Abstract
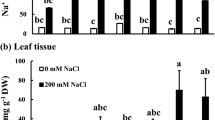
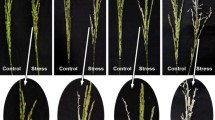
Salt affected soil inhibits plant growth, development and productivity, especially in case of rice crop. Ion homeostasis is a candidate defense mechanism in the salt tolerant plants or halophyte species, where the salt toxic ions are stored in the vacuoles. The aim of this investigation was to determine the OsNHX1 (a vacuolar Na+/H+ exchanger) and OsHKT2;1 (Na+/K+ transporter) regulation by salt stress (200 mM NaCl) in two rice cultivars, i.e. Pokkali (salt tolerant) and IR29 (salt susceptible), the accumulation of Na+ in the root and leaf tissues using CoroNa Green® staining dye and the associated physiological changes in test plants. Na+ content was largely increased in the root tissues of rice seedlings cv. Pokkali (15 min after salt stress) due to the higher expression of OsHKT2;1 gene (by 2.5 folds) in the root tissues. The expression of OsNHX1 gene in the leaf tissues was evidently increased in salt stressed seedlings of Pokkali, whereas it was unchanged in salt stressed seedlings of IR29. Na+ in the root tissues of both Pokkali and IR29 was enriched, when subjected to 200 mM NaCl for 12 h and easily detected in the leaf tissues of salt stressed plants exposed for 24 h, especially in cv. Pokkali. Moreover, the overexpression of OsNHX1 gene regulated the translocation of Na+ from root to leaf tissues, and compartmentation of Na+ into vacuoles, thereby maintaining the photosynthetic abilities in cv. Pokkali. Overall growth performance, maximum quantum yield (Fv/Fm), photon yield of PSII (ΦPSII) and net photosynthetic rate (Pn) was improved in salt stressed leaves of Pokkali than those in salt stressed IR29.





Similar content being viewed by others
References
Almeida P, Katschnig D, de Boer A (2013) HKT transporters—state of the art. Int J Mol Sci 14:20359–20385
Basu S, Roychoudhury A (2014) Expression profiling of abiotic stress-inducible genes in response to multiple stresses in rice (Oryza sativa L.) varieties with contrasting level of stress tolerance. BioMed Res Int. Article ID 706890
Basu S, Giri RK, Benazir I, Kumar S, Rajwanshi R, Dwivedi SK, Kumar G (2017) Comprehensive physiological analyses and reactive oxygen species profiling in drought tolerant rice genotypes under salinity stress. Physiol Mol Biol Plants 23:837–850
Bertazzini M, Sacchi GA, Forlani G (2018) A differential tolerance to mild salt stress conditions among six Italian rice genotypes dose not rely on Na+ exclusion from shoots. J Plant Physiol 226:145–153
Cha-um S, Supaibulwatana K, Kirdmanee C (2006) Water relation, photosynthetic ability and growth of Thai jasmine rice (Oryza sativa L. ssp. indica cv. KDML 105) to salt stress by application of exogenous glycinebetaine and choline. J Agron Crop Sci 192:25–36
Fu L, Shen Q, Kuang L, Yu J, Wu D, Zhang G (2018) Metabolite profiling and gene expression of Na/K transporter analyses reveal mechanisms of the difference in salt tolerance between barley and rice. Plant Physiol Biochem 130:248–257
Fujiwara K, Kozai T, Watanabe L (1987) Fundamental studies on environment in plant tissue culture vessel. (3) Measurement of carbon dioxide gas concentration in closed vessels containing tissue cultured plantlets and estimates of net photosynthetic rates of the plantlets. J Agric Methodol 43:21–30
Garciadeblás B, Senn ME, Bañuelos MA, Rodríguez-Navarro A (2003) Sodium transport and HKT transporters: the rice model. Plant J 34:788–801
Ghosh B, Ali NM, Gantait S (2016) Response of rice under salinity stress: a review update. J Res Rice 4:2
Grattan S, Zeng L, Shannon M, Roberts S (2002) Rice is more sensitive to salinity than previously thought. Calif Agric 56:189–198
Hamam AM, Coskun D, Britto DT, Plett D, Kronzucker HJ (2018) Plasma-membrane electrical responses to salt and osmotic gradients contradict radiotracer kinetics, and reveal Na+-transport dynamics in rice (Oryza sativa L.). Planta. https://doi.org/10.1007/s00425-018-3059-7
Han Y, Yin S, Huang L (2015) Towards plant salinity tolerance-implications from ion transporters and biochemical regulation. Plant Growth Regul 76:13–23
Hazman M, Hause B, Eiche E, Riemann M, Nick P (2016) Different forms of osmotic stress evoke qualitatively different responses in rice. J Plant Physiol 202:45–56
Horie T, Costa A, Kim TH, Han MJ, Horie R, Leung HY, Miyao A, Hirochika H, An G, Schroeder JI (2007) Rice OsHKT2;1 transporter mediates large Na+ influx component into K+-starved roots for growth. EMBO J 26:3003–3014
Horie T, Karahara I, Katsuhara M (2012) Salinity tolerance mechanisms in glycophytes: an overview with the central focus on rice plants. Rice 5:11
Hossain GS, Waditee R, Hibino T, Tanaka Y, Takabe T (2006) Root specific expression of Na+/H+ antiporter gene from Synechocystis sp. PCC6803 confers salt tolerance of tobacco plant. Plant Biotechnol 23:275–281
Hossain MM, Imran S, Islam MA, Kader MA, Uddin MI (2017) Expressional analysis of OsNHX1, OsNHX2, OsSOS1 and OsDREB transporters in salt tolerant (FR13A) and salt sensitive rice (BRRI dhan29) induced by salinity stress. IOSR J Agric Veter 10:64–70
Huang S, Spielmeyer W, Lagudah ES, Munns R (2008) Comparative mapping of HKT genes in wheat, barley, and rice, key determinants of Na+ transport, and salt tolerance. J Exp Bot 59:927–937
Hussain S, Zhang JH, Zhong C, Zhu LF, Cao XC, Yu SM, James AB, Hu JJ, Jin QY (2017) Effects of salt stress on rice growth, development characteristics, and the regulating ways: a review. J Integr Agric 16:2357–2374
Hussain M, Ahmad S, Hussain S, Lal R, Ul-Allah S, Nawaz A (2018) Rice in saline soils: physiology, biochemistry, genetics, and management. Adv Agron 148:231–287
Jabnoune M, Espeout S, Mieulet D, Fizames C, Verdeil JL, Conéjéro G, Rodríguez-Navarro A, Sentenac H, Guiderdoni E, Abdelly C, Véry AA (2009) Diversity in expression patterns and functional properties in the rice HKT transporter family. Plant Physiol 150:1955–1971
Jesus JM, Danko AS, Fiúza A, Borges MT (2015) Phytoremediation of salt-affected soils: a review of processes, applicability, and the impact of climate change. Environ Sci Pollut Res 22:6511–6525
Julkowska MM, Testerink C (2015) Tuning plant signaling and growth to survive salt. Trend Plant Sci 20:586–594
Kader MA, Lindberg S (2005) Uptake of sodium in protoplasts of salt-sensitive and salt-tolerant cultivars of rice, Oryza sativa L. determined by the fluorescent dye SBFI. J Exp Bot 56:3149–3158
Kader MA, Seidel T, Golldack D, Lindberg S (2006) Expression of OsHKT1, OsHKT2, and OsVHA are differentially regulated under NaCl stress in salt-sensitive and salt-tolerant rice (Oryza sativa L.) cultivars. J Exp Bot 57:4257–4268
Khare T, Kumar V, Kishor PBK (2015) Na+ and Cl− ions show additive effects under NaCl stress on induction of oxidative stress and the responsive antioxidative defense in rice. Protoplasma 252:1149–1165
Krishnamurthy SL, Sharma PC, Sharma DK, Ravikiran KT, Singh YP, Mishra VK, Burman D, Maji B, Mandal S, Sarangi SK, Gautam RK, Singh PK, Manohara KK, Marandi BC, Padmavathi G, Vanve PB, Patil KD, Thirumeni S, Verma OP, Khan AH, Tiwari S, Geetha S, Shakila M, Gill R, Yadav VK, Roy SKB, Prakash M, Bonifacio J, Ismail A, Gregorio GB, Singh RK (2017) Identification of mega-environments and rice genotypes for general and specific adaptation to saline and alkaline stresses in India. Sci Rep 7:7968
Kumar K, Mosa KA (2015) Ion transporters: a decisive component of salt stress tolerance in plants. In: Wani SH, Anwar M (eds) Managing salt tolerance in plants: molecular and genomic perspectives. CRC Press, Taylor & Francis Group, Boca Raton, pp 373–390
Lakra N, Kaur C, Singla-Pareek SL, Pareek A (2019) Mapping the ‘early salinity response’ triggered proteome adaptation in contrasting rice genotypes using iTRAQ approach. Rice 12:3
Lichtenthaler HK (1987) Chlorophylls and carotenoids: pigments of photosynthetic biomembranes. Method Enzymol 148:350–380
Loggini B, Scartazza A, Brugnoli E, Navari-Izzo F (1999) Antioxidant defense system, pigment composition, and photosynthetic efficiency in two wheat cultivars subjected to drought. Plant Physiol 119:1091–1099
Maathuis FJM (2014) Sodium in plants: perception, signaling, and regulation of sodium fluxes. J Exp Bot 65:849–858
Maxwell K, Johnson GN (2000) Chlorophyll fluorescence-a practical guide. J Exp Bot 51:659–668
Mekawy AMM, Assaha DVM, Yahaki H, Tada Y, Ueda A, Seneoka H (2015) Growth, physiological adaptation, and gene expression analysis of two Egyptian rice cultivars under salt stress. Plant Physiol Biochem 87:17–25
Miyamoto T, Ochiai K, Nonoue Y, Matsubara K, Yano M, Matoh T (2015) Expression level of the sodium transporter gene OsHKT2;1 determines sodium accumulation of rice cultivars under potassium-deficient conditions. Soil Sci Plant Nutr 61:481–492
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol Plant 15:473–497
Ochiai K, Matoh T (2002) Characterization of the Na+ delivery from roots to shoots in rice under saline stress: excessive salt enhances apoplastic transport in rice plants. Soil Sci Plant Nutr 48:371–378
Oh DH, Leidi E, Zhang Q, Hwang SM, Li Y, Quintero FJ, Jiang X, D’Urzo MP, Lee SY, Zhao Y, Bahk JD, Bressan RA, Yun DJ, Pardo JM, Bohnert HJ (2009) Loss of halophytism by interference with SOS1 expression. Plant Physiol 151:210–222
Omisun T, Sahoo S, Saha B, Panda SK (2018) Relative salinity tolerance of rice cultivars native to North East India: a physiological, biochemical and molecular perspective. Protoplasma 255:193–202
Parihar P, Singh S, Singh R, Singh VP, Prasad SM (2015) Effect of salinity stress on plans and its tolerance strategies: a review. Environ Sci Pollut Res 22:4056–4075
Pitman M, Läuchli A (2004) Global impact of salinity and agricultural ecosystems. In: Läuchli A, Lüttge U (eds) Salinity: environment–plants–molecules. Springer, Dordrecht, pp 3–20
Plett DC, Møller IS (2010) Na+ transport in glycophytic plants: what we know and would like to know. Plant Cell Environ 33:612–626
Qadir M, Oster J (2002) Vegetative bioremediation of calcareous sodic soils: history, mechanisms, and evaluation. Irrig Sci 21:91–101
Rahman MA, Thomson MJ, Shah-E-Alam M, de Ocampo M, Egdane J, Ismail AM (2016) Exploring novel genetic sources of salinity tolerance in rice through molecular and physiological characterization. Ann Bot 117:1083–1097
Reddy INBL, Kim BK, Yoon IS, Kim KH, Kwon TR (2017) Salt tolerance in rice: focus on mechanisms and approaches. Rice Sci 24:123–144
Rengasamy P (2010) Soil processes affecting crop production in salt-affected soil. Funct Plant Biol 37:613–620
Roy SJ, Negrão S, Tester M (2014) Salt resistant crop plants. Curr Opin Biotechnol 26:115–124
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory, Cold Spring Harbor
Shabala SN, Shabala SI, Martynenko AI, Babourina O, Newman IA (1998) Salinity effect on bioelectric activity, growth, Na+ accumulation and chlorophyll fluorescence of maize leaves: a comparative survey and prospects for screening. Aust J Plant Physiol 25:609–616
Shabala S, Shabala S, Cuin TA, Pang J, Percey W, Chen Z, Conn S, Eing C, Wegner LH (2010) Xylem ionic relations and salinity tolerance in barley. Plant J 62:839–853
Shabala S, Wu H, Bose J (2015) Salt stress sensing and early signalling events in plant roots: current knowledge and hypothesis. Plant Sci 241:109–119
Shakeela BS, Chachar QI, Chachar SD, Soloangi AB, Solangi JA (2016) Effect of salinity (NaCl) stress on physiological characteristics of rice (Oryza sativa L.) at early seedling stage. Int J Agric Technol 12:263–279
Shen G, Wei J, Qiu X, Hu R, Kuppu S, Auld D, Blumwald E, Gaxiola R, Payton P, Zhang H (2015) Co-overexpression of AVP1 and AtNHX1 in cotton further improves drought and salt tolerance in transgenic cotton plants. Plant Mol Biol Rep 33:167–177
Singh V, Singh AP, Bhadoria J, Giri J, Singh J, Vineeth TV, Sharma PC (2018) Differential expression of salt-tolerant and salt sensitive rice (Oryza sativa L.) at seedling stage. Protoplasma 255:1667–1681
Tanaka K, Ohta K, Haddad PR, Fritz JS, Lee KP, Hasebe K, Ieuji A, Miyanaga A (1999) Acid-rain monitoring in East Asia with a portable-type ion-exclusion-cation-exchange chromatographic analyzer. J Chromatogr 850:311–317
Theerawitaya C, Yamada N, Samphumphuang T, Cha-um S, Kirdmanee C, Takabe T (2015) Evaluation of Na+ enrichment and expression of some carbohydrate related genes in indica rice seedlings under salt stress. Plant Omic J 8:130–140
Wang W, Vinocur B, Altman A (2003) Plant responses to drought, salinity and extreme temperatures: towards genetic engineering for stress tolerance. Planta 218:1–14
Wang H, Zhang MS, Guo R, Shi DC, Liu B, Lin XY, Yang CW (2012) Effects of salt stress on ion balance and nitrogen metabolism of old and young leaves in rice (Oryza sativa L.). BMC Plant Biol 12:194
Wu H (2018) Plant salt tolerance and Na+ sensing and transport. Crop J 6:215–225
Wu Y, Hu Y, Xu G (2009) Interactive effects of potassium and sodium on root growth and expression of K/Na transporter genes in rice. Plant Growth Regul 57:271–280
Yao X, Horie T, Xue S, Leung HY, Katsuhara M, Brodsky DE, Wu Y, Schroeder JI (2010) Differential sodium and potassium transport selectivities of the rice OsHKT2;1 and OsHKT2;2 transporters in plant cells. Plant Physiol 152:341–355
Yasmin F, Biswas S, Nurnabi GM, Jewel NA, Elias SM, Seraj ZI (2015) Constitutive overexpression of the plasma membrane Na+/H+ antiporter for conferring salinity tolerance in rice. Plant Tiss Cult Biotechnol 25:257–272
Zhang Y, Fang J, Wu X, Dong L (2018) Na+/K+ balance and transport regulatory mechanisms in weedy and cultivated rice (Oryza sativa L.) under salt stress. BMC Plant Biol 18:375
Zhu M, Zhou M, Shabala L, Shabala S (2017) Physiological and molecular mechanisms mediating xylem Na+ loading in barley in the context salinity stress tolerance. Plant Cell Environ 40:1009–1020
Acknowledgement
The funding was provided by BIOTEC (Grant number P1750079).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Theerawitaya, C., Tisarum, R., Samphumphuang, T. et al. Expression levels of the Na+/K+ transporter OsHKT2;1 and vacuolar Na+/H+ exchanger OsNHX1, Na enrichment, maintaining the photosynthetic abilities and growth performances of indica rice seedlings under salt stress. Physiol Mol Biol Plants 26, 513–523 (2020). https://doi.org/10.1007/s12298-020-00769-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12298-020-00769-3