Abstract
Background
The molecular characteristics and antimicrobial susceptibility of Staphylococcus aureus (S. aureus) in general pediatric wards and county-level hospitals were rarely reported in China.
Methods
Staphylococcus aureus was isolated from children hospitalized with respiratory tract infection (RTI) in Zhongjiang and Youyang counties in 2015. All isolates were typed by multilocus sequence, staphylococcal protein A, accessory gene regulator (agr), and staphylococcal cassette chromosome mec [SCCmec, for methicillin-resistant S. aureus (MRSA) only]. Polymerase chain reaction was used to screen 21 super-antigen (SAg) genes and panton–valentine leukocidin (pvl). Antimicrobial susceptibility testing was performed by E test.
Results
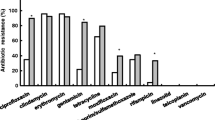
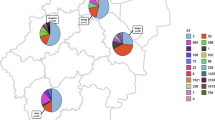
A total of 2136 children were enrolled. Overall, 125 (5.9%) children carried S. aureus, among which MRSA accounted for 42.4%. ST59-SCCmec type IV-t437-agr group I (58.5%) was the most prevalent genotype in MRSA, and ST188-t189-agr group I (22.2%) was the top genotype in methicillin-sensitive S. aureus (MSSA). The pvl carriage rate in MRSA and MSSA was 15.1% and 9.7%, respectively (P = 0.4112). About 96.8% of S. aureus isolates were positive for at least one SAg gene. The most common SAg gene profile in the dominant ST59 clone was seb–sek–seq (42.8%). All S. aureus isolates were resistant to penicillin and erythromycin (minimum inhibitory concentration 90 was > 32 and 256 mg/L to penicillin and erythromycin, respectively), but usually susceptible to other tested non-β-lactam antimicrobials.
Conclusions
Staphylococcus aureus and MRSA were detected with a high frequency in children with RTI in county-level hospitals of China. ST59-SCCmec type IV-t437-agr group I was the dominant MRSA clone. The S. aureus isolates exhibited high resistance to penicillin and erythromycin.

Similar content being viewed by others
References
Laux C, Peschel A, Krismer B. Staphylococcus aureus colonization of the human nose and interaction with other microbiome members. Microbiol Spectr. 2019. https://doi.org/10.1128/microbiolspec.GPP3-0029-2018.
Sakr A, Brégeon F, Mège JL, Rolain JM, Blin O. Staphylococcus aureus nasal colonization: an update on mechanisms, epidemiology, risk factors, and subsequent infections. Front Microbiol. 2018;9:2419.
Butterly A, Schmidt U, Wiener-Kronish J. Methicillin-resistant Staphylococcus aureus colonization, its relationship to nosocomial infection, and efficacy of control methods. Anesthesiology. 2010;113:1453–9.
Grumann D, Nübel U, Bröker BM. Staphylococcus aureus toxins—their functions and genetics. Infect Genet Evol. 2014;21:583–92.
Zollner TM, Wichelhaus TA, Hartung A, Von Mallinckrodt C, Wagner TO, Brade V, et al. Colonization with superantigen-producing Staphylococcus aureus is associated with increased severity of atopic dermatitis. Clin Exp Allergy. 2000;30:994–1000.
Gesualdo F, Bongiorno D, Rizzo C, Bella A, Menichella D, Stefani S, et al. MRSA nasal colonization in children: prevalence meta-analysis, review of risk factors and molecular genetics. Pediatr Infect Dis J. 2013;32:479–85.
Lyles RD, Trick WE, Hayden MK, Lolans K, Fogg L, Logan LK. Regional epidemiology of methicillin-resistant Staphylococcus aureus among critically ill children in a state with mandated active surveillance. J Pediatric Infect Dis Soc. 2016;5:409–16.
Zervou FN, Zacharioudakis IM, Ziakas PD, Mylonakis E. MRSA colonization and risk of infection in the neonatal and pediatric ICU: a meta-analysis. Pediatrics. 2014;133:1015–23.
Li S, Sun J, Zhang J, Li X, Tao X, Wang L, et al. Comparative analysis of the virulence characteristics of epidemic methicillin-resistant Staphylococcus aureus (MRSA) strains isolated from Chinese children: ST59 MRSA highly expresses core gene-encoded toxin. APMIS. 2014;122:101–14.
Enright MC, Day NP, Davies CE, Peacock SJ, Spratt BG. Multilocus sequence typing for characterization of methicillin-resistant and methicillin-susceptible clones of Staphylococcus aureus. J Clin Microbiol. 2000;38:1008–155.
Koreen L, Ramaswamy SV, Graviss EA, Naidich S, Musser JM, Kreiswirth BN. spa typing method for discriminating among Staphylococcus aureus isolates: implications for use of a single marker to detect genetic micro- and macrovariation. J Clin Microbiol. 2004;42:792–9.
Gilot P, Lina G, Cochard T, Poutrel B. Analysis of the genetic variability of genes encoding the RNA III-activating components Agr and TRAP in a population of Staphylococcus aureus strains isolated from cows with mastitis. J Clin Microbiol. 2002;40:4060–7.
Milheirico C, Oliveira DC, de Lencastre H. Multiplex PCR strategy for subtyping the staphylococcal cassette chromosome mec type IV in methicillin-resistant Staphylococcus aureus: SCCmec IV multiplex. J Antimicrob Chemother. 2007;60:42–8.
Holtfreter S, Grumann D, Schmudde M, Nguyen HT, Eichler P, Strommenger B, et al. Clonal distribution of superantigen genes in clinical Staphylococcus aureus isolates. J Clin Microbiol. 2007;45:2669–800.
Hesje CK, Sanfilippo CM, Haas W, Morris TW. Molecular epidemiology of methicillin-resistant and methicillin-susceptible Staphylococcus aureus isolated from the eye. Curr Eye Res. 2011;36:94–102.
The European Committee on Antimicrobial Susceptibility Testing. Breakpoint tables for interpretation of MICs and zone diameters. Version 7.1. 2017. https://www.eucast.org. Accessed 19 Sep 2017.
Clinical and Laboratory Standards Institute. Performance standards for antimicrobial susceptibility testing; 24th informational supplement. CLSI Stand Antimicrob Suscept Test. 2014;34:1–226.
Lin J, Peng Y, Xu P, Zhang T, Bai C, Lin D, et al. Methicillin-resistant Staphylococcus aureus nasal colonization in Chinese children: a prevalence meta-analysis and review of influencing factors. PLoS One. 2016;11:e0159728.
Fu JJ, Ye XH, Fan YP, Chen SD. Analysis on risk factors of Staphylococcus aureus nasal vestibule colonization among Guangzhou healthy children. ChongQing Medicine. 2015;44:3669–711.
Pan H, Cui B, Huang Y, Yang J, Ba-Thein W. Nasal carriage of common bacterial pathogens among healthy kindergarten children in Chaoshan region, southern China: a cross-sectional study. BMC Pediatr. 2016;16:161.
Ho PL, Chiu SS, Chan MY, Gan Y, Chow KH, Lai EL, et al. Molecular epidemiology and nasal carriage of Staphylococcus aureus and methicillin-resistant S. aureus among young children attending day care centers and kindergartens in Hong Kong. J Infect. 2012;64:500–6.
Regev-Yochay G, Raz M, Carmeli Y, Shainberg B, Navon-Venezia S, Pinco E, et al. Parental Staphylococcus aureus carriage is associated with staphylococcal carriage in young children. Pediatr Infect Dis J. 2009;28:960–5.
Eibach D, Nagel M, Hogan B, Azuure C, Krumkamp R, Dekker D, et al. Nasal carriage of Staphylococcus aureus among children in the Ashanti region of Ghana. PLoS One. 2017;12:e0170320.
Lo WT, Wang CC, Lin WJ, Wang SR, Teng CS, Huang CF, et al. Changes in the nasal colonization with methicillin-resistant Staphylococcus aureus in children: 2004–2009. PLoS One. 2010;5:e15791.
Pan HH, Huang YC, Chen CJ, Huang FL, Ting PJ, Huang JY, et al. Prevalence of and risk factors for nasal methicillin-resistant Staphylococcus aureus colonization among children in central Taiwan. J Microbiol Immunol Infect. 2019;52:45–53.
Mediavilla JR, Chen L, Mathema B, Kreiswirth BN. Global epidemiology of community-associated methicillin resistant Staphylococcus aureus (CA-MRSA). Curr Opin Microbiol. 2012;15:588–95.
Chen CJ, Huang YC. New epidemiology of Staphylococcus aureus infection in Asia. Clin Microbiol Infect. 2014;20:605–23.
Kale P, Dhawan B. The changing face of community-acquired methicillin-resistant Staphylococcus aureus. Indian J Med Microbiol. 2016;34:275–85.
Yang X, Qian S, Yao K, Wang L, Liu Y, Dong F, et al. Multiresistant ST59-SCCmec IV-t437 clone with strong biofilm-forming capacity was identified predominantly in MRSA isolated from Chinese children. BMC Infect Dis. 2017;17:733.
Qiao Y, Ning X, Chen Q, Zhao R, Song W, Zheng Y, et al. Clinical and molecular characteristics of invasive community-acquired Staphylococcus aureus infections in Chinese children. BMC Infect Dis. 2014;14:582.
Deurenberg RH, Stobberingh EE. The evolution of Staphylococcus aureus. Infect Genet Evol. 2008;8:747–63.
Deodhar D, Varghese G, Balaji V, John J, Rebekah G, Janardhanan J, et al. Prevalence of toxin genes among the clinical isolates of Staphylococcus aureus and its clinical impact. J Glob Infect Dis. 2015;7:97–102.
Arabestani MR, Rastiyani S, Alikhani MY, Mousavi SF. The relationship between prevalence of antibiotics resistance and virulence factors genes of MRSA and MSSA strains isolated from clinical samples, west Iran. Oman Med J. 2018;33:134–40.
Chang W, Ma X, Gao P, Lv X, Lu H, Chen F. Vancomycin MIC creep in methicillin-resistant Staphylococcus aureus (MRSA) isolates from 2006 to 2010 in a hospital in China. Indian J Med Microbiol. 2015;33:262–6.
Qin Y, Wen F, Zheng Y, Zhao R, Hu Q, Zhang R. Antimicrobial resistance and molecular characteristics of methicillin-resistant Staphylococcus aureus isolates from child patients of high-risk wards in Shenzhen. China Jpn J Infect Dis. 2017;70:479–84.
Chen CJ, Hsu KH, Lin TY, Hwang KP, Chen PY, Huang YC. Factors associated with nasal colonization of methicillin-resistant Staphylococcus aureus among healthy children in Taiwan. J Clin Microbiol. 2011;49:131–7.
Botelho-Nevers E, Gagnaire J, Verhoeven PO, Cazorla C, Grattard F, Pozzetto B, et al. Decolonization of Staphylococcus aureus carriage. Med Mal Infect. 2017;47:305–10.
Poovelikunnel T, Gethin G, Humphreys H. Mupirocin resistance: clinical implications and potential alternatives for the eradication of MRSA. J Antimicrob Chemother. 2015;70:2681–92.
Song Y, Cui L, Lv Y, Li Y, Xue F. Characterisation of clinical isolates of oxacillin-susceptible mecA-positive Staphylococcus aureus in China from 2009 to 2014. J Glob Antimicrob Resist. 2017;11:1–3.
Pournaras S, Sabat AJ, Grundmann H, Hendrix R, Tsakris A, Friedrich AW. Driving forces of mechanisms regulating oxacillin-resistance phenotypes of MRSA: truly oxacillin-susceptible mecA-positive Staphylococcus aureus clinical isolates also exist. Curr Pharm Des. 2015;21:2048–53.
Giannouli S, Labrou M, Kyritsis A, Ikonomidis A, Pournaras S, Stathopoulos C, et al. Detection of mutations in the FemXAB protein family in oxacillin-susceptible mecA-positive Staphylococcus aureus clinical isolates. J Antimicrob Chemother. 2010;65:626–33.
Acknowledgements
We are very grateful to the local clinical laboratory staff, Ping Tang, Hai-Ling Zeng in People's Hospital of Zhongjiang County and Xiao-Ping Cheng in Youyang Hospital, respectively, who kindly helped us in treating and storing the clinical isolates. We also thank pediatricians in the two hospitals involved in collecting the samples.
Funding
This work was supported by the National Natural Science Foundation of China (No. 81571948) and the Beijing Natural Science Foundation (No. 7172075), the medical research project of Chongqing Health and Family Planning Commission (No. 2016ZDX041).
Author information
Authors and Affiliations
Contributions
SYQ and KHY designed the study, reviewed and revised the manuscript. CS and QW conducted the experiments and wrote the manuscript, and contributed equally to this work. WTL, XY, WS and QHM oversaw data analysis planning and execution. DNW and CHC designed the data collection instruments and coordinated sample collection. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethical approval
This study was approved by the Ethics Committee of Beijing Children’s Hospital Affiliated to Capital Medical University. Written informed consents were obtained from the patients’ legal guardian.
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Sun, C., Wang, Q., Li, WT. et al. Molecular characteristics and antimicrobial susceptibility of Staphylococcus aureus among children with respiratory tract infections in southwest China. World J Pediatr 16, 284–292 (2020). https://doi.org/10.1007/s12519-019-00317-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12519-019-00317-4