigation: [Rui-Bin Chen, Zhi- Yi Lin, Guo-Sheng Gu]; validation: [Rui-Bin Chen]; writing - original draft preparation: [Rui-Bin Chen, Zhi-Yi Lin, Guo-Sheng Gu]; writing -review & editing: [Rui-Bin Chen, Guo-Bo Xie, Zhi-Yi Lin, Guo-Sheng Gu, Yi Yu, Jun-Rui Yu, Zhen-Guo Liu]; funding acquisition: [Guo-Bo Xie, Zhi-Yi Lin, Zhen-Guo Liu]; resources: [Yi Yu, Jun-Rui Yu, Zhen-Guo Liu].
Corresponding authors
Ethics declarations
Conflict of Interest
The authors have no relevant financial or non-financial interests to disclose.This paper does not contain any studies with human participants or animals performed by any of the authors.
Consent to participate
All listed authors have read the submitted manuscript and agree to its submission.
Consent to publish
All listed authors consent for this publication.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chen, R., Xie, G., Lin, Z. et al. Predicting Microbe-Disease Associations Based on a Linear Neighborhood Label Propagation Method with Multi-order Similarity Fusion Learning. Interdiscip Sci Comput Life Sci (2024). https://doi.org/10.1007/s12539-024-00607-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12539-024-00607-0