Abstract
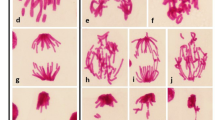
Ribosomal DNA is an important repeated sequence that forms the nucleolus at the interphase. Its transcription into ribosomal RNA for ribosome biogenesis also represents a transitional point for several cellular processes, including cell-cycle progression, gene-silencing, and formation of the ribonucleoprotein complex. The levels of rDNA acetylation have an important role in regulating structural changes in rDNA chromatin and transcriptional activity. Using root-tip samples from Brassica campestris, we determined that some rDNA chromatin is located in the heterochromatin regions while some is de-condensed and found in euchromatin regions. Immuno-staining results showed that histone H4K5 acetylation and H4 tetra-acetylation signals are dispersed within the euchromatin. Analysis of the promoter and exon regions of rDNA via chromatin immuno-precipitation (ChIP) revealed a connection between histone acetylation and rDNA conformation.
Similar content being viewed by others
References
Claypool JA, French SL, Johzuka K, Eliason K, Vu L, Dodd JA, Beyer AL, Nomura M (2004) Tor pathway regulates Rrn3p-dependent recruitment of yeast RNA polymerase I to the promoter but does not participate in alteration of the number of active genes. Mol Biol Cell 15:946–956
Clayton AL, Hazzalin CA, Mahadevan LC (2006) Enhanced histone acetylation and transcription: a dynamic perspective. Mol Cell 23:289–296
Crane-Robinson C, Myers FA, Hebbes TR, Clayton AL, Thorne AW (1999) Chromatin immunoprecipitation assays in acetylation mapping of higher eukaryotes. Meth Enzymol 304:533–547
Fuchs J, Demidov D, Houben A, Schubert I (2006) Chromosomal histone modification patterns—from conservation to diversity. Trends Plant Sci 11:199–208
Grummt I (2003) Life on a planet of its own: regulation of RNA polymerase I transcription in the nucleolus. Genes Dev 17:1691–1702
Grummt I (2007) Different epigenetic layers engage in complex crosstalk to define the epigenetic state of mammalian rRNA genes. Hum Mol Genet 16:Spec No 1 R21–R27
Haring M, Offermann S, Danker T, Horst I, Peterhansel C, Stam M (2007) Chromatin immunoprecipitation: optimization, quantitative analysis and data normalization. Plant Meth 3:11
Hu Y, Zhang L, Zhao L, Li J, He S, Zhou K, Yang F, Huang M, Jiang L, Li L (2011) Trichostatin A Selectively Suppresses the Cold- Induced Transcription of the ZmDREB1 Gene in Maize. PLoS ONE 6:e22132
Jasencakova Z, Meister A, Walter J, Turner BM, Schubert I (2000) Histone H4 acetylation of euchromatin and heterochromatin is cell cycle dependent and correlated with replication rather than with transcription. Plant Cell 12:2087–2100
Jenuwein T, Allis CD (2001) Translating the histone code. Science 293:1074–1080
Kornberg RD (1977) Structure of chromatin. Annu Rev Biochem 46:931–954
Krebs JE, Kuo MH, Allis CD, Peterson CL (1999) Cell cycle-regulated histone acetylation required for expression of the yeast HO gene. Genes Dev 13:1412–1421
Lawrence RJ, Earley K, Pontes O, Silva M, Chen ZJ, Neves N, Viegas W, Pikaard CS (2004) A concerted DNA methylation/histone methylation switch regulates rRNA gene dosage control and nucleolar dominance. Mol Cell 13:599–609
Li J, He S, Zhang L, Hu Y, Yang F, Ma L, Huang J, Li L (2012) Telomere and 45S rDNA sequences are structurally linked on the chromosomes in Chrysanthemum segetum L. Protoplasma 249:207–215
Li L, Yang J, Tong Q, Zhao L, Song Y (2005) A novel approach to prepare extended DNA fibers in plants. Cytometry A 63:114–117
Loidl P (2004) A plant dialect of the histone language. Trends Plant Sci 9:84–90
Lusser A, Kolle D, Loidl P (2001) Histone acetylation: lessons from the plant kingdom. Trends Plant Sci 6:59–65
Mathieu O, Jasencakova Z, Vaillant I, Gendrel AV, Colot V, Schubert I, Tourmente S (2003) Changes in 5S rDNA chromatin organization and transcription during heterochromatin establishment in Arabidopsis. Plant Cell 15:2929–2939
McStay B (2006) Nucleolar dominance: a model for rRNA gene silencing. Genes Dev 20:1207–1214
Moss T, Stefanovsky VY (2002) At the center of eukaryotic life. Cell 109:545–548
Moss T, Langlois F, Gagnon-Kugler T, Stefanovsky V (2007) A housekeeper with power of attorney: the rRNA genes in ribosome biogenesis. Cell Mol Life Sci 64:29–49
Murayama A, Ohmori K, Fujimura A, Minami H, Yasuzawa-Tanaka K, Kuroda T, Oie S, Daitoku H, Okuwaki M, Nagata K, Fukamizu A, Kimura K, Shimizu T, Yanagisawa J (2008) Epigenetic control of rDNA loci in response to intracellular energy status. Cell 133:627–639
Peng JC, Karpen GH (2008) Epigenetic regulation of heterochromatic DNA stability. Curr Opin Genet Dev 18:204–211
Preuss S, Pikaard CS (2007) rRNA gene silencing and nucleolar dominance: insights into a chromosome-scale epigenetic on/off switch. Biochim Biophys Acta 1769:383–392
Sandmeier JJ, French S, Osheim Y, Cheung WL, Gallo CM, Beyer AL, Smith JS (2002) RPD3 is required for the inactivation of yeast ribosomal DNA genes in stationary phase. EMBO J 21:4959–4968
Santoro R (2005) The silence of the ribosomal RNA genes. Cell Mol Life Sci 62:2067–2079
Turner BM (2002) Cellular memory and the histone code. Cell 111:285–291
Yang F, Zhang L, Li J, Huang J, Wen R, Ma L, Zhou D, Li L (2010) Trichostatin A and 5-azacytidine both cause an increase in global histone H4 acetylation and a decrease in global DNA and H3K9 methylation during mitosis in maize. BMC Plant Biol 10:178
Zhang J, Xu F, Hashimshony T, Keshet I, Cedar H (2002) Establishment of transcriptional competence in early and late S phase. Nature 420:198–202
Zhang L, Qiu Z, Hu Y, Yang F, Yan S, Zhao L, Li B, He S, Huang M, Li J, Li L (2011) ABA treatment of germinating maize seeds induces VP1 gene expression and selective promoter-associated histone acetylation. Physiol Plant 143:287–296
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Qiu, Z., Zhang, L., Hu, Y. et al. The acetylation level of rDNA in Brassica campestris . J. Plant Biol. 55, 298–302 (2012). https://doi.org/10.1007/s12374-011-0317-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12374-011-0317-7