Abstract
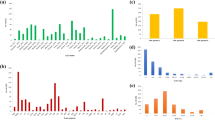
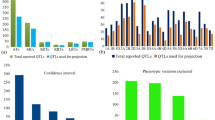
Meta-QTL (MQTL) analysis for drought tolerance was undertaken in bread wheat to identify consensus and robust MQTLs using 340 known QTLs from 11 earlier studies; 13 MQTLs located on 6 chromosomes (1D, 3B, 5A, 6D, 7A and 7D) were identified, with maximum of 4 MQTLs on chromosome 5A. Mean confidence intervals for MQTLs were much narrower (mean, 6.01 cM; range 2.07–19.46 cM), relative to those in original QTLs (mean, 13.6 cM; range, 1.0–119.1 cM). Two MQTLs, namely MQTL4 and MQTL12, were major MQTLs with potential for use in marker-assisting breeding. As many as 228 candidate genes (CGs) were also identified using 6 of the 13 MQTLs. In-silico expression analysis of these 228 CGs allowed identification of 14 important CGs, with + 3 to − 8 fold change in expression under drought (relative to normal conditions) in a tolerant cv. named TAM107. These CGs encoded proteins belonging to the following families: NAD-dependent epimerase/dehydratase, protein kinase, NAD(P)-binding domain protein, heat shock protein 70 (Hsp70), glycosyltransferase 2-like, etc. Important MQTLs and CGs identified in the present study should prove useful for future molecular breeding and for the study of molecular basis of drought tolerance in cereals in general and wheat in particular.



Similar content being viewed by others
References
Acuña-Galindo MA, Mason RE, Subramanian NK, Hays DB (2015) Meta-analysis of wheat QTL regions associated with adaptation to drought and heat stress. Crop Sci 55:477–492
Arcade A, Labourdette A, Falque M, Mangin B, Chardon F, Charcosset A, Joets J (2004) BioMercator: integrating genetic maps and QTL towards discovery of candidate genes. Bioinformatics 20:2324–2326
Ayalew H, Liu H, Börner A, Kobiljski B, Liu C, Yan G (2018) Genome-wide association mapping of major root length QTLs under PEG induced water stress in wheat. Front Plant Sci 9:1759
Ballini E, Morel JB, Droc G, Price A, Courtois B, Notteghem JL, Tharreau D (2008) A genome-wide meta-analysis of rice blast resistance genes and quantitative trait loci provides new insights into partial and complete resistance. Mol Plant Microbe Interact 21:859–868
Bennett D, Izanloo A, Reynolds M, Kuchel H, Langridge P, Schnurbusch T (2012) Genetic dissection of grain yield and physical grain quality in bread wheat (Triticum aestivum L.) under water-limited environments. Theor Appl Genet 125:255–271
Cattivelli L, Rizza F, Badeck FW, Mazzucotelli E, Mastrangelo AM, Francia E, Mare C, Tondelli A, Stanca AM (2008) Drought tolerance improvement in crop plants: an integrated view from breeding to genomics. Field Crops Res 105:1–14
Chardon F, Virlon B, Moreau L, Falque M, Joets J, Decousset L, Murigneux A, Charcosset A (2004) Genetic architecture of flowering time in maize as inferred from quantitative trait loci meta-analysis and synteny conservation with the rice genome. Genetics 168:2169–2185
Chen L, An Y, Li Y-x, Li C, Shi Y, Song Y, Zhang D, Wang T, Li Y (2017) Candidate loci for yield-related traits in maize revealed by a combination of metaQTL analysis and regional association mapping. Front Plant Sci 8:2190
Cho EK, Choi YJ (2009) A nuclear-localized HSP70 confers thermoprotective activity and drought-stress tolerance on plants. Biotechnol Lett 31:597–606
Coque M, Martin A, Veyrieras JB, Hirel B, Gallais A (2008) Genetic variation for N-remobilization and postsilking N-uptake in a set of maize recombinant inbred lines. 3. QTL detection and coincidences. Theor Appl Genet 117:729–747
Courtois B, Ahmadi N, Khowaja F, Price AH, Rami JF, Frouin J, Hamelin C, Ruiz M (2009) Rice root genetic architecture: meta-analysis from a drought QTL database. Rice 2:115–128
Danan S, Veyrieras JB, Lefebvre V (2011) Construction of a potato consensus map and QTL meta-analysis offer new insights into the genetic architecture of late blight resistance and plant maturity traits. BMC Plant Biol 1:16
Darvasi A, Soller M (1997) A simple method to calculate resolving power and confidence interval of QTL map location. Behav Genet 27:125–132
El-Feki WM, Byrne PF, Reid SD, Haley SD (2018) Mapping quantitative trait loci for agronomic traits in winter wheat under different soil moisture levels. Agronomy 8:133
Fang Y, Xie K, Xiong L (2014) Conserved miR164-targeted NAC genes negatively regulate drought resistance in rice. J Exp Bot 65:2119–2135
Farooq M, Hussain M, Siddique KH (2014) Drought stress in wheat during flowering and grain filling periods. Crit Rev Plant Sci 33:331–349
Gahlaut V, Jaiswal V, Tyagi BS, Singh G, Sareen S, Balyan HS, Gupta PK (2017) QTL mapping for nine drought-responsive agronomic traits in bread wheat under irrigated and rain-fed environments. PLoS ONE 12: e0182857
Gegas VC, Nazari A, Griffiths S, Simmonds J, Fish L, Orford S, Sayers L, Doonan JH, Snape JW (2010) A genetic framework for grain size and shape variation in wheat. Plant Cell 22:1042–1046
Goffinet B, Gerber S (2000) Quantitative trait loci: a meta-analysis. Genetics 155:463–473
Griffiths S, Simmonds J, Leverington M, Wang Y, Fish L, Sayers L, Alibert L, Orford S, Wingen L, Herry L, Faure S, Laurie D, Bilham L, Snape J (2009) Meta-QTL analysis of the genetic control of ear emergence in elite European winter wheat germplasm. Theor Appl Genet 119:383–395
Griffiths S, Simmonds J, Leverington M, Wang Y, Fish L, Sayers L, Alibert L, Orford S, Wingen L, Snape J (2012) Meta-QTL analysis of the genetic control of crop height in elite European winter wheat germplasm. Mol Breed 29:159–171
Guo B, Sleper DA, Lu P, Shannon JG, Nguyen HT, Arelli PR (2006) QTLs associated with resistance to soybean cyst nematode in soybean: meta-analysis of QTL locations. Crop Sci 46:595–602
Guo J, Chen L, Li Y, Shi Y, Song Y, Zhang D, Li Y, Wang T, Yang D, Li C (2018) Meta-QTL analysis and identification of candidate genes related to root traits in maize. Euphytica 2014:223
Gupta PK, Mir RR, Mohan A, Kumar J (2008) Wheat genomics: present status and future prospects. Int J Plant Genomics 2008:896451
Gupta A, Rico-Medina A, Ana I, Caño-Delgado AI (2020a) The physiology of plant responses to drought. Science 368:266–269
Gupta PK, Balyan HS, Sharma S, Kumar R (2020b) Genetics of yield, abiotic stress tolerance and biofortification in wheat (Triticum aestivum L). Theor Appl Genet 133:1569–1602
Häberle J, Holzapfel J, Schweizer G, Hartl L (2009) A major QTL for resistance against Fusarium head blight in European winter wheat. Theor Appl Genet 119:325–332
Hanocq E, Laperche A, Jaminon O, Laine AL, Le Gouis J (2007) Most significant genome regions involved in the control of earliness traits in bread wheat, as revealed by QTL meta-analysis. Theor Appl Genet 114:569–584
Hao Z, Li X, Liu X, Xie C, Li M, Zhang D, Zhang S (2010) Meta-analysis of constitutive and adaptive QTL for drought tolerance in maize. Euphytica 174:165–177
Hong Y, Chen X, Liang X, Liu H, Zhou G, Li S, Wen S, Holbrook CC, Guo B (2010) A SSR-based composite genetic linkage map for the cultivated peanut. (Arachis hypogaea L) genome. BMC Plant Biol 10:17
Hu H, Xiong L (2014) Genetic engineering and breeding of drought-resistant crops. Annu Rev Plant Biol 65:715–741
Islam MS, Ontoy J, Subudhi PK (2019) Meta-analysis of quantitative trait loci associated with seedling-stage salt tolerance in rice (Oryza sativa L). Plants 8:33
Khahani B, Tavakol E, Shariati V, Fornara F (2020) Genome wide screening and comparative genome analysis for meta-QTLs, ortho-MQTls and candidate genes controlling yield and yield-related traits in rice. BMC Genom 21:294
Khowaja FS, Norton GJ, Courtois B, Price AH (2009) Improved resolution in the position of drought-related QTLs in a single mapping population of rice by meta-analysis. BMC Genom 10:276
Kosina P, Reynolds M, Dixon J, Joshi A (2007) Stakeholder perception of wheat production constraints, capacity building needs, and research partnerships in developing countries. Euphytica 157:475–483
Kumar S, Sehgal SK, Kumar U, Prasad PV, Joshi AK, Gill BS (2012) Genomic characterization of drought tolerance-related traits in spring wheat. Euphytica 186:265–276
Lanaud C, Fouet O, Clément D, Boccara M, Risterucci AM, Surujdeo-Maharaj S, Legavre T, Argout X (2009) A meta-QTL analysis of disease resistance traits of Theobroma cacao L. Mol Breed 24:361–374
Li JZ, Zhang ZW, Li YL, Wang QL, Zhou YG (2011) QTL consistency and meta-analysis for grain yield components in three generations in maize. Theor Appl Genet 122:771–782
Li WT, Liu C, Liu YX, Pu ZE, Dai SF, Wang JR, Lan XJ, Zheng YL, Wei YM (2013) Meta-analysis of QTL associated with tolerance to abiotic stresses in barley. Euphytica 189:31–49
Li S, Chen N, Li F, Mei F, Wang Z, Cheng X, Kang Z, Mao H (2020) Characterization of wheat homeodomain-leucine zipper family genes and functional analysis of TaHDZ5-6A in drought tolerance in transgenic Arabidopsis. BMC Plant Biol 20:50
Liu S, Hall MD, Griffey CA, McKendry AL (2009) Meta-analysis of QTL associated with Fusarium head blight resistance in wheat. Crop Sci 49:1955–1968
Liu S, Zenda T, Wang X, Liu G, Jin H, Yang Y, Dong A, Duan H (2019) Comprehensive meta-analysis of maize QTLs associated with grain yield, flowering date and plant height under drought conditions. J Agric Sci 11:1–19
Loffler M, Schon CC, Miedaner T (2009) Revealing the genetic architecture of FHB resistance in hexaploid wheat (Triticum aestivum L.) by QTL meta-analysis. Mol Breed 23:473–488
Malik S, Malik TA (2015) Genetic mapping of potential QTLs associated with drought tolerance in wheat. J Anim Plant Sci 25: 1032–1040
Marandel G, Salava J, Abbott A, Candresse T, Decroocq V (2009) Quantitative trait loci meta-analysis of Plum pox virus resistance in apricot (Prunus armeniaca L.) new insights on the organization and the identification of genomic resistance factors. Mol Plant Pathol 10:347–360
Mathews KL, Malosetti M, Chapman S, McIntyre L, Reynolds M, Shorter R et al (2008) Multi-environment QTL mixed models for drought stress adaptation in wheat. Theor Appl Genet 117:1077–1091
McIntyre CL, Mathews KL, Rattey A, Chapman SC, Drenth J, Ghaderi M et al (2010) Molecular detection of genomic regions associated with grain yield and yield-related components in an elite bread wheat cross evaluated under irrigated and rainfed conditions. Theor Appl Genet 120:527–541
Metsalu T, Vilo J (2015) ClustVis: a web tool for visualising clustering of multivariate data using principal component analysis and heatmap. Nucleic Acids Res 43:426–570
Min DH, Zhao Y, Huo DY, Li LC, Chen M, Xu ZS, Ma YZ (2013) Isolation and identification of a wheat gene encoding a zinc finger protein (TaZnFP) responsive to abiotic stresses. Acta Physiol Plant 35:1597–1604
Nezhadahmadi A, Zakaria HP, Golam F (2013) Drought tolerance in wheat. Sci World J 2013:610721
Norton GJ, Aitkenhead MJ, Khowaja FS, Whalley WR, Price AH (2008) A bioinformatic and transcriptomic approach to identifying positional candidate genes without fine mapping: an example using rice root-growth QTLs. Genomics 92:344–352
Onyemaobi I, Ayalew H, Liu H, Siddique KH, Yan G (2018) Identification and validation of a major chromosome region for high grain number per spike under meiotic stage water stress in wheat (Triticum aestivum L.). PLoS ONE 13:3
Peleg Z, Fahima T, Krugman T, Abbo S, Yakir D, Korol AB et al (2009) Genomic dissection of drought resistance in durum wheat × wild emmer wheat recombinant inbred line population. Plant, Cell Environ 32:758–779
Pinto RS, Reynolds MP, Mathews KL, McIntyre CL, Olivares-Villegas JJ, Chapman SC (2010) Heat and drought adaptive QTL in a wheat population designed to minimize confounding agronomic effects. Theor Appl Genet 121:1001–1021
Quraishi UM, Murat F, Abrouk M, Pont C, Confolent C, Oury FX, Ward J, Boros D, Gebruers K, Delcour JA, Courtin CM, Bedo Z, Saulnier L, Guillon F, Balzergue S, Shewry PR, Feuillet C, Charmet G, Salse J (2010) Combined meta-genomics analyses unravel candidate genes for the grain dietary fiber content in bread wheat (Triticum aestivum L.). Funct Integr Genomics 11:71–83
Quraishi UM, Abrouk M, Murat F et al (2011) Cross-genome map based dissection of a nitrogen use efficiency ortho-metaQTL in bread wheat unravels concerted cereal genome evolution. Plant J 65:745–756
Quraishi UM, Pont C, Ain Q-U, Flores R, Burlot L, Alaux M, Quesneville H, Salse J (2017) Combined genomic and genetic data integration of major agronomical traits in bread wheat (Triticum aestivum L.). Front Plant Sci 8:1843
Rong J, Feltus FA, Waghmare VN, Pierce GJ, Chee PW, Draye X, Saranga Y, Wright RJ, Wilkins TA, May OL, Smith CW (2007) Meta-analysis of polyploid cotton QTL shows unequal contributions of subgenomes to a complex network of genes and gene clusters implicated in lint fiber development. Genetics 176:2577–2588
Sakamoto Y, Ishiguro M, Kitagawa G (1986) Akaike information criterion statistics. D. Reidel, Dordrecht, p 81
Shabala S, Pottosin I (2014) Regulation of potassium transport in plants under hostile conditions: implications for abiotic and biotic stress tolerance. Physiol Plant 151:257–279
Shi J, Li R, Qiu D, Jiang C, Long Y, Morgan C, Bancroft I, Zhao J, Meng J (2009) Unraveling the complex trait of crop yield with quantitative trait loci mapping in Brassica napus. Genetics 182:851–861
Shirasawa K, Bertioli DJ, Varshney RK, Moretzsohn MC, Leal-Bertioli SC, Thudi M, Pandey MK, Rami JF, Foncéka D, Gowda MV, Qin H (2013) Integrated consensus map of cultivated peanut and wild relatives reveals structures of the A and B genomes of Arachis and divergence of the legume genomes. DNA Res 20:173–184
Somers DJ, Isaac P, Edwards K (2004) A high-density microsatellite consensus map for bread wheat (Triticum aestivum L.). Theor Appl Genet 109:1105–1114
Soriano JM, Royo C (2015) Dissecting the genetic architecture of leaf rust resistance in wheat by QTL meta-analysis. Phytopathology 105:1585–1593
Sun YN, Pan JB, Shi XL, Du XY, Wu Q, Qi ZM, Jiang HW, Xin DW, Liu CY, Hu GH, Chen QS (2012) Multi-environment mapping and meta-analysis of 100-seed weight in soybean. Mol Biol Rep 39:9435–9443
Swamy BM, Vikram P, Dixit S, Ahmed HU, Kumar A (2011) Meta-analysis of grain yield QTL identified during agricultural drought in grasses showed consensus. BMC Genom 12:319
Truntzler M, Barrière Y, Sawkins MC, Lespinasse D, Betran J, Charcosset A, Moreau L (2010) Meta-analysis of QTL involved in silage quality of maize and comparison with the position of candidate genes. Theor Appl Genet 121:1465–1482
Tuberosa R (2012) Phenotyping for drought tolerance of crops in the genomics era. Front Physiol 3:347
Tyagi S, Gupta PK (2012) Meta-analysis of QTLs involved in pre-harvest sprouting tolerance and dormancy in bread wheat. Triticeae Genom Genet 3:9–24
Verma V, Foulkes MJ, Worland AJ, Sylvester-Bradley R, Caligari PDS, Snape JW (2004) Mapping quantitative trait loci for flag leaf senescence as a yield determinant in winter wheat under optimal and drought-stressed environments. Euphytica 135:255–263
Wang Y, Yao J, Zhang ZF, Zheng YL (2006) The comparative analysis based on maize integrated QTL map and meta-analysis of plant height QTLs. Chin Sci Bull 51:2219–2230
Wu F, Sheng P, Tan J, Chen X, Lu G, Ma W, Heng Y, Lin Q, Zhu S, Wang J, Wang J, Guo X, Zhang X, Lei C, Wan J (2015) Plasma membrane receptor-like kinase leaf panicle 2 acts downstream of the DROUGHT AND SALT TOLERANCE transcription factor to regulate drought sensitivity in rice. J Exp Bot 66:271–281
Xu J, Dai X, Ramasamy RK, Wang L, Zhu T, McGuire PE, Jorgensen CM, Dehghani H, Gulick PJ, Luo MC, Müller HG (2019) Aegilops tauschii genome sequence: a framework for meta-analysis of wheat QTLs. G3 Genes Genomes Genet 9:841–853
Yang D, Liu Y, Cheng H, Chang L, Chen J, Chai S, Li M (2016) Genetic dissection of flag leaf morphology in wheat (Triticum aestivum L) under diverse water regimes. BMC Genet 17:94
Zhang LY, Liu DC, Guo XL, Yang WL, Sun JZ, Wang DW, Zhang A (2010) Genomic distribution of quantitative trait loci for yield and yield-related traits in common wheat. J Integr Plant Biol 52:996–1007
Zhao X, Peng Y, Zhang J, Fang P, Wu B (2018) Identification of QTLs and meta-QTLs for seven agronomic traits in multiple maize populations under well-watered and water-stressed conditions. Crop Sci 58:507–520
Zheng BS, Le Gouis J, Leflon M, Rong WY, Laperche A, Brancourt- Hulmel M (2010) Using probe genotypes to dissect QTL environment interactions for grain yield components in winter wheat. Theor Appl Genet 121:1501–1517
Acknowledgements
The work was carried out, when AK, IJ and KK held JRF positions under a research project funded by Department of Biotechnology, New Delhi, India and GS held SRF position under NASF-ICAR program of Government of India. PKG was awarded Hony Scientist position and HSB was awarded Senior Scientist position both from Indian National Science Academy (INSA). Head, Department of Genetics and Plant Breeding, CCS University, Meerut, provided the necessary infrastructure. Authors are also thankful to Dr. Sandhya Tyagi, Division of Genetics, IARI, New Delhi for providing inputs to AK for conducting MQTL analysis.
Author information
Authors and Affiliations
Contributions
PKG, HSB and PKS conceived the study and also edited and finalized the manuscript. AK conducted MQTL analysis and wrote the first draft of the MS jointly with GS; IJ and KK helped AK in preparing the files for MQTL analysis.
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kumar, A., Saripalli, G., Jan, I. et al. Meta-QTL analysis and identification of candidate genes for drought tolerance in bread wheat (Triticum aestivum L.). Physiol Mol Biol Plants 26, 1713–1725 (2020). https://doi.org/10.1007/s12298-020-00847-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12298-020-00847-6