Abstract
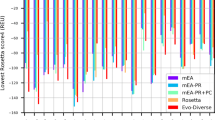
The breakthrough of AlphaFold2 and the publication of AlphaFold DB represent a significant advance in the field of predicting static protein structures. However, AlphaFold2 models tend to represent a single static structure, and multiple-conformation prediction remains a challenge. In this work, we proposed a method named MultiSFold, which uses a distance-based multi-objective evolutionary algorithm to predict multiple conformations. To begin, multiple energy landscapes are constructed using different competing constraints generated by deep learning. Subsequently, an iterative modal exploration and exploitation strategy is designed to sample conformations, incorporating multi-objective optimization, geometric optimization and structural similarity clustering. Finally, the final population is generated using a loop-specific sampling strategy to adjust the spatial orientations. MultiSFold was evaluated against state-of-the-art methods using a benchmark set containing 80 protein targets, each characterized by two representative conformational states. Based on the proposed metric, MultiSFold achieves a remarkable success ratio of 56.25% in predicting multiple conformations, while AlphaFold2 only achieves 10.00%, which may indicate that conformational sampling combined with knowledge gained through deep learning has the potential to generate conformations spanning the range between different conformational states. In addition, MultiSFold was tested on 244 human proteins with low structural accuracy in AlphaFold DB to test whether it could further improve the accuracy of static structures. The experimental results demonstrate the performance of MultiSFold, with a TM-score better than that of AlphaFold2 by 2.97% and RoseTTAFold by 7.72%. The online server is at http://zhanglab-bioinf.com/MultiSFold.
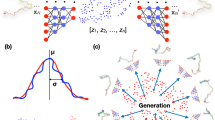
Graphical Abstract






Similar content being viewed by others
Data Availability
All data needed to evaluate the conclusions are present in the paper and the Supplementary Materials. The online server at http://zhanglab-bioinf.com/MultiSFold.
References
Jumper J, Evans R, Pritzel A et al (2021) Highly accurate protein structure prediction with AlphaFold. Nature 596:583–589. https://doi.org/10.1038/s41586-021-03819-2
Subramaniam S, Kleywegt GJ (2022) A paradigm shift in structural biology. Nat Methods 19:20–23. https://doi.org/10.1038/s41592-021-01361-7
Jones DT, Thornton JM (2022) The impact of AlphaFold2 one year on. Nat Methods 19:15–20. https://doi.org/10.1038/s41592-021-01365-3
Varadi M, Anyango S, Deshpande M et al (2022) AlphaFold protein structure database: massively expanding the structural coverage of protein-sequence space with high-accuracy models. Nucleic Acids Res 50:D439–D444. https://doi.org/10.1093/nar/gkab1061
Henzler-Wildman KA, Thai V, Lei M et al (2007) Intrinsic motions along an enzymatic reaction trajectory. Nature 450:838–844. https://doi.org/10.1038/nature06410
Greener JG, Filippis I, Sternberg MJE (2017) Predicting protein dynamics and allostery using multi-protein atomic distance constraints. Structure 25:546–558. https://doi.org/10.1016/j.str.2017.01.008
Tunyasuvunakool K, Adler J, Wu Z et al (2021) Highly accurate protein structure prediction for the human proteome. Nature 596:590–596. https://doi.org/10.1038/s41586-021-03828-1
Thornton JM, Laskowski RA, Borkakoti N (2021) AlphaFold heralds a data-driven revolution in biology and medicine. Nat Med 27:1666–1669. https://doi.org/10.1038/s41591-021-01533-0
Baek M, DiMaio F, Anishchenko I et al (2021) Accurate prediction of protein structures and interactions using a three-track neural network. Science 373:871–876. https://doi.org/10.1126/science.abj8754
Ramanathan A, Savol A, Burger V et al (2014) Protein conformational populations and functionally relevant substates. Acc Chem Res 47:149–156. https://doi.org/10.1021/ar400084s
Weis WI, Kobilka BK (2018) The molecular basis of G protein–coupled receptor activation. Annu Rev Biochem 87:897–919. https://doi.org/10.1146/annurev-biochem-060614-033910
Modi V, Dunbrack RL (2019) Defining a new nomenclature for the structures of active and inactive kinases. Proc Nat Acad Sci 116:6818–6827. https://doi.org/10.1073/pnas.1814279116
Xie T, Saleh T, Rossi P et al (2020) Conformational states dynamically populated by a kinase determine its function. Science 370:eabc2754. https://doi.org/10.1126/science.abc2754
Skolnick J, Gao M, Zhou H et al (2021) AlphaFold 2: why it works and its implications for understanding the relationships of protein sequence, structure, and function. J Chem Inf Model 61:4827–4831. https://doi.org/10.1021/acs.jcim.1c01114
Boehr DD, Nussinov R, Wright PE (2009) The role of dynamic conformational ensembles in biomolecular recognition. Nat Chem Biol 5:789–796. https://doi.org/10.1038/nchembio.232
Shaw DE, Maragakis P, Lindorff-Larsen K et al (2010) Atomic-level characterization of the structural dynamics of proteins. Science 330:341–346. https://doi.org/10.1126/science.1187409
Cournia Z, Allen TW, Andricioaei I et al (2015) Membrane protein structure, function, and dynamics: a perspective from experiments and theory. J Membr Biol 248:611–640. https://doi.org/10.1007/s00232-015-9802-0
Campbell E, Kaltenbach M, Correy GJ et al (2016) The role of protein dynamics in the evolution of new enzyme function. Nat Chem Biol 12:944–950. https://doi.org/10.1038/nchembio.2175
del Alamo D, Sala D, McHaourab HS et al (2022) Sampling alternative conformational states of transporters and receptors with AlphaFold2. Elife 11:e75751. https://doi.org/10.7554/eLife.75751
Zacharias M (2017) Predicting allosteric changes from conformational ensembles. Structure 25:393–394. https://doi.org/10.1016/j.str.2017.02.006
de Groot BL, van Aalten DMF, Scheek RM et al (1997) Prediction of protein conformational freedom from distance constraints. Proteins 29:240–251. https://doi.org/10.1002/(SICI)1097-0134(199710)29:2%3c240::AID-PROT11%3e3.0.CO;2-O
de Groot BL, Hayward S, van Aalten DMF et al (1998) Domain motions in bacteriophage T4 lysozyme: a comparison between molecular dynamics and crystallographic data. Proteins 31:116–127. https://doi.org/10.1002/(SICI)1097-0134(19980501)31:2%3c116::AID-PROT2%3e3.0.CO;2-K
de Groot BL, Vriend G, Berendsen HJC (1999) Conformational changes in the chaperonin GroEL: new insights into the allosteric mechanism11 edited by A. R. Fersht. J Mol Biol 286:1241–1249. https://doi.org/10.1006/jmbi.1998.2568
Seeliger D, Haas J, de Groot BL (2007) Geometry-based sampling of conformational transitions in proteins. Structure 15:1482–1492. https://doi.org/10.1016/j.str.2007.09.017
Feng Q, Hou M, Liu J et al (2022) Construct a variable-length fragment library for de novo protein structure prediction. Brief Bioinform 23:bbac086. https://doi.org/10.1093/bib/bbac086
Zhao KL, Xia YH, Zhang FJ et al (2023) Protein structure and folding pathway prediction based on remote homologs recognition using PAthreader. Commun Biol 6:243. https://doi.org/10.1038/s42003-023-04605-8
Haliloglu T, Hacisuleyman A, Erman B (2022) Prediction of allosteric communication pathways in proteins. Bioinformatics 38:3590–3599. https://doi.org/10.1093/bioinformatics/btac380
Zhao KL, Liu J, Zhou XG et al (2021) MMpred: a distance-assisted multimodal conformation sampling for de novo protein structure prediction. Bioinformatics 37:4350–4356. https://doi.org/10.1093/bioinformatics/btab484
Meng Z, Yıldız BS, Li G et al (2023) Application of state-of-the-art multiobjective metaheuristic algorithms in reliability-based design optimization: a comparative study. Struct Multidiscip Optim 66:191. https://doi.org/10.1007/s00158-023-03639-0
Panagant N, Pholdee N, Bureerat S et al (2021) A comparative study of recent multi-objective metaheuristics for solving constrained truss optimisation problems. Arch Comput Method Eng 28:4031–4047. https://doi.org/10.1007/s11831-021-09531-8
Günaydın AC, Yıldız AR, Kaya N (2022) Multi-objective optimization of build orientation considering support structure volume and build time in laser powder bed fusion. Mater Test 64:323–338. https://doi.org/10.1515/mt-2021-2075
Anosri S, Panagant N, Champasak P et al (2023) A comparative study of state-of-the-art metaheuristics for solving many-objective optimization problems of fixed wing unmanned aerial vehicle conceptual design. Arch Comput Method Eng 30:3657–3671. https://doi.org/10.1007/s11831-023-09914-z
Hong Z, Yu L, Zhang G (2010) A novel method for adaptive determination clusters number based on N-order nearest neighbor. In: Proceedings of the 29th Chinese control conference. IEEE, p 3007–3011. https://ieeexplore.ieee.org/abstract/document/5573321
Liu J, Zhou XG, Zhang Y et al (2020) CGLFold: a contact-assisted de novo protein structure prediction using global exploration and loop perturbation sampling algorithm. Bioinformatics 36:2443–2450. https://doi.org/10.1093/bioinformatics/btz943
Liu J, Zhao KL, He GX et al (2021) A de novo protein structure prediction by iterative partition sampling, topology adjustment and residue-level distance deviation optimization. Bioinformatics 38:99–107. https://doi.org/10.1093/bioinformatics/btab620
Zhang G, Hou M, Peng C et al (2021) An overview of multi-domain protein structure prediction methods. J Univ Electron Sci Technol China. https://doi.org/10.12178/1001-0548.2022132
Peng CX, Zhou XG, Liu J et al (2023) Multiple conformational states assembly of multidomain proteins using evolutionary algorithm based on structural analogues and sequential homologues. bioRxiv. https://doi.org/10.1101/2023.01.15.524086
Kabsch W, Sander C (1983) Dictionary of protein secondary structure: Pattern recognition of hydrogen-bonded and geometrical features. Biopolymers 22:2577–2637. https://doi.org/10.1002/bip.360221211
Fu L, Niu B, Zhu Z et al (2012) CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics 28:3150–3152. https://doi.org/10.1093/bioinformatics/bts565
Boocock GRB, Morrison JA, Popovic M et al (2003) Mutations in SBDS are associated with Shwachman–Diamond syndrome. Nat Genet 33:97–101. https://doi.org/10.1038/ng1062
Senger B, Lafontaine DLJ, Graindorge J-S et al (2001) The nucle(ol)ar Tif6p and Efl1p are required for a late cytoplasmic step of ribosome synthesis. Mol Cell 8:1363–1373. https://doi.org/10.1016/S1097-2765(01)00403-8
Finch AJ, Hilcenko C, Basse N et al (2011) Uncoupling of GTP hydrolysis from eIF6 release on the ribosome causes Shwachman–Diamond syndrome. Genes Dev 25:917–929. https://doi.org/10.1101/gad.623011
Weis F, Giudice E, Churcher M et al (2015) Mechanism of eIF6 release from the nascent 60S ribosomal subunit. Nat Struct Mol Biol 22:914–919. https://doi.org/10.1038/nsmb.3112
Nicoludis JM, Gaudet R (2018) Applications of sequence coevolution in membrane protein biochemistry. Biochim Biophys Acta Biomembr 1860:895–908. https://doi.org/10.1016/j.bbamem.2017.10.004
Garcia CK, Goldstein JL, Pathak RK et al (1994) Molecular characterization of a membrane transporter for lactate, pyruvate, and other monocarboxylates: Implications for the Cori cycle. Cell 76:865–873. https://doi.org/10.1016/0092-8674(94)90361-1
Ritzhaupt A, Wood IS, Ellis A et al (1998) Identification and characterization of a monocarboxylate transporter (MCT1) in pig and human colon: its potential to transport l-lactate as well as butyrate. J Physiol 513:719–732. https://doi.org/10.1111/j.1469-7793.1998.719ba.x
Wang N, Jiang X, Zhang S et al (2021) Structural basis of human monocarboxylate transporter 1 inhibition by anti-cancer drug candidates. Cell 184:370–383. https://doi.org/10.1016/j.cell.2020.11.043
Acknowledgements
This work is supported by the National Key R&D Program of China (2022ZD0115103), the National Nature Science Foundation of China (No. 62173304), and the Key Project of Zhejiang Provincial Natural Science Foundation of China (No. LZ20F030002).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no confict of interest.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hou, M., Jin, S., Cui, X. et al. Protein Multiple Conformation Prediction Using Multi-Objective Evolution Algorithm. Interdiscip Sci Comput Life Sci (2024). https://doi.org/10.1007/s12539-023-00597-5
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12539-023-00597-5