Abstract
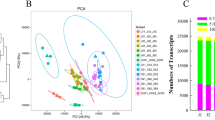
To screen candidate marker genes implicated in vernalization, bolting-related agronomic traits and the expression of genes homologous to the bolting and flowering-related genes in Arabidopsis thaliana were investigated in KWS9147 during 16-week (w) chilling. Bolting-related agronomic traits showed that less than 10-w chilling was not sufficient for bolting. A transitional period was determined between 10 and 14 weeks when vernalization time was enough for bolting in some seedlings. After 14-w chilling, nearly all seedlings were well vernalized. At the transcriptional level, the expression of 19 candidate genes was significantly changed in response to chilling (change fold > 2 and P value < 0.05). In particular, three genes (BvGI, BvBTC1 and BvFVE1) were notable, as their expression continuously increased or decreased during 8–16-w chilling, when BI and BD were significantly changed. On the basis of correlation analysis, a co-expression network containing two modules was constructed. BvGI, BvBTC1 and BvFVE1 were located at the centre of the first module, which co-expressed with the other 3 photoperiodic genes. In the second module, BvFT1 and BvFT2 were associated with BvLHY, BvGATA22 and BvFVE2. Our study on the temporal expression pattern of bolting and flowering-related genes during vernalization provides valuable information for preliminary screening of key regulators and may facilitate the future molecular breeding of vernalization-insensitive varieties.





Similar content being viewed by others
Availability of Data and Material
Not applicable.
References
Abe, Jun, G.P. Guan, and Y. Shimamoto. 1997. A gene complex for annual habit in sugar beet (Beta vulgaris L.). Euphytica 94 (2): 129–135.
Abou-Elwafa, S.F., B. Büttner, T. Chia, G. Schulze-Buxloh, U. Hohmann, E. Mutasa-Göttgens, C. Jung, and A.E. Muller. 2011. Conservation and divergence of autonomous pathway genes in the flowering regulatory network of Beta vulgaris. Journal of Experimental Botany 62 (10): 3359–3374.
Amasino, R. 2010. Seasonal and developmental timing of flowering. Plant Journal 61 (6): 1001–1013.
Ausín, I., C. Alonso-Blanco, J.A. Jarillo, L. Ruiz-García, and J. Martínez-Zapater. 2004. Regulation of flowering time by FVE, a retinoblastoma-associated protein. Nature Genetics 36 (2): 162–166.
Biancardi, E. 2005. Genetics and breeding of sugar beet. Boca Raton: CRC Press.
Chia, T.Y.P., A. Müller, C. Jung, and E.S. Mutasa-Gcettgens. 2008. Sugar beet contains a large CONSTANS-LIKE gene family including a CO homologue that is independent of the early-bolting (B) gene locus. Journal of Experimental Botany 59 (10): 2735–2748.
Chow, B.Y., A. Helfer, D.A. Nusinow, and S.A. Kay. 2012. ELF3 recruitment to the PRR9 promoter requires other Evening Complex members in the Arabidopsis circadian clock. Plant Signaling & Behavior 7 (2): 170–173.
Dai, S., X. Wei, L. Pei, R.L. Thompson, Y. Liu, J.E. Heard, T.G. Ruff, and R.N. Beachy. 2011. BROTHER OF LUX ARRHYTHMO is a component of the Arabidopsis circadian clock. Plant Cell 23 (3): 961–972.
Dally, N., K. Xiao, D. Holtgräwe, and C. Jung. 2014. The B2 flowering time locus of beet encodes a zinc finger transcription factor. Proceedings of the National Academy of Sciences of the United States of America 111 (28): 10365–10370.
Ginestet, C. 2011. ggplot2: elegant graphics for data analysis. Journal of the Royal Statistical Society: Series A (Statistics in Society) 174 (1): 245–246.
Goodstein, D.M., S. Shu, R. Howson, R. Neupane, R.D. Hayes, J. Fazo, T. Mitros, W. Dirks, U. Hellsten, N. Putnam, and D.S. Rokhsar. 2012. Phytozome: a comparative platform for green plant genomics. Nucleic Acids Research 40 (Database issue): D1178–D1186.
Hayama, R., S. Yokoi, S. Tamaki, M. Yano, and K. Shimamoto. 2003. Adaptation of photoperiodic control pathways produces short-day flowering in rice. Nature 422 (6933): 719–722.
He, Z., H. Zhang, S. Gao, M.J. Lercher, W.H. Chen, and S. Hu. 2016. Evolview v2: an online visualization and management tool for customized and annotated phylogenetic trees. Nucleic Acids Research 44 (Web Server issue): W236–W241.
Hecht, V., F. Foucher, C. Ferrandiz, R. Macknight, C. Navarro, J. Morin, M.E. Vardy, N. Ellis, J.P. Beltran, C. Rameau, and J.L. Weller. 2005. Conservation of Arabidopsis flowering genes in model Legumes. Plant Physiology 137 (4): 1420–1434.
Hoffmann, C.M., and S. Kluge-Severin. 2010. Light absorption and radiation use efficiency of autumn and spring sown sugar beets. Field Crops Research 119 (2): 238–244.
Hong, S.Y., S. Lee, P.J. Seo, M.S. Yang, and C.M. Park. 2010. Identification and molecular characterization of a Brachypodium distachyon GIGANTEA gene: functional conservation in monocot and dicot plants. Plant Molecular Biology 72 (4–5): 485–497.
Johnson, L.S., S.R. Eddy, and E. Portugaly. 2010. Hidden Markov model speed heuristic and iterative HMM search procedure. BMC Bioinformatics 11 (1): 431.
Jung, C., and A.E. Müller. 2009. Flowering time control and applications in plant breeding. Trends in Plant Science 14 (10): 563–573.
Kalyaanamoorthy, S., B.Q. Minh, T.K.F. Wong, A.V. Haeseler, and L.S. Jermiin. 2017. ModelFinder: fast model selection for accurate phylogenetic estimates. Nature Methods 14 (6): 587–589.
Kassambara, A. and F. Mundt. 2016. Factoextra: extract and visualize the results of multivariate data analyses. R package version 1 (3):2016.
Kim, D., and S. Sung. 2014. Genetic and epigenetic mechanisms underlying vernalization. The Arabidopsis Book/American Society of Plant Biologists 12: e0171.
Kim, H.J., Y. Hyun, J. Park, M. Park, M. Park, M.D. Kim, H. Kim, M.H. Lee, J. Moon, I. Lee, and J. Kim. 2004. A genetic link between cold responses and flowering time through FVE in Arabidopsis thaliana. Nature Genetics 36 (2): 167–171.
Kim, W.Y., S. Fujiwara, S. Suh, J. Kim, Y. Kim, L.Q. Han, K. David, J. Putterill, H.G. Nam, and D.E. Somers. 2007. ZEITLUPE is a circadian photoreceptor stabilized by GIGANTEA in blue light. Nature 449 (7160): 356–360.
Kohl, M., S. Wiese, and B. Warscheid. 2011. Cytoscape: software for bisualization and analysis of biological networks. Methods in Molecular Biology 696: 291–303.
Kumar, S., G. Stecher, M. Li, C. Knyaz, and K. Tamura. 2018. MEGA X: molecular evolutionary genetics analysis across computing platforms. Molecular Biology and Evolution 35 (6): 1547–1549.
Lê, S., J. Josse, and F. Husson. 2008. FactoMineR: an R package for multivariate analysis. Journal of Statistical Software 25 (1): 1–18.
Li, W.B., T. Wang, Y.H. Zhang, and Y.G. Li. 2015. Overexpression of soybean miR172c confers tolerance to water deficit and salt stress, but increases ABA sensitivity in transgenic Arabidopsis thaliana. Journal of Experimental Botany 67 (1): 175–194.
Madeira, F., Y.M. Park, J. Lee, N. Buso, T. Gur, N. Madhusoodanan, P. Basutkar, A.R.N. Tivey, S.C. Potter, and R.D. Finn. 2019. The EMBL-EBI search and sequence analysis tools APIs in 2019. Nucleic Acids Research 47 (W1): W636–W641.
Milford, G.F.J. 2006. Plant structure and crop physiology. Oxford: Blackwell Publishing Ltd.
Nguyen, L.T., H.A. Schmidt, A. Von Haeseler, and B.Q. Minh. 2015. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Molecular Biology and Evolution 32 (1): 268–274.
Pfeiffer, N., A.E. Müller, C. Jung, and F.J. Kopisch-Obuch. 2017. QTL for delayed bolting after winter detected in leaf beet (Beta vulgaris L.). Plant Breeding 136 (2): 237–244.
Pin, P.A., R. Benlloch, D. Bonnet, E. Wremerth-Weich, T. Kraft, J.J.L. Gielen, and O. Nilsson. 2010. An antagonistic pair of FT homologs mediates the control of flowering time in sugar beet. Science 330 (6009): 1397–1400.
Pin, P.A., W.Y. Zhang, S.H. Vogt, N. Dally, B. Büttner, G. Schulze-Buxloh, N.S. Jelly, T.Y.P. Chia, E.S. Mutasa-Göttgens, J.C. Dohm, H. Himmelbauer, B. Weisshaar, J. Kraus, J.J.L. Gielen, M. Lommel, G. Weyens, B. Wahl, A. Schechert, O. Nilsson, C. Jung, T. Kraft, and A.E. Müller. 2012. The role of a pseudo-response regulator gene in life cycle adaptation and domestication of beet. Current Biology 22 (12): 1095–1101.
Reeves, P.A., Y.H. He, R.J. Schmitz, R.M. Amasino, L.W. Panella, and C.M. Richards. 2007. Evolutionary conservation of the FLOWERING LOCUS C-mediated vernalization response: evidence from the sugar beet (Beta vulgaris). Genetics 176 (1): 295–307.
Richter, R., C. Behringer, M. Zourelidou, and C. Schwechheimer. 2013. Convergence of auxin and gibberellin signalling on the regulation of the GATA transcription factors GNC and GNL in Arabidopsis thaliana. Proceedings of the National Academy of Sciences of the United States of America 110 (32): 13192–13197.
Searle, I., and G. Coupland. 2004. Induction of flowering by seasonal changes in photoperiod. The EMBO Journal 23: 1217–1222.
Somers, D.E., and S. Fujiwara. 2009. Thinking outside the F-box: novel ligands for novel receptors. Trends in Plant Science 14 (4): 206–213.
Sung, S., and R.M. Amasino. 2004. Vernalization in Arabidopsis thaliana is mediated by the PHD finger protein VIN3. Nature 427 (6970): 159–164.
Tao, Z., L. Shen, X. Gu, Y. Wang, H. Yu, and Y. He. 2017. Embryonic epigenetic reprogramming by a pioneer transcription factor in plants. Nature 551 (7678): 124–128.
Vogt, S.H., G. Weyens, M. Lefbvre, B. Bork, A. Schechert, and A.E. Mller. 2014. The FLC-like gene BvFL1 is not a major regulator of vernalization response in biennial beets. Frontiers in Plant Science 5: 146.
Wang, J. 2014. Regulation of flowering time by the miR156-mediated age pathway. Journal of Experimental Botany 65 (17): 4723–4730.
Wei, T., and V. Simko. 2013. corrplot: visualization of a correlation matrix. R Package Version 230 (231): 11.
Xu, S., and K. Chong. 2018. Remembering winter through vernalisation. Nature Plants 4 (12): 997–1009.
Ye, J., G. Coulouris, I. Zaretskaya, I. Cutcutache, S. Rozen, and T.L. Madden. 2012. Primer-BLAST: a tool to design target-specific primers for polymerase chain reaction. BMC Bioinformatics 13 (1): 134.
Yu, J.W., V. Rubio, N.Y. Lee, S. Bai, S.Y. Lee, S.S. Kim, L. Liu, Y. Zhang, M.L. Irigoven, J.A. Sulivan, Y. Zhang, I. Lee, Q. Xie, N.C. Paek, and X.W. Deng. 2009. COP1 and ELF3 control circadian function and photoperiodic flowering by regulating GI stability. Molecular Cell 32 (5): 617–630.
Funding
This work was supported by the Fundamental Research Fund for the Provincial Universities Basal Research Project in Heilongjiang Province (RCCXYJ201810), the China Agriculture Research System (CARS-170111), and the Project of Species and National Sugar Beet Germplasm Resources Platform (NICGR-2019-017).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest.
Ethics Approval
Not applicable.
Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Code Availability
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Pi, Z., Xing, W., Zhu, X. et al. Temporal Expression Pattern of Bolting-Related Genes During Vernalization in Sugar Beet. Sugar Tech 23, 146–157 (2021). https://doi.org/10.1007/s12355-020-00886-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12355-020-00886-z