Abstract
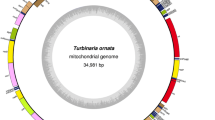
Chlorosarcinopsis eremi is a member of Chlamydomonadales algae which is isolated from terrestrial environments. In this study, the mitochondrial genome of C. eremi isolated from desert region of Iran, was represented for the first time. Following sequencing, assembly and annotation, comparative analyses of C. eremi and other available Chlamydomonadales algae complete mitochondrial genomes were performed. The mitochondrial genome of C. eremi was circular, had a low number of genes coding in the same strand with a minor amount of repeated sequences; same as other non-Reinhardtinia species of Chlamydomonadales algae. GC content of C. eremi mitochondrial genome was in normal range when compared with non-Chlamydomonadales organisms, but among Chlamydomonadales algae, C. eremi had a low GC content mitochondrial genome. C. eremi had the highest percent of non-coding sequences in comparison with other available Chlamydomonadales mitochondrial genomes which was related to intergenic regions. Identity analysis of protein-coding sequences of Chlamydomonadales mitochondrial genomes showed more divergences and may be related to the high mutation rate of mitochondrial genome reported in microbial eukaryotes.





Similar content being viewed by others
References
Bankevich A, Nurk S, Antipov D et al (2012) SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol 19:455–477
Buchheim MA, Lemieux C, Otis C et al (1996) Phylogeny of the Chlamydomonadales (Chlorophyceae): a comparison of ribosomal RNA gene sequences from the nucleus and the chloroplast. Mol Phylogenet Evol 5:391–402
Burger G, Gray MW, Lang BF (2003) Mitochondrial genomes: anything goes. Trends Genet 19:709–716
Camacho C, Madden T, Coulouris G et al (2008) BLAST command line applications user manual
Castresana J (2000) Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol 17:540–552
Chantanachat S, Bold HC (1962) Some algae from arid soils, vol 2. University of Texas, Texas
Cherdchukeattisak P, Fraser PD, Purton S et al (2018) Detection and enhancement of ketocarotenoid accumulation in the newly isolated sarcinoid green microalga chlorosarcinopsis PY02. Biology. https://doi.org/10.3390/biology7010017
Darling AE, Mau B, Perna NT (2010) progressiveMauve: multiple genome alignment with gene gain, loss and rearrangement. PLoS ONE 5:e11147
Del Vasto M, Figueroa-Martinez F, Featherston J et al (2015) Massive and widespread organelle genomic expansion in the green algal genus dunaliella. Genome Biol Evol 7:656–663. https://doi.org/10.1093/gbe/evv027
DePriest MS, Bhattacharya D, López-Bautista JM (2014) The mitochondrial genome of Grateloupia taiwanensis (Halymeniaceae, Rhodophyta) and comparative mitochondrial genomics of red algae. Biol Bull 227:191–200
Dhanalakshmi M (2013) Phytochemistry and antibacterial activity of Chlorosarcinopsis species. IJSTR 2:315–321
Dierckxsens N, Mardulyn P, Smits G (2016) NOVOPlasty: de novo assembly of organelle genomes from whole genome data. Nucleic Acids Res 45:e18–e18
Friedman JR, Nunnari J (2014) Mitochondrial form and function. Nature 505:335
Fulnečková J, Hasíková T, Fajkus J et al (2012) Dynamic evolution of telomeric sequences in the green algal order Chlamydomonadales. Genome Biol Evol 4:248–264
Gärtner G, Hofer A, Ingolić E (1988) Morphological and Taxonomical Observations on Some Strains of Chlorosarcina, Chlorosarcinopsis and Planophila (Chlorophyta, Chlorosarcinales), with Special Reference to “Vegetative Cell Division”. Arch Protistenkunde 135:119–131. https://doi.org/10.1016/S0003-9365(88)80058-7
Gouy M, Guindon S, Gascuel O (2009) SeaView version 4: a multiplatform graphical user interface for sequence alignment and phylogenetic tree building. Mol Biol Evol 27:221–224
Guiry M, Guiry G (2018) AlgaeBase. World-wide electronic publication National University of Ireland. http://www.algaebase.org. Accessed 30 May 2018
Haag-Liautard C, Coffey N, Houle D et al (2008) Direct estimation of the mitochondrial DNA mutation rate in Drosophila melanogaster. PLoS Biol 6:e204
Hadi SI, Santana H, Brunale PP et al (2016) DNA barcoding green microalgae isolated from neotropical inland waters. PLoS ONE 11:e0149284
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucl Acids Symp Ser 41:95–98
Hall J, Fučíková K, Lo C et al (2010) An assessment of proposed DNA barcodes in freshwater green algae. Cryptogam Algol 31:529–555
Hamaji T, Smith DR, Noguchi H et al (2013) Mitochondrial and plastid genomes of the colonial green alga Gonium pectorale give insights into the origins of organelle DNA architecture within the volvocales. PLoS ONE 8:e57177. https://doi.org/10.1371/journal.pone.0057177
Hamaji T, Kawai-Toyooka H, Toyoda A et al (2017) Multiple independent changes in mitochondrial genome conformation in chlamydomonadalean algae. Genome Biol Evol 9:993–999. https://doi.org/10.1093/gbe/evx060
Hebert PD, Ratnasingham S, de Waard JR (2003) Barcoding animal life: cytochrome c oxidase subunit 1 divergences among closely related species. Proc Royal Soc Lond 270:S96–S99
Joshi N, Fass J (2011) Sickle: a sliding-window, adaptive, quality-based trimming tool for FastQ files (Version 1.33)[Software]
Juy-abad FK, Mohammadi P, Zarrabi M (2018) The identification of some phototrophic microorganisms from a semi-arid ecosystem in Iran. C R Acad Bulg 71
Kim KM, Park J-H, Bhattacharya D et al (2014) Applications of next-generation sequencing to unravelling the evolutionary history of algae. Int J Syst Evol Microbiol 64:333–345
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Kurtz S, Choudhuri JV, Ohlebusch E et al (2001) REPuter: the manifold applications of repeat analysis on a genomic scale. Nucleic Acids Res 29:4633–4642
Lang BF, Laforest M-J, Burger G (2007) Mitochondrial introns: a critical view. Trends Genet 23:119–125
Le Gall L, Saunders GW (2010) DNA barcoding is a powerful tool to uncover algal diversity: a case study of the Phyllophoraceae (Gigartinales, Rhodophyta) in the Canadian flora 1. J Phycol 46:374–389
Lemieux C, Vincent AT, Labarre A et al (2015) Chloroplast phylogenomic analysis of chlorophyte green algae identifies a novel lineage sister to the Sphaeropleales (Chlorophyceae). BMC Evol Biol 15:264. https://doi.org/10.1186/s12862-015-0544-5
Letunic I, Bork P (2016) Interactive tree of life (iTOL) v3: an online tool for the display and annotation of phylogenetic and other trees. Nucleic Acids Res 44:W242–W245
Lewis LA, Lewis PO (2005) Unearthing the molecular phylodiversity of desert soil green algae (Chlorophyta). Syst Biol 54:936–947. https://doi.org/10.1080/10635150500354852
Lohse M, Drechsel O, Kahlau S et al (2013) OrganellarGenomeDRAW—a suite of tools for generating physical maps of plastid and mitochondrial genomes and visualizing expression data sets. Nucleic Acids Res 41:W575–W581
Muhammad Tahir H, Akhtar S (2016) Services of DNA barcoding in different fields. Mitochondrial DNA Part A 27:4463–4474
Nakada T, Misawa K, Nozaki H (2008) Molecular systematics of Volvocales (Chlorophyceae, Chlorophyta) based on exhaustive 18S rRNA phylogenetic analyses. Mol Phylogenet Evol 48:281–291. https://doi.org/10.1016/j.ympev.2008.03.016
Nguyen L-T, Schmidt HA, von Haeseler A et al (2014) IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol Biol Evol 32:268–274
Pogoda CS, Keepers KG, Hamsher SE et al (2018) Comparative analysis of the mitochondrial genomes of six newly sequenced diatoms reveals group II introns in the barcoding region of cox1. Mitochondrial DNA A DNA Mapp Seq Anal 1:1–9. https://doi.org/10.1080/24701394.2018.1450397
Prasad AKSK (1982) Notes on Soil Algae: chlorosarcinopsisHerndon (Chlorosarcinales, Chlorophyceae) in India. Arch Protistenkunde 126:273–282. https://doi.org/10.1016/s0003-9365(82)80038-9
Robba L, Russell SJ, Barker GL et al (2006) Assessing the use of the mitochondrial cox1 marker for use in DNA barcoding of red algae (Rhodophyta). Am J Bot 93:1101–1108
Ronquist F, Teslenko M, Van Der Mark P et al (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61:539–542
Rupprecht J (2009) From systems biology to fuel—Chlamydomonas reinhardtii as a model for a systems biology approach to improve biohydrogen production. J Biotechnol 142:10–20
Ševčíková T, Klimeš V, Zbránková V et al (2016) A comparative analysis of mitochondrial genomes in eustigmatophyte algae. Genome Biol Evol 8:705–722
Singh SP, Rastogi RP, Häder D-P et al (2010) An improved method for genomic DNA extraction from cyanobacteria. World J Microbiol Biotechnol 27:1225–1230. https://doi.org/10.1007/s11274-010-0571-8
Smith DR (2015) Mutation rates in plastid genomes: they are lower than you might think. Genome Biol Evol 7:1227–1234
Smith DR (2016) The past, present and future of mitochondrial genomics: have we sequenced enough mtDNAs? Brief Funct Genom 15:47–54. https://doi.org/10.1093/bfgp/elv027
Smith DR, Lee RW (2008) Mitochondrial genome of the colorless green alga Polytomella capuana: a linear molecule with an unprecedented GC content. Mol Biol Evol 25:487–496. https://doi.org/10.1093/molbev/msm245
Smith D, Lee RW, Cushman JC et al (2010) The Dunaliella salina organelle genomes: large sequences, inflated with intronic and intergenic DNA. BMC Plant Biol 10:83. https://doi.org/10.1186/1471-2229-10-83
Smith DR, Hua J, Archibald JM et al (2013) Palindromic genes in the linear mitochondrial genome of the nonphotosynthetic green alga Polytomella magna. Genome Biol Evol 5:1661–1667
Tan J, Lim P-E, Phang S-M et al (2012) Assessment of four molecular markers as potential DNA barcodes for red algae Kappaphycus Doty and Eucheuma J. Agardh (Solieriaceae, Rhodophyta). PLoS ONE 7:e52905
Thompson JD, Gibson TJ, Higgins DG (2003) Multiple sequence alignment using ClustalW and ClustalX. Curr Protoc Bioinformatics 1:2–3
Tillich M, Lehwark P, Pellizzer T et al (2017) GeSeq–versatile and accurate annotation of organelle genomes. Nucleic Acids Res 45:W6–W11
Trivedi S, Aloufi AA, Ansari AA et al (2016) Role of DNA barcoding in marine biodiversity assessment and conservation: an update. Saudi J Biol Sci 23:161–171
Watanabe S, Mitsui K, Nakayama T et al (2006) Phylogenetic relationships and taxonomy of Sarcinoid Green Algae: chlorosarcinopsis, Desmotetra, Sarcinochlamys Gen. Nov., Neochlorosarcina, and Chlorosphaeropsis (Chlorophyceae, Chlorophyta)1. J Phycol 42:679–695. https://doi.org/10.1111/j.1529-8817.2006.00196.x
Wolfsberg TG, Schafer S, Tatusov RL et al (2001) Organelle genome resources at NCBI. Trends Biochem Sci 26:199–203
Wongsnansilp T, Tansakul P, Arunyanart M (2007) Factors affecting growth and betacarotene content of Chlorosarcinopsis sp. (PSU/CHL20) in batch culture. Kasetsart J Nat Sci 41:153–157
Yang EC, Kim KM, Kim SY et al (2015) Highly conserved mitochondrial genomes among multicellular red algae of the Florideophyceae. Genome Biol Evol 7:2394–2406
Yumoto K, Kasai F, Kawachi M (2013) Taxonomic re-examination of Chlamydomonas strains maintained in the NIES-collection. Microbiol Cult Collect 29:1–12
Acknowledgements
The authors would like to thank Dr Sayed-Amir Marashi for his cooperation to provide his Systems Biology laboratory’s facilities. This project was carried out at Shayesteh Sepehr Laboratories and funded by Vice Chancellor of Alzahra University.
Author information
Authors and Affiliations
Contributions
Parisa Mohammadi was responsible for the study concept. Acquisition of data including DNA extraction and bioinformatics analysis was done by Fatemeh Khani-Juyabad. The interpretation of data was implemented by Fatemeh Khani-Juyabad, Parisa Mohammadi and Mahbubeh Zarrabi. Fatemeh Khani-Juyabad prepared the draft of the manuscript. The critical revision of the manuscript for important intellectual content as well as supervision of the study was performed by Parisa Mohammadi and Mahbubeh Zarrabi. Administrative and material support was provided by Parisa Mohammadi.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Khani-Juyabad, F., Mohammadi, P. & Zarrabi, M. Comparative analysis of Chlorosarcinopsis eremi mitochondrial genome with some Chlamydomonadales algae. Physiol Mol Biol Plants 25, 1301–1310 (2019). https://doi.org/10.1007/s12298-019-00696-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12298-019-00696-y